1. Background
The hepatitis E virus (HEV) is a member of the Hepeviridae family, according to the ninth ICTV report. The genome of the virus is the RNA with positive sense (1). Hepatitis E is an important human gastrointestinal disease that has a worldwide spread. This disease can cause sporadic or epidemic cases of acute hepatitis (2). The HEV can be excreted from the feces of infected individuals and animals and may infect others mostly through drinking contaminated water. The disease is usually self-limiting. In some cases, there is a dire illness named acute liver failure, and some patients may die from it (3). The prevalence of hepatitis E depends on poor public health and the entry of the virus into drinking water sources (1). Other modes of transmission have been identified, but they cover only a very small fraction of the causes of disease. The consumption of uncooked infected meat or meat products, injection of contaminated blood products, vertical transmission from the pregnant mother to the fetus, and consumption of raw shellfish are some other ways of contracting the disease (4, 5).
The isolates of this virus can be classified into at least four phylogenetic or genotype groups based on geographical distribution, host diversity, and infection pattern. These genotypes are indistinguishable by serological tests. Genotype I is common in Asia and Africa, and its host is only the human. This genotype is related to the occurrence of the disease caused by drinking contaminated water (6, 7). Genotype II was first isolated from neither hepatitis A nor hepatitis B cases in 1986 in Mexico, and genotype III is common in pig farms in the USA, Europe, Canada, Mexico, New Zealand, South Korea, Japan, and Thailand. The HEV in sporadic and foodborne cases in non-native areas can cause severe hepatitis, especially in people dealing with pigs, resulting in an anti-HEV serological titer. Human and swine hepatitis E isolates are genetically very close in the United States, Britain, and Japan. This suggests that pigs could be a repository for the virus in humans. Genotype IV has been identified in pigs since 1985 and in humans since 1993 in China, Japan, India, Indonesia, and Vietnam have been found to have a wider range of hosts. This genotype is also associated with sporadic and malignant cases of hepatitis, but like genotype I, it does not cause a higher incidence of hepatitis E due to contaminated water consumption. Just like genotype III, genotype IV isolates are genetically very similar in humans and pigs (8, 9).
2. Objectives
Due to the limited reports on HEV detection in deer and the potential role of this animal in maintaining the virus as a reservoir, in the present study, the genome of the virus was searched in the fecal samples of Persian gazelle.
3. Methods
3.1. Sampling
Sampling was performed between the winter of 2017 and the winter of 2019 from the deer of the Moteh protected area and the deserts around Meymeh City near Isfahan by environmentalists and environmental activists of the Environmental Protection and Natural Resources Department of Meymeh City of Isfahan. Sampling was done from two groups of animals, including over one year and under one year of age, and from both males and females. Fresh fecal samples and gastrointestinal swabs taken from the animals were placed in sterile PBS and transported to the laboratory under ice, and the samples were kept at -70°C until the samples were tested. In this study, fecal samples and gastrointestinal swabs were prepared from 50 deer.
3.2. RNA Extraction and Reverse Transcription
To extract RNA from the prepared samples, the RNA extraction kit of RNA Biotechnology Company (Iran) was used, and the extraction process was performed according to the instructions of the manufacturer. Briefly, 1 mL of sample was added to 500 µL of the lysis buffer, and the mixture was placed at room temperature for five minutes. Then, 200 µL of chloroform was added and shaken vigorously for 10 - 15 seconds. Afterward, it was incubated at 25°C for six minutes. In the next step, the mixture was centrifuged for five minutes at 9,000 rpm. The supernatant was then transferred to another tube, and after adding 1000 µL of cold isopropanol, it was incubated at -20°C overnight. After centrifugation under the conditions of the previous step, the supernatant was discarded. Washing was then performed with 80% ethanol, and after discarding ethanol and evaporating remaining ethanol, the RNA was dissolved in 30 µL of deionized sterile water and stored at -70°C until further steps.
The cDNA synthesis was performed with a TaqMan RT Kit (Invitrogen, Cat no.: N8080234). For this purpose, the RT master mix consisted of 1 µL of 10× RT buffer, 2.4 µL of 25 mM MgCl2, 1.8 µL of 10 mM dNTPs, 0.4 µL of Random hexamers, 0.3 µL of RNase inhibitors, 0.3 µL of reverse transcriptase (MMLV RT) (50 U/µL), the RNA template (0.4 - 0.5 µg/µL), and deionized sterile water. The reaction was carried out at 25°C for 10 min, 48°C for 30 min, and 94°C for 5 min.
