Abstract
Background:
Acute lymphoblastic leukemia (ALL) is a malignant disorder of lymphoid progenitor cells that affects both children and adults. B-lineage acute lymphoblastic leukemia (B-ALL) that derived from primary B cell precursors is a common subtype of ALL. PAX5 is a member of PAX gene family. This gene is located at 9p13.2, encoding the B-cell lineage specific activator protein (BSAP). BSAP is an essential regulator of B lymphocytes identity and function which plays an important role in part of B cell specific genes.Objectives:
The aim of this study is to screen probable variants in flanking regions of introns 1 and 2 near the exons 1, 2 and 3 of PAX5 gene among B-ALL patients from Khuzestan province.Patients and Methods:
In this descriptive study, blood samples were collected from 50 patients with clinical symptoms of B-ALL in Khuzestan province. In order to identify the probable variants in introns 1 and 2 near the exons 1, 2 and 3 of PAX5 gene, flanking regions of introns amplified by PCR and the products were sequenced for any probable change.Results:
Two variants in nine patients were identified including IVS2-43T > C and IVS2 + 11T > G. IVS2-43T > C variant was found as a heterozygous form in one patient and IVS2 + 11T > G was found as a homozygous variant in 8 patients with B-ALL.Conclusions:
The overall frequency of variants in intron 2 of PAX5 gene was 18%. IVS2 + 11T > G variant of PAX5 gene probably do not associated with B-ALL risk in the population.Keywords
1. Background
Acute lymphoblastic leukemia (ALL) is a neoplasm of lymphoid progenitor cells and is the most common malignancy of pediatric [1]. Worldwide incidence projected is 1 - 4.75 per 100,000 people [2]. In patients with ALL, the incidence, severity, clinical symptoms and response to treatment are different [3, 4]. Despite significant advances in the successful treatment of ALL, the disease causes cancer-related death in many young people . Poor prognostic outcome of ALL is associated with white blood cell counts above 50000/mL [5]. The disease is determined due to genetic changes that arrest B and T lymphocytes differentiation, and causing aberrant cell proliferation and survival [6]. Paired box 5 (PAX5) is a transcription factor that plays a role in commitment of hematopoietic cells to B lymphocytes [7]. It is involved in B lineage cell development of several ways, including: activating the transcription of many of specific genes in B cell lines, suppressing the expression of lineage-inappropriate genes and regulation of VHdiversity-joining) recombination in immunoglobulin heavy chain (variable) to DJH (diversity-joining) recombination in immunoglobulin heavy chain gene (IgH) [8, 9]. The human PAX5 gene, which encodes the PAX5 protein, also known as the B-cell specific activator protein (BSAP), is situated at the 9p13 locus [10]. In the hematopoietic system, this protein exclusively expresses from the pro-B to the mature B cell stage, where it controls the differentiation, function, and development of B lymphocytes [11, 12]. Several studies have demonstrated that PAX5 is the main target of somatic mutations in B-ALL. A genome analysis from pediatric ALL patients was performed by Mullighan et al. and was identified deletion, amplification, point mutation and structural rearrangement in genes encoding development and differentiation of B lymphocytes that among them, PAX5 gene was main target of somatic mutation in 31.7% of children [13]. Familiades et al. examine a cohort of 117 adult B-ALL (B-lineage acute lymphoblastic leukemia) patients using oligo SNP arrays. They demonstrated that PAX5 is mutated in 34% of adult B-ALL [14].
It is well known that most eukaryotic genes have introns [15], which are integral elements of the genome that exert diverse important functions such as alternative splicing and also actively take part in gene regulation and evolution [16]. Majewski and Ott had shown that 20 nucleotides of introns from both ends could save less SNPs (single nucleotide polymorphism) in comparison to the other parts of introns [17]. So they concluded that these parts of introns may have role in regulation or process of splicing. Intron point mutations can sometimes lead to aberrant mRNA splicing that cause a protein truncated or representing an alternative mechanism for inactivation [16]. Therefore, it seems that mutation in some parts of intron would be very important. This study is performed to identify probable variants in flanking regions of introns 1 and 2 near the exons 1, 2 and 3 of PAX5 gene in patients with acute lymphoblastic leukemia in Khuzestan province, Southwest Iran.
