Abstract
Background:
Muscle loss occurs in some conditions such as aging, sarcopenia, and cancer. The interaction between protein synthesis and degradation signaling components induced by high-intensity interval training (HIIT) is not well studied.Objectives:
The purpose of the present study was to simultaneously examine the effect of eight-week HIIT on the gene expression of both signaling components.Methods:
Sixteen male Wistar rats were randomly assigned to HIIT and non-exercise control groups. The HIIT group ran on a treadmill, five days/week for eight weeks, with 0º slope, including five interval sets of high and low intensity. Forty-eight hours after the last exercise session, dissected soleus muscles were stored at -80°C for later analyses.Results:
The gene expression of Akt1, mTORC1, and S6K1 were increased in the HIIT group compared with the control group (All P ≤ 0.031) concomitant with the suppression of eIF4EBP1. The results of the S6K1 and eIF4EBP1 mRNA were also confirmed by the Western blotting. According to the inhibitory effect of Akt1, the gene expressions of FoxO3a and, consequently, MuRF1 and LC3A were significantly inhibited (All P ≤ 0.003). Western blot analysis did not confirm the LC3A protein expression, which may underline the role of LC3A in autophagy to promote cell survival.Conclusions:
The intensities and durations of the exercise training protocol are sufficient to increase protein synthesis signaling components and especially inhibit the atrophy-related gene expression, which may lead to attenuating muscle loss and increasing muscle mass. Accordingly, it may be considered for rehabilitation and/or prevention of some conditions such as sarcopenia and cachexia.Keywords
Signaling pathway Gene and Protein Expression Hypertrophy Atrophy
1. Background
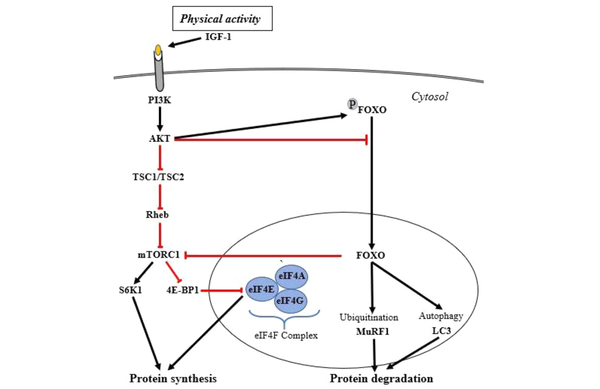
The percentage of the aging population is rising, and it is estimated that 80 million individuals will be over the age of 65 years by the year 2050. Sarcopenia, termed age-related skeletal muscle loss, which is a geriatric syndrome and public health issue that will spread between 10 to 20% and along with cachexia, muscle loss related to disease and immobility, may lead to disability, poor quality of life, and mortality. Skeletal muscle mass is controlled by the balance between muscle protein synthesis, termed “hypertrophy” and protein degradation, termed “atrophy”. As shown in Figure 1, IGF-1/PI3K/Akt signaling pathway regulates protein synthesis and muscle growth. IGF-1 activates the Akt through the activation of PI3K, and consequently, mTORC1 is activated by Akt to play a crucial role in protein synthesis and skeletal muscle growth by phosphorylating and activating the S6K1 (1, 2). In addition, mTORC1 phosphorylates and suppresses the activity of eIF4EBP1; otherwise, non-phosphorylated eIF4EBP1 inhibits the formation of the eIF4F complex (eIF4E, eIF4A, and eIF4G) by binding to the eIF4E (3). Furthermore, skeletal muscle protein degradation is controlled by two proteolytic systems, ubiquitin-proteasome, and autophagy-lysosomal pathways. FoxO transcription factors regulate both pathways by inducing two critical atrophy-related E3 ligases, MuRF1 and MAFbx, and also several autophagy-related genes, including MAP1LC3 (4). Activation of both systems, e.g., in some conditions such as cancer and heart failure, leads to protein degradation and skeletal muscle loss (5).
Protein synthesis and degradation signalings, including ubiquitination and autophagy. The left cascade is related to protein synthesis signaling, and the protein degradation signaling is presented in the right cascade. The red lines indicate the inhibitory effect of each molecule on protein synthesis, and the black arrows depict signaling components involved in the activation of protein synthesis. Abbreviations: IGF-1, Insulin-like growth factor 1; PI3K, Phosphatidylinositol 3‑kinase; Akt, Protein kinase B; TSC1/TSC2, Tuberous Sclerosis 1/2; Rheb, Ras homolog enriched in brain; mTOR, Mammalian Target of Rapamycin; S6K1 (RPS6K1), Ribosomal Protein S6 Kinase1; eIF4E eukaryotic translation Initiation Factor 4E; eIF4EBP1, eIF4E-Binding Protein 1 (also known as 4E-BP1); FOXO Forkhead box O; MuRF1, Muscle Ring Finger 1; MAFbx, Muscle Atrophy F-box.
