Abstract
Background:
Cancerous cells proliferate as fast as possible without a proper surveillance system. This rapid cell division leads to enormous mutation rates, which help a tumor establish.Objectives:
This study evaluated the potential of inducing apoptosis using Noxa and Puma in a hepatocarcinoma cell line.Methods:
The current study generated two recombinant lentiviruses, pLEX-GCN and pLEX-GCP, bearing Noxa and Puma, respectively. Transduction of both genes to hepatocarcinoma (HepG2) was verified using fluorescent microscopic analysis, western blotting, and quantitative real-time polymerase chain reaction (PCR). To evaluate the potential of Noxa and Puma to initiate apoptosis, a caspase-9 real-time, MTT assay, and a 4’, 6-diamidino-2-phenylindole (DAPI) reagent were performed to stain apoptotic cells.Results:
The data verified successful transduction to HepG2 and HEK293T. Higher relative expression of Noxa and Puma rather than the untransduced cell line showed these genes are expressed more in HepG2 in comparison to HEK293T. The results of the real-time PCR, MTT assay, and DAPI reagent illustrated that higher cells initiated apoptosis following Puma transduction rather than Noxa.Conclusions:
In this approach, the suicide gene was transferred to transformed cells and ignited apoptosis to exterminate them. Puma is a more potent killer gene and has higher capabilities to start intrinsic apoptosis pathway.Keywords
1. Background
The link between apoptosis and cellular-surveillance mechanisms, that is, checkpoints, is thought to provide a critical defense against the development of cancer. Flaws in apoptotic pathways, with a failure to abolish cells harboring DNA damage, viral infection, or aberrations in the mitotic apparatus, have accompanied the inception and progression of cancer, and mutations in proapoptotic genes may present augmented tumor invasiveness and resistance to therapy.
Apoptosis regulators consist of three members: anti-apoptosis (BH1-4), pro-apoptosis (BH1 3) and BH3-only proteins. BH3-only proteins drive cells toward apoptosis by limiting anti-apoptotic (e.g., BCL-2, BCL-xL, and/or MCL-1) and activating pro-apoptosis proteins (i.e., BAK and/or BAX). Among BH3-only members, Bim and Bid inhibit anti-apoptotic proteins and Puma, Bad, and Noxa activate pro-apoptotic proteins, which lead cells toward apoptosis. Activated pro-apoptotic BH3-only proteins lead to apoptosome assembly by means of facilitating cytochrome-C release to cytoplasm from the intermembrane space of the mitochondria (internal apoptosis pathway) (1, 2).
The p53 tumor-suppressor gene is disabled in approximately 50% of human tumors. This tumor suppressor is phosphorylated following reversible cellular damage and causes cell-cycle arrests by activating ataxia telangiectasia-mutated (ATM), ataxia telangiectasia, and Rad3-related (ATR) (3, 4). Apoptosis is induced by BH3-only proteins of the Bcl-2 family, of which Noxa and Puma have been well characterized. These two proteins function in cooperation as downstream factors to promote a p53-mediated apoptotic response. Following irreparable damage, the activated p53 subsequently activates Puma and Noxa, which lead the damaged cell to apoptosis to prevent transformation (5-7).
2. Objectives
This study evaluates the best killer gene for the hepatocarcinoma killer gene. There are two widely used killer genes in cancer gene therapy: Noxa and Puma. Both genes are activated following irreparable genetic damage, which is the hallmark of rapidly dividing cancerous cells. To reduce side effects of these killer genes in the normal cell line, both suicidal genes were cloned under control of the cancer-specific promoter CXCR1. Enhanced green fluorescent protein (eGFP) made it possible to monitor and also verify all the steps.
3. Methods
3.1. Cell Culture
Human embryonic kidney (HEK) 293T cells (ATCC CRL-1573) were grown in Dulbecco’s Modified Eagle’s Medium (DMEM, Sigma-Aldrich), supplemented with 10% FBS, 100 units/mL penicillin, and 100 mg/ml streptomycin incubated in 5% CO2 at 37°C.
Hepatocarcinoma (HepG2), the hepatocellular (ATCC HB-8065) cancerous cell line, was grown in DMEM-f12 15% FBS, supplemented with pen/strept and insulin.
