1. Background
Acute kidney injury (AKI) is defined as an abrupt loss of kidney function or damage within hours or days (1). AKI is one of the most significant complications after surgery, especially after cardiac surgery (CS-AKI) and major non-cardiac surgery (2). As it accompanies poor short- and long-term outcomes and high costs of care, prevention of its occurrence could be a target for health care systems (3).
Every year more than 2 million cardiac surgeries are performed, and the occurrence of CS-AKI is reported to be from 5 to 47% (2, 4). CS-AKI is considered the second most common cause of AKI (after sepsis) (4). It is also associated with increased mortality, morbidity, length of stay, and higher medical costs (3). Based on statistics, more than 25% of patients who develop severe AKI will die within a year. Even after complete renal recovery, the mortality rate remains high for over ten years (1).
Creatinine and urine output are the critical determinants of stages of AKI. These two variables describe risk, injury, failure, loss, end-stage kidney disease (RIFLE), and kidney disease improving global outcomes (KDIGO). Unfortunately, postoperative creatinine levels may arise due to dehydration or muscle injury, which does not allow it to become a reliable indicator for AKI (5). On the other hand, reliance on creatinine as a diagnostic tool for AKI would hasten the treatment as it may rise several hours after injury (6). Therefore, any diagnostic tool to predict the occurrence of AKI, especially after cardiac surgery, would result in a better outcome for the patient and the health care system.
Although the pathophysiology of CS-AKI remains complex and completely understood, multiple pathways are proposed to participate in these complications (4). Hypoperfusion, ischemia-reperfusion injury, neurohumoral activation, inflammation, oxidative stress, nephrotoxins, and mechanical factors are pathways for CS-AKI development (4). Therefore, many preoperative, intraoperative, and postoperative factors could be described as risk factors.
Due to the complexity of CS-AKI pathophysiology (3), traditional statistical methods are unsuitable for this enormous amount of information. Therefore, machine learning (ML) approaches are proposed to stratify and specify recognizable factors. Although artificial intelligence (AI) and ML have become frontier technologies worldwide (7), these tools remain embryonic in the healthcare discipline (8).
Studies have proposed different learning models for the prediction of AKI after severe burns (8), thoracoabdominal aortic aneurysms (7), and continuous risk prediction of future deterioration in patients (9).
2. Objectives
This study was conducted to develop a predictive tool for CS-AKI. In this regard, we have tried to use interpretable ML methods to predict CS-AKI.
The rest of this paper is organized as follows. Section 2 describes the materials, the proposed method, study variables, and evaluation metrics. Section 3 presents the results of the different experiments separately. Finally, section 4 discusses the results and section 5 is the conclusion.
3. Methods
3.1. Study Design
The Research Ethics Committee of Shahid Beheshti University of Medical Sciences approved this retrospective/prospective multi-center cohort study in March 2019 (Ethics ID: IR.SBMU.MSP.REC.1398.175). Therefore, the records of 1435 patients undergoing heart surgeries, including coronary artery bypass grafting (CABG), valvular, transplant, and aortic, between 2019 and 2021 were reviewed, of which information of 1250 patients was retrospectively collected (by A. E. and M. A.). The data of 185 patients were added to the study prospectively after obtaining their consent. The required data were collected from three hospitals: Imam Hossein, Shahid Modarres, and Masih Daneshvari, located in Tehran. Patients are anonymous at all stages of this study.
3.2. Study Variables
The data collected in this study are based on the variables listed in Table 1. These variables are considered in six categories, including demographic and surgery type characteristics, past medical history (PMH), drug history (DH), lab data, anesthesia and surgery information, and postoperative variables, and are inspired by previous studies (10). The only exclusion criterion for this study is age < 18.