Given that exercise training has beneficial effects on the prevention and/or improvement of some diseases such as obesity, hypertension, cachexia, and sarcopenia (6), the exercise training-induced alteration of gene expressions related to skeletal muscle protein synthesis and degradation signaling components have less been studied, especially simultaneously. Recently, a study investigated the effects of six-week high-intensity interval training (HIIT) on muscle atrophy-related genes and concluded HIIT may improve muscle atrophy (7). In addition, the effect of HIIT on both signaling components was recently examined in a study in which the exercise protocol was designed according to the studies of patients with heart failure (8). HIIT has been known as an effective modality for improving muscle size (9). Considering the lack of information, particularly gene mRNAs related to ubiquitin-proteasome and autophagy-lysosome system, simultaneously comparison the alteration and interaction of both protein synthesis and degradation signaling components may reveal the potential therapeutic target and/or help to design a more effective exercise training program for the prevention and/or rehabilitation of related diseases.
2. Objectives
The purpose of the present study was to investigate the alterations of Akt1, mTORC1, S6K1, eIF4EBP1, FoxO3a, MuRF1, MAP1LC3A (Hereafter referred to as LC3A) gene expression induced by HIIT. In addition, the protein expressions of S6K1, eIF4EBP1, and LC3A were evaluated by Western blotting for further confirmation of the mRNA results.
3. Methods
3.1. Study Design and Subjects
All research protocols were performed in accordance with the declaration of Helsinki 1975. Sixteen male Wistar rats, 8-month-old, were randomly divided into two groups, including HIIT (217.38 ± 10.88 gm) and control (219.86 ± 13.79 gm). They were kept, 4 rats per cage, in a light-dark 12 h cycle, 22 - 24°C temperature, 60 ± 5% humidity, with free access to standard rat chow and water. Two-week familiarization and adaptation to the main exercise protocol, at a gradual increase in running time and speed, was conducted. To determine exercise intensity, the maximal oxygen uptake (VO2max) test was performed as previously described and was repeated once every two weeks throughout the study (10, 11). Briefly, after 20 min warm-up at 50 - 60% VO2max, the velocity of the treadmill was increased by 0.03 m/s every two min until the rat was not able to run further.
3.2. Exercise Training Intervention
Eight-week of HIIT, five days per week, included five interval sets of high (85 - 90% VO2max) and low intensity (50 - 60% VO2max). The running time for high-intensity running on the treadmill with 0° slope was 4 min in the 1st week, 6 min in the 2nd week, and 8 min from 3rd to 8th week, whereas low-intensity running time was considered two min throughout the intervention. Each exercise session consisted of five min warm-up and cool-down at 50 - 60% VO2max (11, 12). An identical condition was considered for the control group except exercise training. Due to the death, the final number of seven rats remained in the control group. Forty-eight hours after the last exercise session, the soleus muscle was dissected from the hindlimb of each anesthetized rat and stored at -80°C for qRT-PCR and Western blot analyses.
3.3. Gene Expression
Quantitative real-time polymerase chain reaction (qRT-PCR) was used to assess the gene expression levels. Then 40 - 50 mg of soleus muscle was homogenized, and according to the manufacturer’s instruction (QIAGEN, Germany), total RNA was isolated using TRizol reagent (Qiazol, (cat. no. 79306, USA). RNA purity and concentration were determined spectrophotometrically by NanoDrop 2000 (Thermo Scientific, Rockford, USA), and RNA integrity was checked electrophoretically by 1% agarose gel stained with Nancy-520 (Sigma-Aldrich, Sao Paulo, SP, Brazil). Then cDNA was synthesized using a cDNA synthesis kit (Fermentas, Glen Burnie, MD, USA), and mRNA levels of the target genes and reference gene (GAPDH) were assessed separately by qRT-PCR. Forty cycles were considered for each RT-PCR cycle at the temperatures of 94°C maintained for 20s, 58 - 60°C for 30s, and 72°C for 30s. Melting point dissociation curves were used to confirm the purity of the amplification products (13). The sequences of the gene’s primer are shown in Table 1.