3.2. Constructing Recombinant pLEX MCS
3.2.1. Cloning of CXCR1 Promoter into pLEX MSC
The CXCR1 sequence was evaluated in Genbank (Gene ID: 3577). This gene is allocated in chromosome 2, region 3. To detect this gene’s promoter and regulatory sequences, the CXCR1 gene and 5000 bases upstream of CXCR1 were analyzed by P-MATCH software, the Trasnfac database, and the NNPP database. After thorough investigation, 490 base pairs, -320 to -810, were considered as the promoter region of CXCR1. To amplify the CXCR1 promoter, genomic DNA was extracted and polymerase chain reaction (PCR) amplified the region -1000 to + 100. The amplified CXCR1 promoter fragment was double digested by HindIII and BamHI and then ligated into the same site in pUC19.
3.2.2. Cloning of Puma and Noxa Downstream of CXCR1 Promoter
The coding sequence of Noxa (Genbank accession number NM_021127) and Puma (Genbank accession number NM_001127240) were PCR amplified using extracted mRNA and synthesized cDNA. Each suicide gene was double digested by SmaI and KpnI and then ligated downstream of the CXCR1 promoter in pUC19.
After construction of the suicide genes under regulation of the cancer-specific CXCR1 promoter, these fragments, CXCR1-Puma and CXCR1-Noxa, were double digested and were subcloned into the NotI-XhoI site of pLEX MCS, pL-CX-Puma, and pL-CX-Noxa, respectively. Recombinant pLEX, pL-CX-Puma, and pL-CX-Noxa were confirmed by sequencing (Pishgam company, Tehran, Iran).
Primers Sequence for Noxa and Puma Cloning
| Name | Primer sequence | Restriction site | Tag |
|---|---|---|---|
| Puma-For | gctCCCGGGgaggcgattgcgattgggtg | SmaI | |
| Puma-Rev | GCTGGTACCccactgttccaatctgattttattgaaaagCATCATCACCATCACCAC | KpnI | Hist-tag |
| Noxa-For | GCTCCCGGGactggacaaaagcgtggtctctg | SmaI | |
| Noxa-Rev | GCTGGTACCaacagtgaaaacttttaatagtttactgccCATCATCACCATCACCAC | KpnI | Hist-tag |
3.3. Transduction Evaluation
3.3.1. Microscopic Analysis of GFP Expression
In this study, eGFP played a role as a positive control. eGFP expression means successful viral production and transduction. Therefore, to analyze this reporter gene expression, 1 x 105 of the transduced cells were seeded in a 24-well flask. After 12 hours, 24 hours, 48 hours, and 72 hours, the cells were detached from the flask using 0.25% trypsin-EDTA and analyzed by fluorescent microscopy. Fluorescent microscopy used 395 nm to excite eGFP, and its emission peak was at 595 nm.
3.3.2. Western Blotting
HepG2 and HEK293 cells transduced by pLEX-Puma and pLEX-Noxa were harvested and lysed in lysis buffer (50 mM Tris-HCl (pH 7.5), 150 mM NaCl, 1% Nonidet P-40, and 5 mM EDTA). Total protein was resolved on 4% - 12% SDS-polyacrylamide gels and transferred to nitrocellulose membranes at 300 mA for 3 hours on ice. The membranes were blocked and then incubated with anti-his–tag antibody (Abcam, Cambridge, United Kingdom) overnight at 4°C. After washing, the membranes were incubated with horseradish peroxidase-conjugated secondary antibody for 1 h at room temperature and then washed again. ECL reagent (Thermo Scientific, Massachusetts, United States) uncovered recombinant his-tagged Noxa and Puma.
3.3.3. Puma and Noxa Real-Time
Quantitative real-time PCR was performed on extracted RNA of respective transduced cell lines to corroborate the expression of the Puma and Noxa gene. cDNA was synthesized from isolated RNA using RNeasy Mini Kit (Qiagen, Hilden, Germany), and then real-time PCR was carried out on both genes for 24 hours and 48 hours transduction. The relative expression ratio of the Noxa and Puma gene was calculated according to the equation of Pfaffl using untreated cells as reference:
Puma-F, 5’ACGACCTCAACGCACAGTACGA (8),
Puma-R 5’GTAAGGGCAGGAGTCCCATGATGA (8),
Noxa-F, 5’-GCTGGAAGTCGAGTGTGCTA-3’ (9), and
Noxa-R, 5’-CCTGAGCAGAAGAGTTTGGA-3’ (9).