| Features | Frequency (%) | Range of Values | Mean ± SD |
|---|---|---|---|
| Demographic and surgery type characteristics | |||
| Age | 18 - 86 | 58.08 ± 11.26 | |
| Gender | |||
| Female | 570 (39.72) | ||
| Male | 865 (60.28) | ||
| BMI (Kg/m2) | 16.41 - 48.13 | 26.72 ± 3.64 | |
| Types of cardiac surgery | |||
| CABG | 1007 (70.17) | ||
| Valvular | 298 (20.77) | ||
| Transplant | 23 (1.60) | ||
| Aortic | 67 (4.67) | ||
| Other types of surgery | 21 (1.46) | ||
| Missing values | 19 (1.32) | ||
| Surgical settings | |||
| Elective | 1222 (85.16) | ||
| Emergency | 206 (14.35) | ||
| Missing values | 7 (0.49) | ||
| PMH | |||
| HTN | |||
| Positive | 941 (65.57) | ||
| Negative | 468 (32.61) | ||
| Missing values | 26 (1.81) | ||
| DM | |||
| Positive | 681 (47.46) | ||
| Negative | 726 (50.59) | ||
| Missing values | 28 (1.95) | ||
| CKD | |||
| Positive | 182 (12.68) | ||
| Negative | 1221 (85.09) | ||
| Missing values | 32 (2.23) | ||
| PHTN | |||
| Positive | 528 (36.79) | ||
| Negative | 863 (60.14) | ||
| Missing values | 44 (3.07) | ||
| COPD | |||
| Positive | 191 (13.31) | ||
| Negative | 1213 (84.53) | ||
| Missing values | 31 (2.16) | ||
| Stenting | |||
| Positive | 345 (24.04) | ||
| Negative | 1059 (73.80) | ||
| Missing values | 31 (2.16) | ||
| CVA | |||
| Positive | 148 (10.31) | ||
| Negative | 1256 (87.53) | ||
| Missing values | 31 (2.16) | ||
| 3VD | |||
| Positive | 881 (61.39) | ||
| Negative | 531 (37.00) | ||
| Missing values | 23 (1.60) | ||
| DH | |||
| ACEI | |||
| Positive | 478 (33.31) | ||
| Negative | 891 (62.09) | ||
| Missing values | 66 (4.60) | ||
| ARB | |||
| Positive | 512 (35.68) | ||
| Negative | 858 (59.79) | ||
| Missing values | 65 (4.53) | ||
| BB | |||
| Positive | 949 (66.13) | ||
| Negative | 422 (29.41) | ||
| Missing values | 64 (4.46) | ||
| Diuretics | |||
| Positive | 676 (47.11) | ||
| Negative | 693 (48.29) | ||
| Missing values | 66 (4.60) | ||
| CCB | |||
| Positive | 320 (22.30) | ||
| Negative | 1049 (73.10) | ||
| Missing values | 66 (4.60) | ||
| Statin | |||
| Positive | 1020 (71.08) | ||
| Negative | 349 (24.32) | ||
| Missing values | 66 (4.60) | ||
| ASA | |||
| Positive | 1074 (74.84) | ||
| Negative | 296 (20.63) | ||
| Missing values | 65 (4.53) | ||
| NSAIDs | |||
| Positive | 98 (6.83) | ||
| Negative | 1270 (88.50) | ||
| Missing values | 67 (4.67) | ||
| LAB | |||
| EF (%) | 10 - 66 | 47.67 ± 9.66 | |
| Cr (mg/dL) | 0.5 - 12 | 1.26 ± 0.84 | |
| Alb (g/dL) | 1.8 - 6.8 | 3.83 ± 0.66 | |
| BS (mg/dL) | 65 - 569 | 160.21 ± 59.76 | |
| Hb A1C (%) | 2.8 - 13.7 | 6.35 ± 1.47 | |
| Hct (%) | 15.8 - 56 | 39.02 ± 4.66 | |
| Anesthesia and surgery | |||
| Anesthesia time (min) | 130 - 960 | 341.75 ± 75.11 | |
| Crystalloid (Lit) | 0.3 - 4.5 | 1.43 ± 0.20 | |
| Colloid | |||
| Positive | 1080 (75.26) | ||
| Negative | 323 (22.51) | ||
| Missing values | 32 (2.23) | ||
| Colloid type | |||
| HTS | 3 (0.21) | ||
| Albumin | 1079 (75.19) | ||
| HES | 0 (0) | ||
| Gelatin | 0 (0) | ||
| Not prescribed | 323 (22.51) | ||
| Missing values | 30 (2.09) | ||
| Colloid dose (cc) | 5000 - 30000 | 10070.12 ± 1150.54 | |
| P.C | |||
| Positive | 661 (46.06) | ||
| Negative | 762 (53.10) | ||
| Missing values | 12 (0.84) | ||
| FFP | |||
| Positive | 840 (58.54) | ||
| Negative | 585 (40.77) | ||
| Missing values | 10 (0.69) | ||
| Diuretics | |||
| Positive | 1382 (96.31) | ||
| Negative | 39 (2.72) | ||
| Missing values | 14 (0.97) | ||
| CPB time (min) | 19 - 400 | 111.55 ± 37.15 | |
| Hemofiltration | |||
| Positive | 1113 (77.56) | ||
| Negative | 317 (22.09) | ||
| Missing values | 5 (0.35) | ||
| Post-Op | |||
| Inotrope | |||
| Positive | 278 (19.37) | ||
| Negative | 1126 (78.47) | ||
| Missing values | 31 (2.16) | ||
| Cr-1 (mg/dL) | 0.46 - 11.4 | 1.55 ± 0.83 | |
| Cr-7 (mg/dL) | 0.081 - 9 | 1.39 ± 0.80 | |
| Dialysis | |||
| Positive | 101 (7.04) | ||
| Negative | 1299 (90.52) | ||
| Missing values | 35 (2.44) | ||
| Nephrology consultation | |||
| Positive | 304 (21.18) | ||
| Negative | 1095 (76.31) | ||
| Missing values | 36 (2.51) |