The Primer Sequences of Genes
| Gene name | Gene ID | Forward Sequence | Reverse Sequence |
|---|---|---|---|
| Akt1 | 24185 | 5’- TGTGGGAAGATGTGTATGAGAA -3’ | 5’- TTGATGAGGCGGTGTGATGGTGA -3’ |
| mTOR | 56718 | 5’- TGATTTTGGGAGAACAGAAGATGA -3’ | 5’- GAGGTAACAGGATGGTGGAGTG -3’ |
| S6K1 | 83840 | 5’- GTGTTGTGGATTGGTGGAGT -3’ | 5’- TTGTTGTGTGAGGTAGGGAGG -3’ |
| eIF4EBP1 | 116636 | 5’-GGACACAGTGATGCTTCT-3’ | 5’-GGAGGTATGTGCTGGTAT-3’ |
| FoxO3a | 294515 | 5’- GCCTCATCTCAAAGCTGGGT -3’ | 5’- TGCTCTGGAGTAGGGATGCT -3’ |
| MuRF1 | 140939 | 5’- GGTGTGACGAAGGTGGAAGAGA -3 | 5’- CAAGGCAGATGGGGCAGATGAG -3 |
| LC3A | 362245 | 5’- GGGTGGATTAGGCAGAGATGTGA -3 | 5’- GATGAGGGGCAAGATGGGTAGA -3 |
| GAPDH | 108351137 | 5’-AAGTTCAACGGCACAGTCAAGG-3’ | 5’-CATACTCAGCACCAGCATCACC-3’ |
3.4. Western Blot Analysis
Homogenized soleus muscles in RIPA buffer (Cytomatingene) with a protease inhibitor cocktail (Sigma) were centrifuged at 15,000 rpm for 10 min at 4°C. Through collected supernatant, protein content was assessed by the Lowry method. Then, proteins were separated by polyacrylamide gel electrophoresis (Bio-Rad) via 4 - 20% gradient polyacrylamide gels containing 0.1% sodium dodecyl sulfate for ~2 h at 95 V and were transferred to PVDF membranes (Roth) for 80 min at 80 V (Bio-Rad). Nonspecific sites were blocked overnight at 4°C in TBS containing Tween and 5% nonfat milk (sigma). Membranes were then incubated for 2h with primary antibodies directed against the proteins of interest at room temperature. The protein abundance of S6K1, eIF4EBP1, LC3A, and GAPDH (Served as a loading control to normalize protein loading and transfer) were determined in muscle samples. Following incubation with primary antibodies, membranes were washed extensively with PBS-Tween and then incubated with secondary antibodies 1h at room temperature. Then, membranes were developed using DAB, and related images were captured and analyzed using the Image J software (14).
3.5. Statistical Analysis
The results are expressed as mean ± standard error of the mean (SEM). Independent student t-test and Mann-Whitney U test were used to evaluate between-group differences. Statistical significance was accepted at P < 0.05 using SPSS version 24.
4. Results
In this experimental study, the alterations of gene expression involved in protein synthesis and degradation signaling pathways induced by eight-week HIIT were evaluated.
4.1. Protein Synthesis Signaling
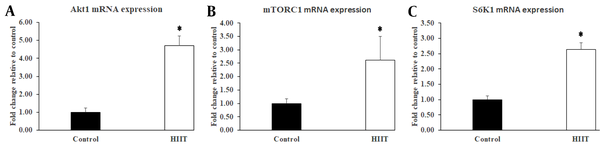
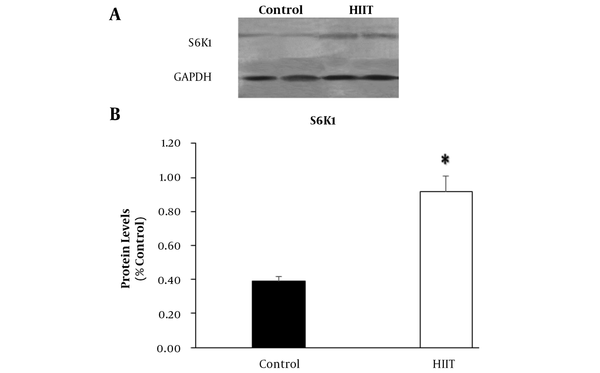
In the present study, the Akt-mTOR-S6K1 signaling pathway was investigated. As shown in Figure 1, compared with the control group, the expression of Akt1 was increased significantly (P < 0.001; F1,13 = 4.122, P = 0.063) in HIIT (Figure 2A), which was followed by a significant increase (P = 0.031; F1,13 = 8.272, P = 0.013) of mTORC1 expression (Figure 2B). Accordingly, the S6K1 expression levels were significantly increased (P < 0.001; F1,13 = 3.873, P = 0.071) in HIIT compared with the control group (Figure 2C). Furthermore, the S6k1 gene expression was confirmed by Western blot analysis (Figure 3A). Similar to the mRNA expression, the HIIT group possessed higher levels of S6K1 protein (P = 0.031) compared with the control group (Figure 3B).
Effect of eight-week HIIT on protein synthesis signaling components in rats’ soleus muscle. The exercise training protocol caused a significant increase in Akt1 (A), mTORC1 (B), and S6K1 (C) mRNA expressions in the HIIT group (n = 8). Data are expressed as mean ± SEM. * Indicates significant difference (P ≤ 0.031) compared with the control group (n = 7).