3.4. Quantifying Apoptosis Response
3.4.1. Caspase-9 Real-Time
Total RNA was isolated from HepG2 and HEK293 cells using RNeasy (Qiagen, Hilden, Germany). mRNA was reverse transcribed (cDNA synthesized), and quantitative PCR was performed using a TaqMan Gold RT-PCR kit (Applied Biosystems, California, United States) on an ABI Prism 7700 thermal cycler (Applied Biosystems, California, United States). The primers were as follows: Casp-9, 5’-ATGACCACCACAAAGCAGTCC-3’ (forward primer), 5’-CGTGACCATTTTCTTGGCAG-3’ (reverse primer), and glyceraldehyde-3-phosphate dehydrogenase, 5’-ATGTGTCCGTCGTGGATCTGAC-3’ (forward primer), 5’-TCAAGAAGGTGGTGAAGCAGG-3’ (reverse primer).
3.4.2. Cell Viability
The MTT assay was performed as explained by Deniziot and Lang (10). Briefly, transduced cancerous and normal cell lines using pLEX-GCN and pLEX-GCP with MOI of 100 were cultured for 48 hours at 37°C, 5% CO2 in complete media. Then the culture media was changed with free serum culture media. The MTT (Invitrogen, California, United States), dissolved in phosphate buffer saline (PBS) (500 µg/mL MTT), was added to each well. The MTT in the free serum media was incubated for 3 hours. After this interval, the free serum culture media, including the MTT reagent, was removed, and DMSO was added to each well to dissolve the formazan crystals. The optical densities were measured at 490 nm spectral wavelength using a microtiter plate reader. The MTT assays were performed in triplicate.
3.4.3. 4’, 6-Diamidino-2-Phenylindole (DAPI) Staining
To evaluate the apoptotic cells, 4’, 6-diamidino-2-phenylindole (DAPI) reagent-stained cells underwent programmed cell death. Transduced cells were centrifuged at 15,000 g for 5 minutes. Then, harvested cells were resuspended in the fixative solution (0.7% formaldehyde, 0.5% Nonidet P-40, and 10 µg/mL DAPI in PBS). Apoptosis was assessed by methods of microscopic visualization of condensed chromatin and micronucleation.
4. Results
4.1. Construction of Suicide pLEX Vector
The suicide construct, the suicide gene under the cancer-specific promoter, was constructed in two steps. First, the CXCR1 promoter region was amplified from the isolated genomic DNA. This cancer-specific promoter was subcloned into puC19. An 1100 bp band in the colony PCR and double digestion revealed successful subcloning.
Second, Noxa and Puma sequences were ligated downstream of the CXCR1 promoter to generate two distinctive killer genes: CXCR1-Noxa and CXCR1-Puma, respectively. Noxa and Puma cDNA, 1954 bp and 1839 bp, respectively, were PCR amplified. The Noxa and Puma sequences were cloned downstream of the CXCR1 promoter in pUC19. The colony PCR and double digestion revealed appropriate bands respective to each gene. Final verification was the sequencing result, which showed no unintentional mutation during the cloning process. Finally, CXCR1-Noxa and CXCR1-Puma were subcloned into pLEX-G to generate pLEX-GCN and pLEX-GCP, respectively.
4.2. Transduction of HepG2 by Recombinant Lentivirus
To certify that each killer gene delivered by a lentiviral vector was expressed in the target cells, fluorescent microscopy analysis was performed to reveal the eGFP expression, and western blotting was performed using the anti-his-tag antibody (Abcam, Cambridge, United Kingdom).
4.2.1. Fluorescent Microscopy Revealed eGFP Expression
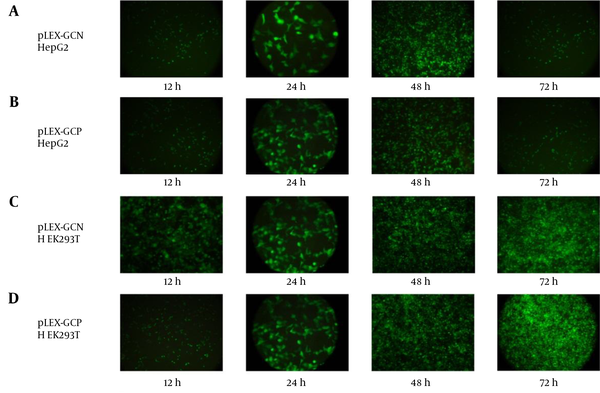
In the hepatocarcinoma cell line HepG2, more cells showed eGFP expression from 12 hours to 48 hours, but at 72 hours, significant eGFP signals were lost. This sudden drop of fluorescent signal was because of expression of Noxa and Puma under the cancer-specific CXCR1 promoter and induction of the apoptosis pathway in this cell line (Figure 1).