Abbreviations: BMI, body mass index; CABG, coronary artery bypass graft; PMH, past medical history; HTN, hypertension; DM, diabetes mellitus; CKD, chronic kidney disease; PHTN, pulmonary hypertension; COPD, chronic obstructive pulmonary disease; CVA, cerebrovascular accident; 3VD, three vessels disease; DH, drug history; ACEI, angiotensin convertase enzyme inhibitor; ARB, angiotensin receptor blocker; BB, beta blocker; CCB, calcium channel blocker; ASA, acetylsalicylic acid; NSAIDs, nonsteroidal anti-inflammatory drugs; EF, ejection fraction; Cr, creatinine; Alb, albumin; BS, blood sugar; Hb A1C, glycated hemoglobin; Hct, hematocrit; HTS, hetastarch; HES, hydroxyethyl starch; P.C, packed red cells; FFP, fresh frozen plasma; CPB time, cardiopulmonary bypass time; Cr-1, creatinine day 1 postoperative; Cr-7, creatinine day 7 postoperative.
Cr-1 and Cr-7 variables are related to serum creatinine levels in patients in the first and seventh days after surgery, respectively. These variables and variable Cr, which shows preoperative serum creatinine levels in patients, are used to determine stages of AKI based on the KDIGO criterion. Based on this criterion, labels related to predicting the incidence of AKI in patients after cardiac surgery are determined. The number of patients based on their AKI stage in the first and seventh days after surgery is shown in Table 2.
| Stage of AKI | ||||
|---|---|---|---|---|
| No AKI | Stage 1 | Stage 2 | Stage 3 | |
| First postoperative day | 763 | 571 | 94 | 7 |
| Seventh postoperative day | 1022 | 367 | 41 | 5 |
3.3. Proposed Method
This study used several machine learning approaches to determine the best machine learning technique for predicting AKI after cardiac surgery. Well-known machine learning methods were considered for this prediction, such as support vector machine (SVM) (11), multi-layer perceptron (MLP) (12), decision tree (DT) (13), random forest (RF) (14), logistic regression (15), XGBoost (16) and AdaBoost (17). All these methods were applied to classify AKI in patients, and their results were obtained in the form of the area under the curve (AUC), precision (PR), recall (RE), F1-score, and accuracy (ACC). Sklearn and XGBoost packages were used in Python to generate codes related to machine learning techniques. The variables in Table 1 were used as input features to machine learning methods for AKI prediction in patients undergoing cardiac surgery. The two corresponding labels for each patient were calculated based on preoperative and postoperative Cr values according to KDIGO criteria. Due to the imbalance in the number of data for each AKI stage, two classes could be considered for each patient's labels, including no AKI or AKI.
Since some data were not recorded in patients' medical files when registering the information, it was necessary to address the missing values correctly. For this purpose, various methods were used to handle the missing values. The issue of missing values was investigated by the mean, median, k-nearest neighbors (KNN) (18), iterative multiple imputations, and dropping methods, and the results were reported for all these methods. Synthetic minority oversampling technique (SMOTE) (19) and class weight methods were also used to balance the two classes' data. Besides, for reliable results, all experiments in this study were performed in 10-fold cross-validation.
One of the main contributions of this study was to address interpretable machine learning models. For this purpose, after conducting experiments and selecting the best machine learning model for predicting AKI after cardiac surgery, examining the interpretability of this model was also on the agenda. The interpretability of the machine learning model determines whether the model has been predicted based on appropriate characteristics. For this purpose, the feature ranking method, local interpretable model-agnostic explanations (LIME) (20), and Shapley (21) were used to examine the interpretability of machine learning models in this article.
3.4. Evaluation Metrics
Various metrics, including PR, RE, F1-score, ACC, and AUC, were used to evaluate and compare the machine learning methods.