2. Objectives
The aim of this study is to screen probable variants in flanking regions of introns 1 and 2 near the exons 1, 2 and 3 of PAX5 gene among B-ALL patients from Khuzestan province.
3. Patients and Methods
In this descriptive study, 50 patients selected with B-ALL (31 males and 19 females) that referred to Shafa hospital of Ahvaz. Among patients with leukemia, only patients with B-ALL were selected. The mean age of patients was 9.4 years (1 to 27 years old). The primary diagnosis of B-ALL was established on the basis of clinical phenotype followed by analysis of bone marrow sample using flow cytometry. After obtaining consent from parents of patients, blood samples were collected in tubes containing 0.5 M EDTA (CinnaGen company, Iran) solution and were maintained at -70°C and DNA was extracted using DNA extraction kit from Gen Fanavaran Company, Iran. Primers (Gen Fanavaran, Iran) for flanking regions of introns 1 and 2 near the exons 1, 2 and 3 were designed by primer 3 plus software (www.bioinformatics.nl/ primer3plus/). Primer sequences and annealing temperature for each primer pair are listed in Table 1. PCR was performed using thermocycler (Flex thermocycler, Germany) and in 35 rounds with the following condition: initial denaturation at 94°C during 5 minutes followed by 35 cycles at 94°C for 30 seconds, at 60°C for 45 seconds and at 72°C for 30 seconds, followed by one cycle of 72°C for 3 minutes. To confirm the size of the PCR products, samples were loaded on 1.5% agarose (CinnaGen Company, Iran) gel. Sequencing was performed for 50 samples using big dye terminator cycle sequencing ready reaction kit (Applied Biosystems, USA) on an ABI Prism 3500 automated genetic analyzer (applied Biosystems, USA). The reactions were performed using reverse primer in Table 1. Sequencing results were analysed using Chromas program and compared with the normal PAX5 gene sequence by NCBI BLAST tool.
After observation the variant in patients, blood sample were taken from 10 of parents of the patients (5 males and 5 females) and DNA extraction, PCR and sequencing was performed according to the above procedure. Statistical analysis was performed using SPSS-16 software (SPSS, Inc., Chicago, IL).
Sequence of the Forward and Reverse Primers Used for Flanking Regions of Introns 1 and 2 Near the Exons 1, 2 and 3
| Exon | Primer | Annealing Temperature, °C |
|---|---|---|
| 1 | 60 | |
| F | 5'- ATTATTCCGACCTGTGAGCG-3' | |
| R | 5'- GTGTCCCGGAGTTTGCAC-3' | |
| 2 | 60 | |
| F | 5'-AGCGGTGCTTCTCCTATGTG-3' | |
| R | 5'-ACAGCTGCTGGGTCATGTTT-3' | |
| 3 | 60 | |
| F | 5'-TAGCCCGTTATTTTGTTGCC-3' | |
| R | 5'- ATCCCTCCAAATCCCCAAC-3' |
4. Results
After the extraction of DNA from the total blood of 50 ALL patients, PCR of the mentioned regions was performed. The lengh of intron regions amplified in reaction for the initial and final parts of intron 1 was respectively 26 bp and 50 bp and for the initial and final parts of intron 2 was 49 bp and 54 bp. Direct sequencing of the regions was done and after comparison with PAX5 reference sequence (NG-033894), two variants were identified in 9 patients (Table 2).
Detected Variants From the Studied Samples
| Mutation | Location | Relative Frequency, % | |
|---|---|---|---|
| DNA level | Protein level | ||
| c.212 + 11T > G | IVS2 + 11T > G | Intron 2 | 16 |
| c.213 - 43T > C | IVS2 - 43T > C | Intron 2 | 2 |
Patients were seven males and two females from age 3 to 27 years. IVS2 - 43T > C variant was heterozygous in one patient. IVS2 + 11T > G variant was homozygous in 8 patients, therefore normal people had two T nucleotides and patients had two G nucleotides in two alleles of PAX5 gene. In Figure 1, the results of sequencing of variants have been shown
The Result of Sequencing
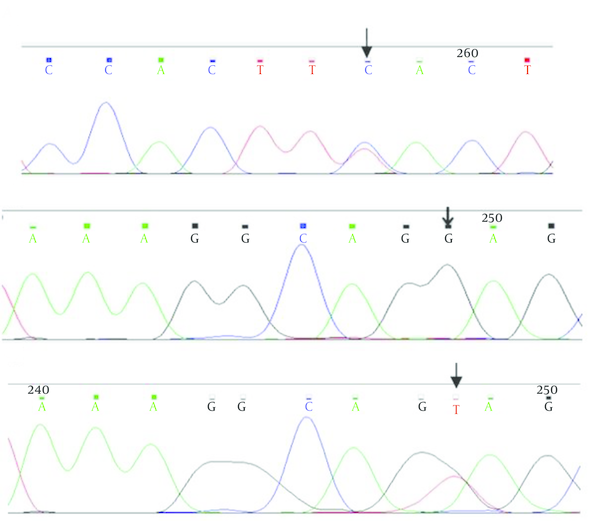
Sequencing of the PAX5 gene for parents of the patients revealed that, IVS2 + 11T > G homozygous variant exist in some of them too.