On the other hand, eGFP signals increased in the normal cell line HEK293T as time passed from 12 hours to 72 hours. Because the cancer-specific promoter is inactive in normal cells, no Noxa and Puma were expressed, apoptosis was not induced, and more cells showed eGFP expression.
Microscopic Analysis of eGFP Expression in the Target Cells
4.2.2. Puma and Noxa Western Blotting
In order to verify Noxa and Puma expression in the hepatocarcinoma cell line HepG2, Noxa and Puma underwent immune detection. The presence of the C-terminal his-tag provided a means to detect transduced killer genes without interference of cellular ones.
After staining nitrocellulose with the primary anti-his–tag antibody and secondary antibody, ECL reagent detection unveiled the 6 kDa and 26.5 kDa band, which were proof of transduced Noxa and Puma expression, respectively (Figure 2).
Noxa and Puma Western Blotting
4.2.3. Puma and Noxa Real-Time PCR
To ensure successful transduction, real-time PCR was performed on isolated RNA from both transduced HepG2 and HEK293T cell lines. As can be concluded from Figure 3, Noxa is between 44% and 57% relative expression in HepG2 at 24 hours and 48 hours, respectively, even though in HEK293T at both 24 hours and 48 hours it is between 15% and 20%. In the case of Puma, relative expression in HepG2 at 24 hours and 48 hours is 52% and 77%, respectively. Puma basal expression in HEK293T is similar to Noxa.
Real-time PCR quantitatively illustrates Noxa and Puma expression.
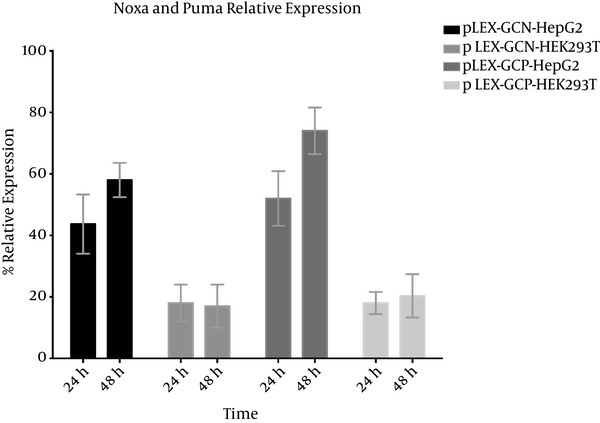
4.3. Apoptosis Induction via the Suicide Gene
In order to evaluate the apoptosis response ensuing pLEX-GCN and pLEX-GCP, three tests were performed. Caspase-9 (the main Noxa and Puma effector and apoptosis initiator) real-time revealed relative expression of Noxa and Puma in comparison to untreated cells. MTT assay evaluated cell viability and DAPI reagent stained apoptotic cell.
4.3.1. Caspase-9 Up-Regulated Following Transduction
In the normal cell line HEK293T, caspase-9 expression following pLEX-GCN transduction was almost equal, 20% compared to 25%. Similar results were obtained after pLEX-GCP transduction on the same cell line, HEK293T. Relative expression was between 18 at 72 hours and 23 at 48 hours after transduction.
On the other hand, caspase-9 relative expression percentages were higher in the hepatocarcinoma cell line HepG2. In this cell line, the CXCR1 promoter is highly active due to its feature. Caspase-9 relative expressions following pLEX-GCN were 57.66%, 61.66%, and 33.33% for 24 hours, 48 hours, and 72 hours, respectively. These percentages were meaningfully higher for pLEX-GCP suicide construct, 80.33%, 91.33% and 42% for 24 hours, 48 hours, and 72 hours, respectively.