The protein expression of S6K1 and GAPDH in rats’ soleus muscle (A). The eight-week HIIT caused a significant increase of S6K1 protein levels in the HIIT group (n = 8). Data are expressed as mean ± SEM. * Indicates significant difference (P = 0.031) compared with the control group (n = 7).
4.2. Protein Degradation Signaling
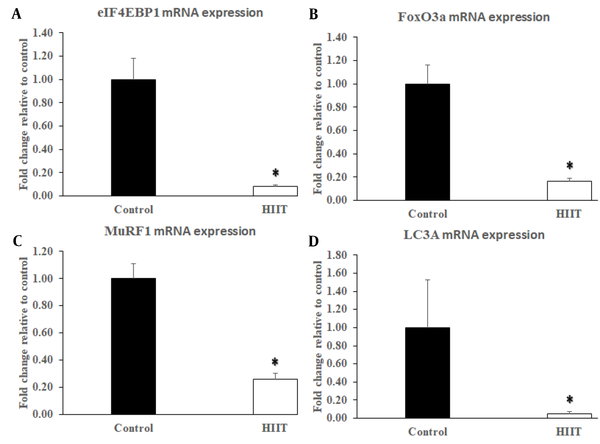
In an atrophy condition, muscle hypertrophy is prevented by binding 4EBP1 to eIF4E. But, the increased levels of Akt1 and mTORC1 in the HIIT group inhibited the eIF4EBP1 to form the 4EBP1-eIF4E complex. Accordingly, the expression of eIF4EBP1 was significantly decreased in the HIIT group (P < 0.001) compared with the control group (Figure 4A). On the other hand, the expression of FoxO3a, as a main regulator of atrophy, was significantly reduced (P < 0.001; F1,13 = 17.564, P = 0.001) in the exercise group compared with the control group (Figure 4B). Accordingly, the gene expression of MuRF1, as a downstream target of the FoxOs, was significantly reduced (P < 0.001; F1,13 = 9.077, P = 0.010) in the HIIT group (Figure 4C), and the expression of autophagy-related gene LC3A, which also is regulated by FoxO3a, was significantly suppressed in the HIIT group (Figure 4D) compared with the control group (P = 0.003; F1,13 = 25.081, P < 0.001).
Effect of eight-week HIIT on the protein degradation signaling components in rats’ soleus muscle. The exercise training protocol caused a significant reduction in eIF4EBP1 (A), FoxO3a (B), MuRF1 (C), and LC3A (D) mRNA expressions in the HIIT group (n = 8). Data are expressed as mean ± SEM. * Indicates significant difference (P ≤ 0.003) compared with the control group (n = 7).
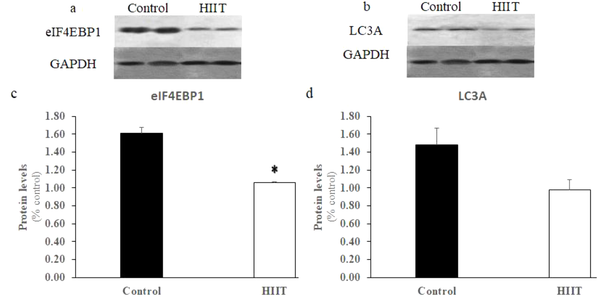
Western blot analysis revealed (Figure 5A), the expression level of eIF4EBP1 was significantly reduced (P = 0.012) in the HIIT group (Figure 5C). Although the protein level of LC3A (Figure 5B) was considerably lower in the HIIT group compared with the control group (Figure 5D), the between-group difference was not significant (P = 0.141). The results indicate that only the protein expression of eIF4EBP1 was significantly inhibited.
The protein expression of eIF4EBP1, LC3A, and GAPDH in rats’ soleus muscle (A and B). The protein levels of eIF4EBP1 were significantly reduced by the eight-week HIIT (C). In comparison to the significant reduction in mRNA expression, the expression of LC3A protein in the HIIT group (n = 8) was not significantly reduced compared with the control group (D). Data are expressed as mean ± SEM. * Indicates significant difference (P < 0.05) compared with the control group (n = 7).
5. Discussion
The present study simultaneously examines the alteration of gene expressions involved in protein synthesis and degradation signaling induced by HIIT.