The Result of Sequencing of Homozygous IVS2 + 11T > G Variant in Intron 2 in the Patient’s Parents
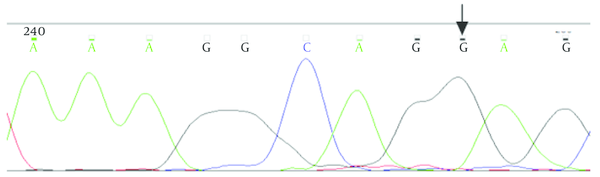
5. Discussion
In this study, we tried to find the probable variants in flanking regions of introns 1 and 2 near the exons 1, 2 and 3 of PAX5 gene in the population of Khuzestan patients with B-ALL. The flanking regions of introns and exons were sequenced directly and two variants were found that located in intron 2 of PAX5 gene. These two variants including IVS2 + 11T > G and IVS2-43T > C were identified in 9 of 50 patients, while in the 41 other patients, did not observe any mutation in these regions. So frequency of patients with variants in intron 2 of PAX5 gene in the population was 18%. IVS2 + 11T > G variant was shown in 6 males and 2 females in age range 3 to 27 years from different ethnic group and IVS2 - 43T > C variant was found in one 15 years-old boy with Arab ethnicity.
The PAX family of DNA binding proteins plays an important role in growth regulation in higher eukaryotes. Their importance is characterized by PAX gene mutations that caused developmental abnormalities such as Waardenburg syndrome and cancer [18]. As mentioned PAX5 is a member of PAX gene family [19] that is required for B-cell development and maintenance [20, 21] and so the gene could be significant target of somatic mutations in B-ALL [22]. On the other hand regulation of splicing of RNA transcripts encoding transcription factors is likely to play an important role during cell development and differentiation [23]. Pre-mRNA splicing and regulation of the processes are essential steps which leads to the removal of introns from immature RNA [24]. As regards alternative splicing occurs in most pre-mRNAs, it is not surprising that disturbance of normal splicing patterns could be cause human diseases [15]. For example it was found that the 20 nucleotides of intron from both ends are very important in the sequence and these areas may have less SNPs than the other parts of introns. This is because of the fact that changes in these areas may have effect on RNA splicing [17]. Since in mRNA splicing, as usual, the introns are removed and exons are joined together, any changes in this process, may effect on the normal removal of an intron (intron retention) or caused to delete an exon (exon skipping). Based on the studies, intron variants may effect on mRNA splicing, gene regulation and DNA-protein interactions especially binding of transcription factors [16, 25]. So sometimes, parts of introns especially near the exonic regions would be important for understanding the reasons of abnormalities.
IVS2 - 43T > C variant was found as a heterozygous variant 43 nucleotides before the AG dinucleotide of the 3'-acceptor splice site of intron 2 of PAX5 gene. This variant was revealed in one patient who was one 15 years old boy with Arab ethnicity.
Another variant which revealed in 8 patients was IVS2 + 11T > G, a homozygous form that located in the eleventh nucleotides after the GT dinucleotide of the 5′-donor splice site of intron 2 of PAX5 gene. These patients were 6 boys and 2 girls in ages between 3 to 27 years. So between these 8 patients, this variant was seen in boys more than girls with different ages. Also with respect to ethnic diversity of patients, this variant was seen in different ethnic groups.