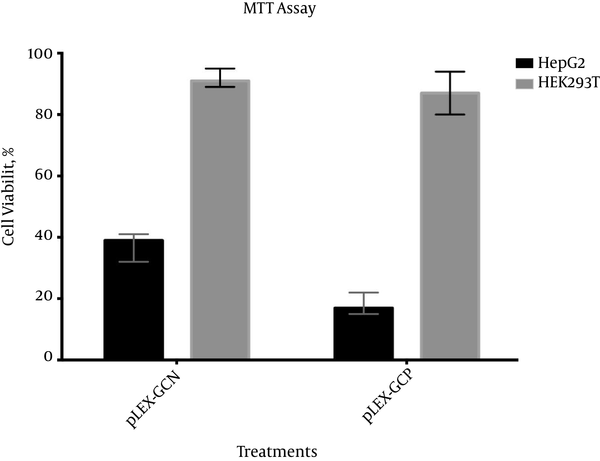
4.3.2. MTT Assay Showed Selective Apoptosis Response in Cancerous Cell Line
MTT assays were carried out four days after transduction. As illustrated in Figure 5, both viral particles were able to initiate the apoptosis response selectively in HepG2. Cell viability of pLEX-GCN in HepG2 was 37.33%, while in the normal cell line where CXCR1 is inactive, cell viability was recorded at 91.66%. In the case of pLEX-GCP suicide construct, 19.33% cell viability for HepG2, but 87% cell viability for HEK293T, was recorded.
Caspase-9 Real-Time PCR Following Noxa and Puma Transduction
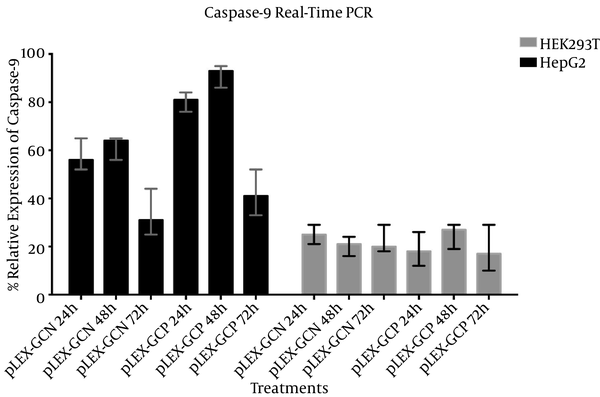
4.3.3. Apoptotic Cell Percentage Determination Using DAPI Staining
Figure 6 shows that HepG2 had a higher percentage of apoptosis compared to HEK293T, while both cell lines were transduced with equal MOI of pLEX-GCN and pLEX-GCP, MOI 100. The apoptotic percentages emanating from pLEX-GCN were 69% and 21% at 72 hours post-transduction for HepG2 and HEK293T, respectively. For the second construct, pLEX-GCP, the apoptotic percentages recorded were 86% and 27% at the same post-induction time for HepG2 and HEK293T, respectively.
MTT Assay Evaluated Cell Viability Following Viral Transduction of pLEX-GCN and pLEX-GCP
DAPI staining

5. Discussion
Hepatocarcinoma is the third cause of death from cancer in the world. One of the top reasons for cancer formation is a malfunctioning apoptosis apparatus. A disabled apoptosis pathway is not able to exterminate a mutated (transformed) cell, which eventually leads to uncontrollable proliferation. An earlier study by the authors (data is being published) showed that Puma protein under an hTERT cancer-specific promoter is highly toxic and induces apoptosis in the breast cancer cell line BT-474. A Zhang et al. review discusses the role of Noxa and Puma in tumor formation. Deletion of both Noxa and Puma are risk factors in tumorigenesis and development (11). In 2012, Tromp et al. (12), on the other hand, concluded that low levels of expression of apoptosis inducers and/or anti-apoptosis Bcl2 proteins, such as Mcl-1, are major factors in chemotherapy resistance. They hypothesized that the approach to overcome chemotherapy resistance is to change the balance of the apoptosis pathway in favor of Noxa and Puma, either by decreasing the Mcl-1 level or increasing the Noxa and Puma level, which will lead to chemotherapy’s synergistic effects against CLL cells in chemoresistant niches (13).
In a concurrent study, the authors of the current study evaluated and compared the top mostly used suicidal gene in hepatocarcinoma gene therapy. Although the earlier study uncovered successful apoptosis induction by Puma in BT-474, the need experimentally proved the best one in hepatocarcinoma therapy.