3.2.1. PCR
For the PCR assay, the primers were designed for HEV with the following sequences: HEVF: TTCCACCACCCAGCAGTATT-3’ and HEVR: GGCATTCTCAACGAGCAGTT-3’. The PCR was carried out at 95°C for four minutes for one cycle and 30 cycles of 95°C for 30 seconds, 54°C for 30 seconds, and 72°C for 30 seconds. The final extension was done at 72°C for four minutes. Afterward, the electrophoresis of PCR products for both controls and test samples was carried out in 1.5% agarose stained by the viewer. Then, the positive samples were sent to Bioneer Co. (Korea) for sequencing.
4. Results
4.1. RT-PCR Test
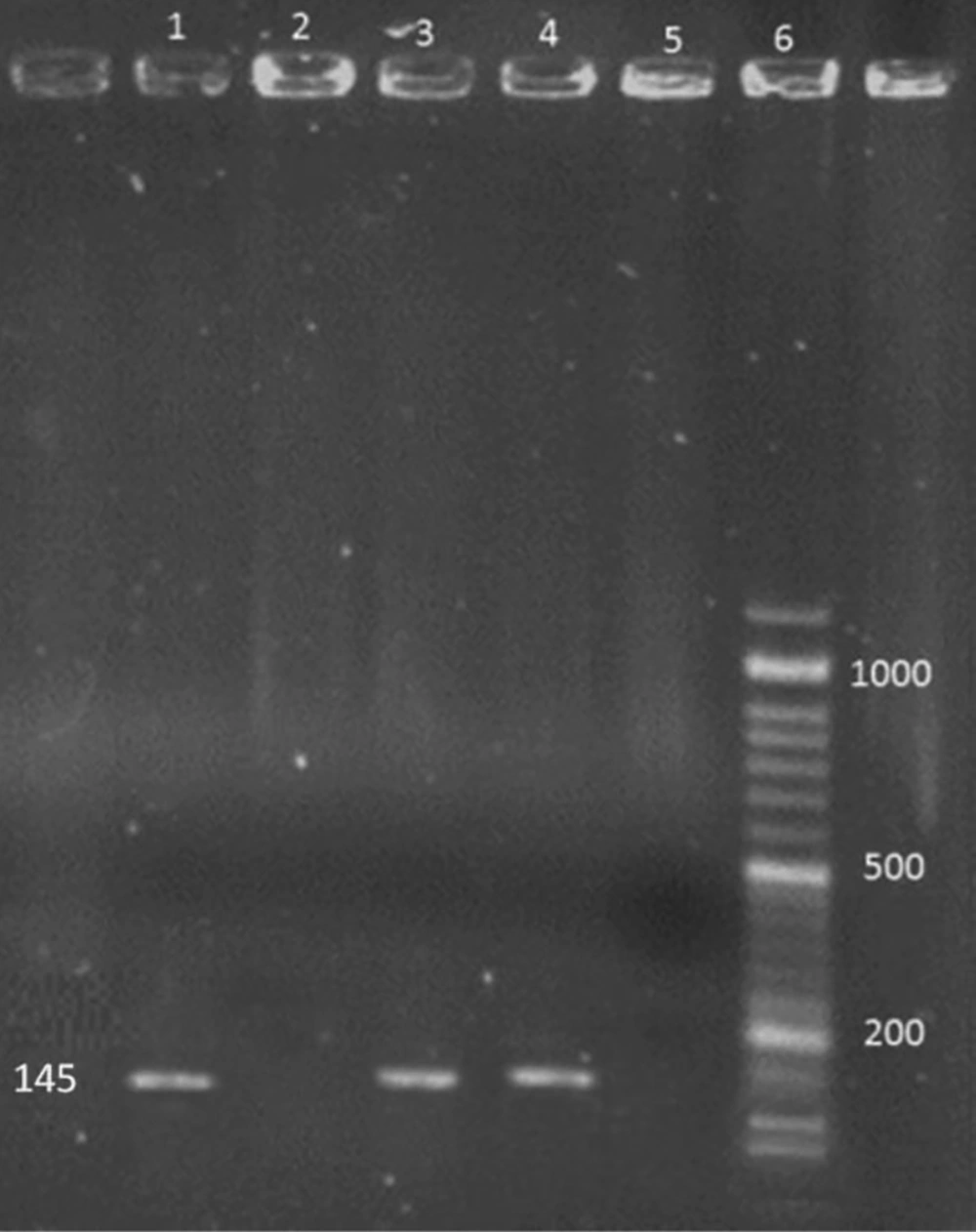
The HEV specific bands with the size of 145 bp were detected in positive controls and positive RT-PCR products, respectively (Figure 1).