Homozygous form of one variant means that change in this region can affect both alleles of this gene. To study the pathogenic effects of this variant, this region was evaluated in 10 of the parents of the patients (5 males and 5 females). In three cases, this variant was found in homozygous form (1 male and 2 females) and in seven cases, it was heterozygous (4 males and 3 females). Because of that 3 subjects that showed the variant as a homozygous form and they are healthy, indicates that this variant, do not have any role on the disease. Therefore, this variant can be one polymorphism that may be used in haplotype studies.
Despite the studies about this variant, to evaluate the frequency of this variant, more comprehensive studies with more samples and analysis of these variant in healthy control samples would help us for more understanding the role of the PAX5 gene in the disorder.
Acknowledgements
References
-
1.
Mullighan CG, Willman CL. Advances in the Biology of Acute Lymphoblastic Leukemia-From Genomics to the Clinic. J Adolesc Young Adult Oncol. 2011;1(2):77-86. [PubMed ID: 23610732]. https://doi.org/10.1089/jayao.2011.0012.
-
2.
Redaelli A, Laskin BL, Stephens JM, Botteman MF, Pashos CL. A systematic literature review of the clinical and epidemiological burden of acute lymphoblastic leukaemia (ALL). Eur J Cancer Care (Engl). 2005;14(1):53-62. [PubMed ID: 15698386]. https://doi.org/10.1111/j.1365-2354.2005.00513.x.
-
3.
Downing JR, Mullighan CG. Tumor-specific genetic lesions and their influence on therapy in pediatric acute lymphoblastic leukemia. Hematology Am Soc Hematol Educ Program. 2006:118-22. 508. [PubMed ID: 17124049]. https://doi.org/10.1182/asheducation-2006.1.118.
-
4.
Figueroa ME, Chen SC, Andersson AK, Phillips LA, Li Y, Sotzen J, et al. Integrated genetic and epigenetic analysis of childhood acute lymphoblastic leukemia. J Clin Invest. 2013;123(7):3099-111. [PubMed ID: 23921123]. https://doi.org/10.1172/JCI66203.
-
5.
Rahimi Z, Rahimi Z, Akramipour R. Prothrombin G20210A Mutation is not a Risk Factor for Pediatric Acute Lymphoblastic Leukemia in Western Iran. Middle East Journal of Cancer. 2013;4(4):139-43.
-
6.
Teitell MA, Pandolfi PP. Molecular genetics of acute lymphoblastic leukemia. Annu Rev Pathol. 2009;4:175-98. [PubMed ID: 18783329]. https://doi.org/10.1146/annurev.pathol.4.110807.092227.
-
7.
Meijerink JP, den Boer ML, Pieters R. New genetic abnormalities and treatment response in acute lymphoblastic leukemia. Semin Hematol. 2009;46(1):16-23. [PubMed ID: 19100364]. https://doi.org/10.1053/j.seminhematol.2008.09.006.
-
8.
Medvedovic J, Ebert A, Tagoh H, Busslinger M. Pax5: a master regulator of B cell development and leukemogenesis. Adv Immunol. 2011;111:179-206. [PubMed ID: 21970955]. https://doi.org/10.1016/B978-0-12-385991-4.00005-2.
-
9.
He T, Hong SY, Huang L, Xue W, Yu Z, Kwon H, et al. Histone acetyltransferase p300 acetylates Pax5 and strongly enhances Pax5-mediated transcriptional activity. J Biol Chem. 2011;286(16):14137-45. [PubMed ID: 21357426]. https://doi.org/10.1074/jbc.M110.176289.
-
10.
Arseneau JR, Laflamme M, Lewis SM, Maicas E, Ouellette RJ. Multiple isoforms of PAX5 are expressed in both lymphomas and normal B-cells. Br J Haematol. 2009;147(3):328-38. [PubMed ID: 19725825]. https://doi.org/10.1111/j.1365-2141.2009.07859.x.
-
11.
Iacobucci I, Lonetti A, Paoloni F, Papayannidis C, Ferrari A, Storlazzi CT, et al. The PAX5 gene is frequently rearranged in BCR-ABL1-positive acute lymphoblastic leukemia but is not associated with outcome. A report on behalf of the GIMEMA Acute Leukemia Working Party. Haematologica. 2010;95(10):1683-90. [PubMed ID: 20534699]. https://doi.org/10.3324/haematol.2009.020792.
-
12.