The PI3K/Akt pathway plays an important role in the regulation of multiple cellular physiological and pathological processes, including proliferation, cell cycle, and growth. Indeed, PI3K is activated by extracellular stimuli such as growth factors and hormones. Serine/threonine kinase Akt then is phosphorylated and activated by PI3K to translocate from the cell membrane to phosphorylate multiple related downstream targets (15). It has been proven that Akt/mTOR pathway is downregulated under muscle atrophy conditions and upregulated during hypertrophy (16), so that 8-week moderate intensity of aerobic exercise training was sufficient to activate IGF-I/Akt/mTOR signaling pathway to prevent muscle wasting in heart failure mice (17). Indeed, Akt indirectly activates mTOR by phosphorylation and inhibition of tuberous sclerosis 2 (TSC2), which, in turn, inactivates the Rheb, as an mTOR inactivator (18). However, other distinct mechanisms, including ERK and leucine have been identified, which may cause a greater stimulation of mTOR (19). The significant increase in gene expression of Akt1 and mTORC1 induced by the HIIT protocol of the present study (Figure 2A and B) are consistent with the finding, which indicates the activation of mTOR is necessary and sufficient to induce skeletal muscle hypertrophy (20). Activated mTORC1, in response to stimuli such as nutrition, stress, and hormones, phosphorylates two downstream substrates S6K1 and eIF4EBP1. The inactive form of S6K1 is bound to the eIF3 complex. When mTORC1 is activated binds to eIF3-S6K1 and phosphorylates S6K1. Activated S6K1 is then released from eIF3-S6K1 and subsequently phosphorylates the translational targets to promote protein synthesis (21). Accordingly, the S6K1 gene expression in the present study, which was confirmed by Western blot analysis (Figure 3A and B), increased significantly in the HIIT group compared with the control group. The findings indicate that the exercise training protocol was capable to increase the expression of genes involved in Akt/mTORC1/S6K1 signaling, which are necessary for protein synthesis and skeletal muscle hypertrophy. Also, mTORC1 phosphorylates and inhibits eIF4EBP1 (4E-BP1) to binds to eukaryotic translation initiator (eIF4E), forming 4EBP1-eIF4E complex. Otherwise, dissociation from the 4EBP1-eIF4E complex, eIF4E binds to eIF4A and eIF4G, forming the eIF4F complex, which is a key step for translation initiation and protein synthesis in skeletal muscle (3). The reduction of eIF4EBP1 gene expression occurs; for example, under growth conditions, when the protein synthesis signaling becomes more active than degradation. Our findings revealed that the exercise training protocol was able to reduce the mRNA and protein expression levels of eIF4EBP1 (Figures 4A and 5A, respectively) compared with the control group (Figure 5C). On the other hand, FoxO3a, as a key regulator of muscle atrophy, modulates a variety of cellular processes, including autophagy and apoptosis, in response to growth factors, nutrition status, and stress signaling (22). In this regard, crosstalk between protein synthesis and protein degradation signaling components has been documented (23). Although the Akt-mTOR-S6K1 signaling pathway initiates the translation to promote muscle hypertrophy (24), Akt can also inhibit the nuclear transcriptional activity of the FoxO to attenuate atrophy and skeletal muscle loss. The role of FoxO3a on proteolysis depends on its cellular location. FoxO3a is inactive when is phosphorylated by Akt at three conserved residues, Thr32, Ser253, Ser315, thereby is excluded from the nucleus to the cytosol to be degraded (4, 22). Consistent with the non-exercise reports, the repressed FoxO3a gene expression in the HIIT group (Figure 4B) was associated with a significant increase of Akt1 gene expression, which may lead to muscle protein synthesis. Based on a suggestion, the skeletal muscle autophagy is not affected by inhibition of mTORC1, whereas inhibition of Akt leads to translocation of FoxO3 to the nucleus, and consequently, autophagy is induced. Although Akt, not mTORC1, has been known as a FoxO3 regulator (25), MST1 (Mammalian sterile 20‑like kinase 1) has also been identified as a positive activator for promoting the nuclear translocation and transcription activity of FoxO3a by phosphorylation and inhibition its association with 14-3-3 protein (26). FoxO3a, in turn, controls muscle protein degradation through two proteolytic systems, ubiquitin-proteasome, and autophagy-lysosome pathways. MuRF1, one of two muscle-specific ubiquitin ligases, has been known as a master gene in muscle wasting and a marker for muscle atrophy. However, an increase in gene expressions of the FoxO3a and MuRF1 in human skeletal muscle immediately after running exercise has been reported (27), the acute effect of exercise should be considered. In agreement with our findings (Figure 4C), it has been shown that both FoxO3a and MuRF1 gene expressions in soleus muscle were suppressed by a short-term (5 consecutive days) endurance treadmill running (14). Recently, four-week and eight-week HIIT did not alter the mRNA expression of AKT, mTOR, 4E-BP1, and MuRF1, possibly because