4.2. Sequencing
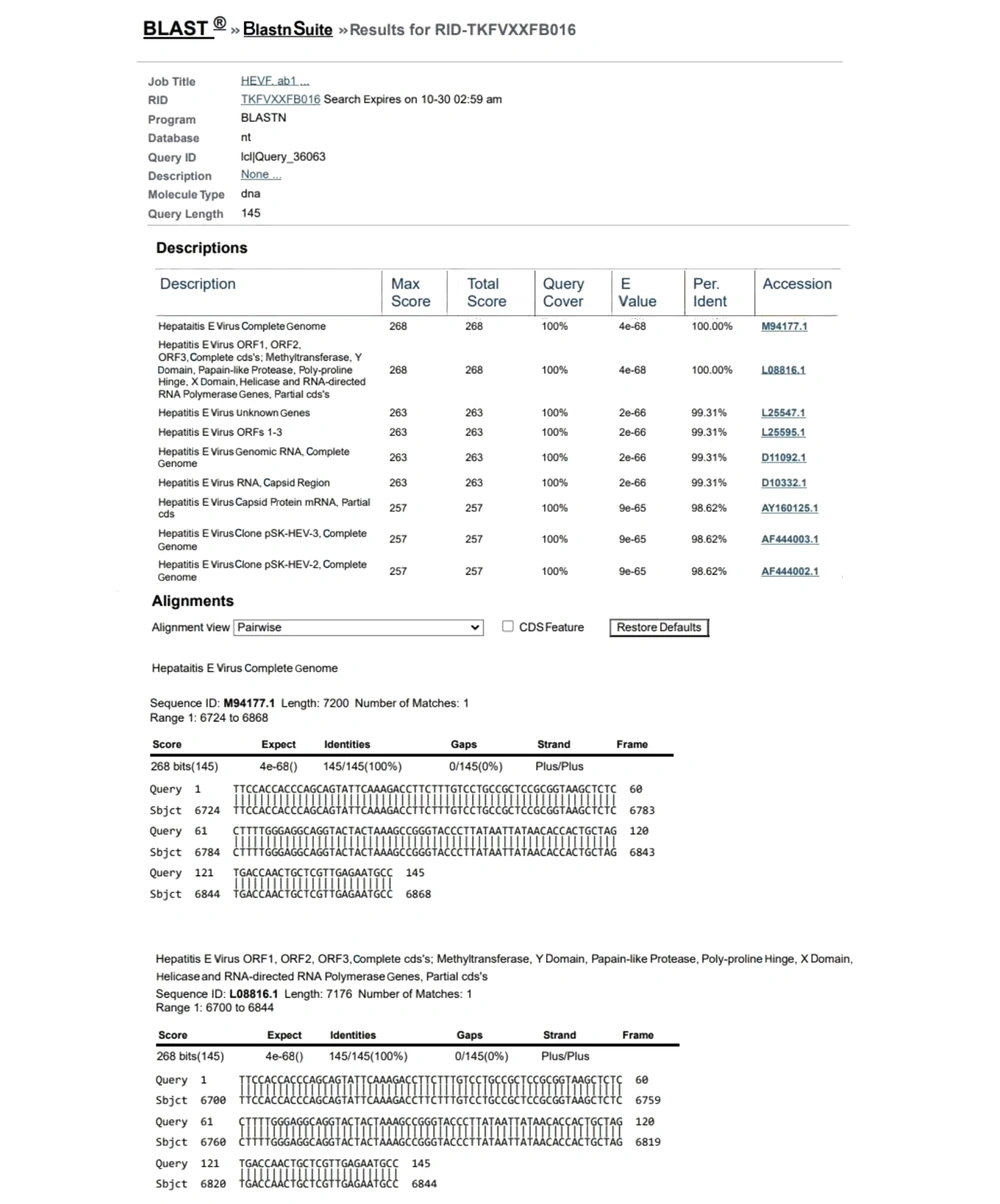
The degree of identity of the sequence extracted from the RT-PCR product reading of the studied genes with the sequences registered in the gene bank for hepatitis E virus was a good indicator of PCR adequacy (Figure 2).
4.3. HEV Prevalence Among Persian Gazelle
In total, out of 50 samples taken from deer in the study area using the RT-PCR test, three samples were HEV RNA-positive. The frequency of infection was two cases in deer above one-year of age and one case in deer under one year. All the three HEV RNA-positive samples were from female deer (Tables 1 and 2).
| Age | Total Frequency | Frequency of HEV-Positive Cases | Percentage of HEV Frequency |
|---|---|---|---|
| Over one year | 38 | 2 | 2 |
| Under one year | 12 | 1 | 2 |
| Total | 50 | 3 | 6 |
| Sex | Total Frequency | Frequency of HEV-Positive Cases | Percentage of HEV Frequency |
|---|---|---|---|
| Male | 18 | 0 | 0 |
| Female | 32 | 3 | 6 |
| Total | 50 | 3 | 6 |
4.4. Statistical Analysis
After entering data in SPSS version 16 software and drawing a two-way agreement table, the relationship between the variables was examined using the chi-square test, and no statistically significant relationship was found between the variables (P > 0.05).