McManus S, Ebert A, Salvagiotto G, Medvedovic J, Sun Q, Tamir I, et al. The transcription factor Pax5 regulates its target genes by recruiting chromatin-modifying proteins in committed B cells. EMBO J. 2011;30(12):2388-404. [PubMed ID: 21552207]. https://doi.org/10.1038/emboj.2011.140.
-
13.
Mullighan CG, Goorha S, Radtke I, Miller CB, Coustan-Smith E, Dalton JD, et al. Genome-wide analysis of genetic alterations in acute lymphoblastic leukaemia. Nature. 2007;446(7137):758-64. [PubMed ID: 17344859]. https://doi.org/10.1038/nature05690.
-
14.
Familiades J, Bousquet M, Lafage-Pochitaloff M, Bene MC, Beldjord K, De Vos J, et al. PAX5 haploinsufficiency is associated with BCR-ABL1 and TCF3-PBX1 fusion genes: a GRAALL study. Leukemia. 2009;23(11):1989-98. [PubMed ID: 19587702]. https://doi.org/10.1038/leu.2009.135.
-
15.
Faustino NA, Cooper TA. Pre-mRNA splicing and human disease. Genes Dev. 2003;17(4):419-37. [PubMed ID: 12600935]. https://doi.org/10.1101/gad.1048803.
-
16.
Najjar Sadeghi R, Damavand B, Vahedi M, Mohebbi SR, Zojazi H, Molaei M, et al. Detection of p53 common intron polymorphisms in patients with gastritis lesions from Iran. Asian Pac J Cancer Prev. 2013;14(1):91-6. [PubMed ID: 23534810].
-
17.
Majewski J, Ott J. Distribution and characterization of regulatory elements in the human genome. Genome Res. 2002;12(12):1827-36. [PubMed ID: 12466286]. https://doi.org/10.1101/gr.606402.
-
18.
Fitzsimmons D, Lukin K, Lutz R, Garvie CW, Wolberger C, Hagman J. Highly cooperative recruitment of Ets-1 and release of autoinhibition by Pax5. J Mol Biol. 2009;392(2):452-64. [PubMed ID: 19616560]. https://doi.org/10.1016/j.jmb.2009.07.028.
-
19.
Blake JA, Ziman MR. Pax genes: regulators of lineage specification and progenitor cell maintenance. Development. 2014;141(4):737-51. [PubMed ID: 24496612]. https://doi.org/10.1242/dev.091785.
-
20.
O'Brien P, Morin PJ, Ouellette RJ, Robichaud GA. The Pax-5 gene: a pluripotent regulator of B-cell differentiation and cancer disease. Cancer Res. 2011;71(24):7345-50. [PubMed ID: 22127921]. https://doi.org/10.1158/0008-5472.CAN-11-1874.
-
21.
An Q, Wright SL, Konn ZJ, Matheson E, Minto L, Moorman AV, et al. Variable breakpoints target PAX5 in patients with dicentric chromosomes: a model for the basis of unbalanced translocations in cancer. Proc Natl Acad Sci U S A. 2008;105(44):17050-4. [PubMed ID: 18957548]. https://doi.org/10.1073/pnas.0803494105.
-
22.
Coyaud E, Struski S, Prade N, Familiades J, Eichner R, Quelen C, et al. Wide diversity of PAX5 alterations in B-ALL: a Groupe Francophone de Cytogenetique Hematologique study. Blood. 2010;115(15):3089-97. [PubMed ID: 20160164]. https://doi.org/10.1182/blood-2009-07-234229.
-
23.
Zwollo P, Arrieta H, Ede K, Molinder K, Desiderio S, Pollock R. The Pax-5 gene is alternatively spliced during B-cell development. J Biol Chem. 1997;272(15):10160-8. [PubMed ID: 9092562].
-
24.
Arenas M, Duley J, Sumi S, Sanderson J, Marinaki A. The ITPA c.94C>A and g.IVS2+21A>C sequence variants contribute to missplicing of the ITPA gene. Biochim Biophys Acta. 2007;1772(1):96-102. [PubMed ID: 17113761]. https://doi.org/10.1016/j.bbadis.2006.10.006.
-
25.
Harland M, Mistry S, Bishop DT, Bishop JA. A deep intronic mutation in CDKN2A is associated with disease in a subset of melanoma pedigrees. Hum Mol Genet. 2001;10(23):2679-86. [PubMed ID: 11726555].