As the results of the current study show, both Noxa and Puma are expressed solely in HepG2 under control of the cancer-specific promoter (CXCR1). Recombinant proapoptotic lentiviruses were transduced to HepG2 and HEK293T. The fluorescent microscopy data attest to successful transduction and gene expression. The interesting finding was reduced fluorescent intensity in HepG2 at 72 hours, while HEK293T showed no loss of eGFP at all. It was assumed that expression of recombinant Noxa and Puma were driven cells toward apoptosis and that eGFP was destroyed following caspase-9 activation. Even though eGFP expression was irrefutable proof for successful viral transduction, western blotting and quantitative PCR also evaluated recombinant Noxa and Puma expressions. Because Noxa and Puma are expressed at low levels in all cells, his-tag was introduced at C-terminal in both genes to make it possible to distinguish between transduced genes and intrinsic ones. Western blotting results and real-time PCR showed that these two killer genes are overexpressed in HepG2, while just a basal expression were detected in the normal cell line control, HEK293T.
After confirmation of Noxa and Puma overexpression in HepG2, it was time to analyze how much these two genes were able to instigate apoptosis. In the intrinsic apoptosis pathway, in response to irreparable DNA damage, p53 is activated, which triggers Noxa and/or Puma activation. Activated Noxa and/or Puma in the next step results in caspase-9 expression and activation. Therefore, amounts of caspase-9 expression would be a good indicator of the potential of each killer gene. In the first phase, caspase-9 real-time PCR could quantitatively show the potential of each killer gene. The real-time PCR of caspase-9 provided evidence that this gene was overexpressed at least two-fold more than untreated cells. As the data showed, caspase-9 expression was meaningfully higher following pLEX-GCP transduction rather than pLEX-GCN. As the first experiment on the intensity of each gene to increase caspase-9 expression provided, Puma was the winner. Caspase-9 expression does not necessarily mean apoptosis induction. This gene is expressed routinely in many cells and needs to be activated prior to apoptosis. Therefore, an MTT assay was performed to evaluate cell viability following Noxa or Puma transduction. As MTT assay results revealed, cell viability in pLEX-GCN-transduced HepG2 was significantly higher than pLEX-GCP, which means more cells underwent apoptosis after pLEX-GCP transduction rather than pLEX-GCN. This outcome was in accordance with the caspase-9 real-time PCR, in which the caspase-9 expression was higher following pLEX-GCP transduction. Therefore, to verify the MTT assay data, DAPI staining needed to be performed. The DAPI reagent is able to stain the apoptotic cell. In the time course of 12 hours and 24 hours, more cells were stained following Noxa transduction, but in the next two, 48 hours and 72 hours, Puma-treated cells experienced more apoptosis than Noxa. In line with earlier results, Puma was more potent for activating caspase-9 and also driving cells toward apoptosis in comparison to Noxa.
This study reveals that augments in the Noxa or Puma proapoptotic protein level leads to an increased cytochrome-C release from the intermembrane space of the mitochondria into the cytoplasm, and finally, caspase-9 activity is induced in HepG2 cells. These results are in accordance with results that melatonin can induce up-regulation of cytochrome-C release and that caspase-9 up-regulation can head cells toward apoptosis in rat prolactinomas, human B-lymphoma cells, or human myeloid cells (14-16).
Besides, this study was able to shed light on the best and the most potent killer genes in hetapocarcinoma gene therapy. Concurrent results illustrate that, although Noxa and Puma are initiators of a similar pathway in apoptosis after irreparable genomic mutation, Puma is more a suitable candidate in hetapocarcinoma gene therapy. A higher apoptosis response is induced following Puma in comparison with its partner, Noxa.
Acknowledgements
References
-
1.
Tang D, Lotze MT, Kang R, Zeh HJ. Apoptosis promotes early tumorigenesis. Oncogene. 2011;30(16):1851-4. [PubMed ID: 21151175]. https://doi.org/10.1038/onc.2010.573.
-
2.
Chipuk JE, Green DR. PUMA cooperates with direct activator proteins to promote mitochondrial outer membrane permeabilization and apoptosis. Cell Cycle. 2009;8(17):2692-6. [PubMed ID: 19652530]. https://doi.org/10.4161/cc.8.17.9412.
-
3.
Dohi T, Beltrami E, Wall NR, Plescia J, Altieri DC. Mitochondrial survivin inhibits apoptosis and promotes tumorigenesis. J Clin Invest. 2004;114(8):1117-27. [PubMed ID: 15489959]. https://doi.org/10.1172/JCI22222.
-
4.