of the exercise intensities and durations; however, a significant difference in the phosphorylated/dephosphorylated ratio in each gene was observed. The exercise protocol included 80% VO2max for 3-min as the high-intensity and 40% VO2max for 3-min as the low-intensity (8). In another study, the HIIT-induced MuRF1 gene expression was not significantly reduced in young male Wistar rats, whereas, in aged rats was suppressed. The exercise training protocol is not well described to interpret the unchanged MuRF1 mRNA but may be influenced by the age differences (7). In addition to MuRF1, FoxO3 also activates several autophagy genes, including LC3 to promote lysosomal protein degradation, which occurs during pathological conditions and causes muscle and myotubes atrophy (4, 5). In the present study, LC3A gene expression was suppressed in the HIIT group compared with the control group (Figure 4D), which did not confirm by Western blot analysis (Figure 5B); however, the protein expression of LC3A was considerably inhibited by the exercise training protocol (Figure 5D). This result may underscore the need for LC3A and autophagy to maintain cell survival (28). Apart from the fact that the repression of LC3A is partly due to the FoxO3a inhibition, MST1 has recently been identified as the LC3 regulator. Indeed, LC3 gene expression increases through activation of FoxO3a by the upregulation of the MST1. In other words, the deletion of MST1 suppresses the gene expression of the Foxo3a, which, in turn, results in a reduction of LC3 mRNA (29). We have recently shown that gene expression of the MST1 in cardiomyocytes, one of the Hippo signaling pathway components, is suppressed by an eight-week endurance treadmill running at 15° inclination (30). Accordingly, it can be interpreted that the significant reduction of FoxO3a and MuRF1 gene expressions in the present study may partly be affected by the HIIT-induced MST1 suppression. In addition to many factors, the alterations of two proteolytic systems components depend on nutrition status (Calorie restriction) and also exercise intensity and duration. It has been shown that nutrition deficiency is a potent stimulus for activation of MuRF1 (Ubiquitin-proteasome system) and LC3A (Autophagy-lysosome system), leading to muscle atrophy, and FoxO3 plays a role as a regulator. In contrast, feeding prevents the expression of both genes (31-33). Accordingly, given free access to standard food, it does not seem that the results of the present study be affected by the energy deficit. In addition, autophagy is influenced by exercise intensity and duration. Exercise intensity (e. g., high Vs. low) and duration (e. g., long Vs. short distance) are more effective to trigger autophagy mainly by activating FoxO3a and AMPK. Moreover, FoxO3a is regulated by many factors as described above. AMPK, as a major regulator of cellular energy homeostasis, mainly is activated by nutrient deprivation and strenuous exercise (34, 35).
5.1. Conclusions
The findings of the present study revealed that the intensity, percentage of VO2max, and duration of exertion of HIIT significantly impact the gene expression levels. The intensities and durations of the HIIT protocol were sufficient to increase the gene expression of protein synthesis signaling components and suppress the atrophy-related gene expressions, which can lead to attenuating muscle loss and increasing muscle mass. Accordingly, the HIIT protocol can be considered for some conditions such as sarcopenia and cachexia.
References
-
1.
Ali S, Garcia JM. Sarcopenia, cachexia and aging: diagnosis, mechanisms and therapeutic options - a mini-review. Gerontology. 2014;60(4):294-305. [PubMed ID: 24731978]. [PubMed Central ID: PMC4112511]. https://doi.org/10.1159/000356760.
-
2.
Yu JS, Cui W. Proliferation, survival and metabolism: the role of PI3K/AKT/mTOR signalling in pluripotency and cell fate determination. Development. 2016;143(17):3050-60. [PubMed ID: 27578176]. https://doi.org/10.1242/dev.137075.
-
3.
Proud CG. Signalling to translation: how signal transduction pathways control the protein synthetic machinery. Biochem J. 2007;403(2):217-34. [PubMed ID: 17376031]. https://doi.org/10.1042/BJ20070024.
-
4.
Mammucari C, Milan G, Romanello V, Masiero E, Rudolf R, Del Piccolo P, et al. FoxO3 controls autophagy in skeletal muscle in vivo. Cell Metab. 2007;6(6):458-71. [PubMed ID: 18054315]. https://doi.org/10.1016/j.cmet.2007.11.001.
-
5.
Zhao J, Brault JJ, Schild A, Cao P, Sandri M, Schiaffino S, et al. FoxO3 coordinately activates protein degradation by the autophagic/lysosomal and proteasomal pathways in atrophying muscle cells. Cell Metab. 2007;6(6):472-83. [PubMed ID: 18054316]. https://doi.org/10.1016/j.cmet.2007.11.004.
-
6.
Egan B, Zierath JR. Exercise metabolism and the molecular regulation of skeletal muscle adaptation. Cell Metab. 2013;17(2):162-84. [PubMed ID: 23395166]. https://doi.org/10.1016/j.cmet.2012.12.012.
-
7.
Tayebi SM, Siahkouhian M, Keshavarz M, Mahdian R, Shamsi MM, Shahbazi S. The Effects of High Intensity Interval Training on Mir-23a Expression and Related Factors Involved in Muscular Atrophy of Aged Rats. Int J Appl Exerc Physiol. 2019;8(1):170-6.
-
8.