5. Discussion
In 2015, Lhomme et al. (10) examined the genomic prevalence of HEV in wild boar (Sus scrofa), mountain deer (Cervus elaphus), wild rabbit (Oryctolagus cuniculus), and Coypu (Myocastor coypus) in all of Southwestern France. Except for Coypu, evidence of HEV was obtained in others. In general, the frequency of HEV RNA isolated from wildlife samples was 3.3%. Thus, this research suggested the potential of these animals as the sources of HEV infection for humans (10). Sonoda et al. (11) in Japan determined HEV infection in wild boar and deer and showed the presence of HEV in boar by genetic methods. Anti-HEV antibodies were detected in 9% of wild boars, and 2% of mountain deer tested, and hepatitis E virus genotype III was isolated from a boar. The results of their study showed that wild boar and deer were infected with HEV in Japan; however, the prevalence of HEV in wild boar was much lower than that of domestic pigs. There was a significant similarity between human and boar HEV isolates that showed HEV transmission to humans through human-animal feed (11). In the present study, the prevalence of hepatitis E infection was 6% in mountain deer, which was higher than in the study by Sonoda et al. (11)
In 2003, Tei et al. (12) reported HEV infection in individuals who ate raw deer meat. A remnant of deer meat, which was frozen, was HEV RNA-positive and had the same nucleotide sequence as the patient's RNA. Family members of patients who did not eat or ate small amounts of deer meat were negative for HEV. Therefore, the authors concluded that hepatitis E is a common zoonosis (12). In the present study, hepatitis E nucleic acid was identified in gastrointestinal and fecal swab samples. In 2020, Trojnar et al. (13) analyzed 108 blood samples and 106 liver samples from yellow deer, red deer, and Sika deer kept on 11 farms in Germany for HEV infection. Using sandwich ELISA, 2.7% of serum samples showed borderline edge results in terms of HEV-specific antibodies; but no positive samples were detected in liver samples. The results showed a low risk of HEV infection in farmed deer in Germany (13).
In 2015, Kukielka et al. (14) evaluated the rotation of the hepatitis virus in Iberian pigs and wild animals in Spain over eight years. By real-time RT-PCR, 12% of wild boar samples and 16.05% of red deer samples were positive. All of the pigs were negative. The serological analysis showed that 43.75% of Iberian pigs, 57.40% of wild boars, and 12.85% of red deer had anti-HEV antibodies. This study showed that the persistent HEV infection of wildlife makes a threat to human health (14). Zheng et al. (9) surveyed two pig farms in China and found that 9.6% of pigs, 0.3% of healthy people working on these farms, and 68.8% of people with proven HEV infections that dealt with pigs excreted the virus in their feces. Most patients showed genotype I, and some showed genotype IV and II. People who came in contact with pigs were 74% more likely to get HEV, and those who lived downstream of pig farms were more likely to be infected than those who lived upstream. Based on these data, the research group concluded that genotype IV of the virus circulates freely between humans and pigs, and the disease is more likely to be transmitted from pigs to humans than reciprocally. Transmission from pigs to humans is possible through direct contact, as well as contact with animal feces (9).
In 2015, Grierson et al. (15) examined the serological prevalence, active infection, and viral diversity of HEV variants in 629 pigs at an abattoir in the United Kingdom. They reported a serological and genomic prevalence of the virus as 92.8 and 15%, respectively (15). In 2004, Kasorndorkbua et al. (8)examined the methods of transmitting hepatitis E to the pig population and infected 27 pigs in a variety of ways. In this study, the main route of transmission was identified through the oral-fecal route, and transmission through infected needles or nasal swabs or tonsils was not confirmed. In 2005, Kasorndorkbua et al. (16) evaluated pig farms in the western United States for HEV contamination. Samples from various sources were assayed for HEV RNA by RT-PCR, and the results showed that seven samples from 28 farms had fecal samples containing HEV (16).
5.1. Conclusions
Overall, the results of the enumerated studies and the present study showed that HEV infection is present in wildlife in several parts of the world, including Iran. The data from the present study and similar data show that virus transmission through wild reservoirs in nature can be an important source of hepatitis E virus infection in humans.