Pusapati RV, Rounbehler RJ, Hong S, Powers JT, Yan M, Kiguchi K, et al. ATM promotes apoptosis and suppresses tumorigenesis in response to Myc. Proc Natl Acad Sci U S A. 2006;103(5):1446-51. [PubMed ID: 16432227]. https://doi.org/10.1073/pnas.0507367103.
-
5.
Yu J, Yue W, Wu B, Zhang L. PUMA sensitizes lung cancer cells to chemotherapeutic agents and irradiation. Clin Cancer Res. 2006;12(9):2928-36. [PubMed ID: 16675590]. https://doi.org/10.1158/1078-0432.CCR-05-2429.
-
6.
Leibowitz BJ, Qiu W, Liu H, Cheng T, Zhang L, Yu J. Uncoupling p53 functions in radiation-induced intestinal damage via PUMA and p21. Mol Cancer Res. 2011;9(5):616-25. [PubMed ID: 21450905]. https://doi.org/10.1158/1541-7786.MCR-11-0052.
-
7.
Wang H, Qian H, Yu J, Zhang X, Zhang L, Fu M, et al. Administration of PUMA adenovirus increases the sensitivity of esophageal cancer cells to anticancer drugs. Cancer Biol Ther. 2006;5(4):380-5. [PubMed ID: 16481741].
-
8.
Garrison SP, Jeffers JR, Yang C, Nilsson JA, Hall MA, Rehg JE, et al. Selection against PUMA gene expression in Myc-driven B-cell lymphomagenesis. Mol Cell Biol. 2008;28(17):5391-402. [PubMed ID: 18573879]. https://doi.org/10.1128/MCB.00907-07.
-
9.
Gomez-Bougie P, Wuilleme-Toumi S, Menoret E, Trichet V, Robillard N, Philippe M, et al. Noxa up-regulation and Mcl-1 cleavage are associated to apoptosis induction by bortezomib in multiple myeloma. Cancer Res. 2007;67(11):5418-24. [PubMed ID: 17545623]. https://doi.org/10.1158/0008-5472.CAN-06-4322.
-
10.
Denizot F, Lang R. Rapid colorimetric assay for cell growth and survival. Modifications to the tetrazolium dye procedure giving improved sensitivity and reliability. J Immunol Methods. 1986;89(2):271-7. [PubMed ID: 3486233].
-
11.
Zhang LN, Li JY, Xu W. A review of the role of Puma, Noxa and Bim in the tumorigenesis, therapy and drug resistance of chronic lymphocytic leukemia. Cancer Gene Ther. 2013;20(1):1-7. [PubMed ID: 23175245]. https://doi.org/10.1038/cgt.2012.84.
-
12.
Tromp JM, Geest CR, Breij EC, Elias JA, van Laar J, Luijks DM, et al. Tipping the Noxa/Mcl-1 balance overcomes ABT-737 resistance in chronic lymphocytic leukemia. Clin Cancer Res. 2012;18(2):487-98. [PubMed ID: 22128299]. https://doi.org/10.1158/1078-0432.CCR-11-1440.
-
13.
Billard C. Development of Noxa-like BH3 mimetics for apoptosis-based therapeutic strategy in chronic lymphocytic leukemia. Mol Cancer Res. 2012;10(6):673-6. [PubMed ID: 22466256]. https://doi.org/10.1158/1541-7786.MCR-12-0015.
-
14.
Trubiani O, Recchioni R, Moroni F, Pizzicannella J, Caputi S, Di Primio R. Melatonin provokes cell death in human B-lymphoma cells by mitochondrial-dependent apoptotic pathway activation. J Pineal Res. 2005;39(4):425-31. [PubMed ID: 16207299]. https://doi.org/10.1111/j.1600-079X.2005.00270.x.
-
15.
Yang QH, Xu JN, Xu RK, Pang SF. Antiproliferative effects of melatonin on the growth of rat pituitary prolactin-secreting tumor cells in vitro. J Pineal Res. 2007;42(2):172-9. [PubMed ID: 17286750]. https://doi.org/10.1111/j.1600-079X.2006.00403.x.
-
16.
Rubio S, Estevez F, Cabrera J, Reiter RJ, Loro J, Quintana J. Inhibition of proliferation and induction of apoptosis by melatonin in human myeloid HL-60 cells. J Pineal Res. 2007;42(2):131-8. [PubMed ID: 17286744]. https://doi.org/10.1111/j.1600-079X.2006.00392.x.