Cui X, Zhang Y, Wang Z, Yu J, Kong Z, Ruzic L. High-intensity interval training changes the expression of muscle RING-finger protein-1 and muscle atrophy F-box proteins and proteins involved in the mechanistic target of rapamycin pathway and autophagy in rat skeletal muscle. Exp Physiol. 2019;104(10):1505-17. [PubMed ID: 31357248]. https://doi.org/10.1113/EP087601.
-
9.
Blue MNM, Smith-Ryan AE, Trexler ET, Hirsch KR. The effects of high intensity interval training on muscle size and quality in overweight and obese adults. J Sci Med Sport. 2018;21(2):207-12. [PubMed ID: 28647284]. [PubMed Central ID: PMC7104622]. https://doi.org/10.1016/j.jsams.2017.06.001.
-
10.
Hoydal MA, Wisloff U, Kemi OJ, Ellingsen O. Running speed and maximal oxygen uptake in rats and mice: practical implications for exercise training. Eur J Cardiovasc Prev Rehabil. 2007;14(6):753-60. [PubMed ID: 18043295]. https://doi.org/10.1097/HJR.0b013e3281eacef1.
-
11.
Wisloff U, Helgerud J, Kemi OJ, Ellingsen O. Intensity-controlled treadmill running in rats: VO(2 max) and cardiac hypertrophy. Am J Physiol Heart Circ Physiol. 2001;280(3):H1301-10. [PubMed ID: 11179077]. https://doi.org/10.1152/ajpheart.2001.280.3.H1301.
-
12.
Kemi OJ, Haram PM, Loennechen JP, Osnes JB, Skomedal T, Wisloff U, et al. Moderate vs. high exercise intensity: differential effects on aerobic fitness, cardiomyocyte contractility, and endothelial function. Cardiovasc Res. 2005;67(1):161-72. [PubMed ID: 15949480]. https://doi.org/10.1016/j.cardiores.2005.03.010.
-
13.
Abdolmaleki A, Zahri S, Bayrami A. Rosuvastatin enhanced functional recovery after sciatic nerve injury in the rat. Eur J Pharmacol. 2020;882:173260. [PubMed ID: 32534070]. https://doi.org/10.1016/j.ejphar.2020.173260.
-
14.
Kavazis AN, Smuder AJ, Powers SK. Effects of short-term endurance exercise training on acute doxorubicin-induced FoxO transcription in cardiac and skeletal muscle. J Appl Physiol (1985). 2014;117(3):223-30. [PubMed ID: 24947024]. [PubMed Central ID: PMC4347740]. https://doi.org/10.1152/japplphysiol.00210.2014.
-
15.
Shi X, Wang J, Lei Y, Cong C, Tan D, Zhou X. Research progress on the PI3K/AKT signaling pathway in gynecological cancer (Review). Mol Med Rep. 2019. https://doi.org/10.3892/mmr.2019.10121.
-
16.
Bodine SC, Stitt TN, Gonzalez M, Kline WO, Stover GL, Bauerlein R, et al. Akt/mTOR pathway is a crucial regulator of skeletal muscle hypertrophy and can prevent muscle atrophy in vivo. Nat Cell Biol. 2001;3(11):1014-9. [PubMed ID: 11715023]. https://doi.org/10.1038/ncb1101-1014.
-
17.
Bacurau AV, Jannig PR, de Moraes WM, Cunha TF, Medeiros A, Barberi L, et al. Akt/mTOR pathway contributes to skeletal muscle anti-atrophic effect of aerobic exercise training in heart failure mice. Int J Cardiol. 2016;214:137-47. [PubMed ID: 27060274]. https://doi.org/10.1016/j.ijcard.2016.03.071.
-
18.
Wan M, Wu X, Guan KL, Han M, Zhuang Y, Xu T. Muscle atrophy in transgenic mice expressing a human TSC1 transgene. FEBS Lett. 2006;580(24):5621-7. [PubMed ID: 16996505]. https://doi.org/10.1016/j.febslet.2006.09.008.
-
19.
Winter JN, Jefferson LS, Kimball SR. ERK and Akt signaling pathways function through parallel mechanisms to promote mTORC1 signaling. Am J Physiol Cell Physiol. 2011;300(5):C1172-80. [PubMed ID: 21289294]. [PubMed Central ID: PMC3093949]. https://doi.org/10.1152/ajpcell.00504.2010.
-
20.
Nader GA, McLoughlin TJ, Esser KA. mTOR function in skeletal muscle hypertrophy: increased ribosomal RNA via cell cycle regulators. Am J Physiol Cell Physiol. 2005;289(6):C1457-65. [PubMed ID: 16079186]. https://doi.org/10.1152/ajpcell.00165.2005.
-
21.
Ma XM, Blenis J. Molecular mechanisms of mTOR-mediated translational control. Nat Rev Mol Cell Biol. 2009;10(5):307-18. [PubMed ID: 19339977]. https://doi.org/10.1038/nrm2672.
-
22.
Fasano C, Disciglio V, Bertora S, Lepore Signorile M, Simone C. FOXO3a from the Nucleus to the Mitochondria: A Round Trip in Cellular Stress Response. Cells. 2019;8(9). [PubMed ID: 31546924]. [PubMed Central ID: PMC6769815]. https://doi.org/10.3390/cells8091110.
-
23.
Sandri M. Signaling in muscle atrophy and hypertrophy. Physiology (Bethesda). 2008;23:160-70. [PubMed ID: 18556469]. https://doi.org/10.1152/physiol.00041.2007.
-
24.
Laplante M, Sabatini DM. mTOR signaling at a glance. J Cell Sci. 2009;122(Pt 20):3589-94. [PubMed ID: 19812304]. [PubMed Central ID: PMC2758797]. https://doi.org/10.1242/jcs.051011.
-
25.
Mammucari C, Schiaffino S, Sandri M. Downstream of Akt: FoxO3 and mTOR in the regulation of autophagy in skeletal muscle. Autophagy. 2008;4(4):524-6. [PubMed ID: 18367868]. https://doi.org/10.4161/auto.5905.
-
26.
Lehtinen MK, Yuan Z, Boag PR, Yang Y, Villen J, Becker EB, et al. A conserved MST-FOXO signaling pathway mediates oxidative-stress responses and extends life span. Cell. 2006;125(5):987-1001. [PubMed ID: 16751106]. https://doi.org/10.1016/j.cell.2006.03.046.
-
27.
Louis E, Raue U, Yang Y, Jemiolo B, Trappe S. Time course of proteolytic, cytokine, and myostatin gene expression after acute exercise in human skeletal muscle. J Appl Physiol (1985). 2007;103(5):1744-51. [PubMed ID: 17823296]. https://doi.org/10.1152/japplphysiol.00679.2007.
-
28.
Botti-Millet J, Nascimbeni AC, Dupont N, Morel E, Codogno P. Fine-tuning autophagy: from transcriptional to posttranslational regulation. Am J Physiol Cell Physiol. 2016;311(3):C351-62. [PubMed ID: 27335173]. https://doi.org/10.1152/ajpcell.00129.2016.
-
29.
Wei B, Dui W, Liu D, Xing Y, Yuan Z, Ji G. MST1, a key player, in enhancing fast skeletal muscle atrophy. BMC Biol. 2013;11:12. [PubMed ID: 23374633]. [PubMed Central ID: PMC3606410]. https://doi.org/10.1186/1741-7007-11-12.
-
30.
Gholipour M, Tabrizi A. The role of Hippo signaling pathway in physiological cardiac hypertrophy. Bioimpacts. 2020;10(4):251-7. [PubMed ID: 32983941]. [PubMed Central ID: PMC7502907]. https://doi.org/10.34172/bi.2020.32.
-
31.
Lysenko EA, Vepkhvadze TF, Lednev EM, Vinogradova OL, Popov DV. Branched-chain amino acids administration suppresses endurance exercise-related activation of ubiquitin proteasome signaling in trained human skeletal muscle. J Physiol Sci. 2018;68(1):43-53. [PubMed ID: 27913948]. https://doi.org/10.1007/s12576-016-0506-8.
-
32.
Nonaka Y, Urashima S, Inai M, Nishimura S, Higashida K, Terada S. Effects of rapid or slow body weight reduction on intramuscular protein degradation pathways during equivalent weight loss on rats. Physiol Res. 2017;66(5):823-31. [PubMed ID: 28730836]. https://doi.org/10.33549/physiolres.933502.
-
33.
Milan G, Romanello V, Pescatore F, Armani A, Paik JH, Frasson L, et al. Regulation of autophagy and the ubiquitin-proteasome system by the FoxO transcriptional network during muscle atrophy. Nat Commun. 2015;6:6670. [PubMed ID: 25858807]. [PubMed Central ID: PMC4403316]. https://doi.org/10.1038/ncomms7670.
-
34.
Schwalm C, Jamart C, Benoit N, Naslain D, Premont C, Prevet J, et al. Activation of autophagy in human skeletal muscle is dependent on exercise intensity and AMPK activation. FASEB J. 2015;29(8):3515-26. [PubMed ID: 25957282]. https://doi.org/10.1096/fj.14-267187.
-
35.
Jamart C, Francaux M, Millet GY, Deldicque L, Frere D, Feasson L. Modulation of autophagy and ubiquitin-proteasome pathways during ultra-endurance running. J Appl Physiol (1985). 2012;112(9):1529-37. [PubMed ID: 22345427]. https://doi.org/10.1152/japplphysiol.00952.2011.