Where TP, FP, TN, and FN indicate true positive, false positive, true negative, and false negative, respectively. AUC denotes the overall success of an experiment where
3.5. Experimental Setup
A set of experiments were designed to predict AKI after cardiac surgery. In the first experiment, machine learning methods, including SVM, MLP, DT, RF, logistic regression, XGBoost, and AdaBoost, were compared for this prediction. This was done to select the most appropriate approach regarding AUC and other evaluation metrics. In the second experiment, methods for overcoming missing values and balancing data were evaluated to reveal these methods' impact on the results obtained from machine learning methods. Finally, in the third experiment, we examined the interpretability of the results to clarify machine learning methods' performance. These experiments were conducted using 80% of the data for model training and 20% for model testing. Additionally, the experiments were done using 10-fold cross-validation, so the results were reliable.
4. Results
4.1. AKI Prediction Using Machine Learning Methods
This section presents the results of the first and second experiments. According to KDIGO criteria, out of 1435 patients studied, 571 patients with AKI stage 1, 94 patients with AKI stage 2, 7 patients with AKI stage 3, and 763 patients without AKI were identified on the first day after cardiac surgery. These values are 367, 41, 5, and 1022 for the seventh day after cardiac surgery. Since the balance of the number of samples is very effective in using machine learning methods, AKI was predicted in patients in two classes with AKI and without AKI. Accordingly, the number of patients in classes with AKI and without AKI on the first day after cardiac surgery is 672 and 763, respectively. These values are 413 and 1022 for the seventh day after cardiac surgery. The results of using machine learning techniques to diagnose AKI after cardiac surgery are shown in Tables 3 and 4 for the first and seventh day after cardiac surgery. Since some patients' information is missing due to inappropriate file registration, the methods for handling missing values have been used; these methods' results are presented in Tables 3 and 4. The results of these experiments based on PR, RE, F1-score, and ACC metrics are given in Appendix 1 and 2 in Supplementary File.
| ML Method | Missing Value Imputation | ||||
|---|---|---|---|---|---|
| Multiple Imputation | Mean | Median | KNN | Drop | |
| SVM | 0.73 ± 0.05 | 0.69 ± 0.04 | 0.71 ± 0.03 | 0.72 ± 0.05 | 0.72 ± 0.04 |
| MLP | 0.73 ± 0.04 | 0.69 ± 0.03 | 0.70 ± 0.03 | 0.69 ± 0.05 | 0.71 ± 0.02 |
| DT | 0.66 ± 0.04 | 0.64 ± 0.03 | 0.61 ± 0.04 | 0.64 ± 0.02 | 0.66 ± 0.04 |
| Logistic regression | 0.75 ± 0.03 | 0.73 ± 0.03 | 0.74 ± 0.03 | 0.74 ± 0.04 | 0.73 ± 0.05 |
| AdaBoost | 0.78 ± 0.03 | 0.76 ± 0.04 | 0.77 ± 0.02 | 0.77 ± 0.03 | 0.80 ± 0.04 |
| XGBoost | 0.80 ± 0.05 | 0.78 ± 0.04 | 0.78 ± 0.03 | 0.79 ± 0.04 | 0.81 ± 0.04 |
| RF | 0.81 ± 0.05 | 0.78 ± 0.03 | 0.79 ± 0.03 | 0.79 ± 0.04 | 0.80 ± 0.04 |
Abbreviations: SVM, support vector machine; MLP, multi-layer perceptron; DT, decision tree; RF, random forest.
| ML Method | Missing Value Imputation | ||||
|---|---|---|---|---|---|
| Multiple Imputation | Mean | Median | KNN | Drop | |
| SVM | 0.73 ± 0.07 | 0.71 ± 0.04 | 0.72 ± 0.04 | 0.73 ± 0.05 | 0.67 ± 0.08 |
| MLP | 0.71 ± 0.07 | 0.69 ± 0.04 | 0.70 ± 0.05 | 0.71 ± 0.04 | 0.65 ± 0.07 |
| DT | 0.65 ± 0.05 | 0.62 ± 0.03 | 0.65 ± 0.04 | 0.62 ± 0.03 | 0.61 ± 0.06 |
| Logistic regression | 0.73 ± 0.05 | 0.71 ± 0.03 | 0.73 ± 0.03 | 0.72 ± 0.05 | 0.66 ± 0.05 |
| AdaBoost | 0.78 ± 0.04 | 0.76 ± 0.03 | 0.77 ± 0.05 | 0.76 ± 0.03 | 0.74 ± 0.06 |
| XGBoost | 0.80 ± 0.05 | 0.77 ± 0.04 | 0.80 ± 0.04 | 0.78 ± 0.03 | 0.76 ± 0.05 |
| RF | 0.79 ± 0.05 | 0.77 ± 0.04 | 0.79 ± 0.05 | 0.78 ± 0.03 | 0.76 ± 0.05 |
Abbreviations: SVM, support vector machine; MLP, multi-layer Perceptron; DT, decision tree; RF, random forest.
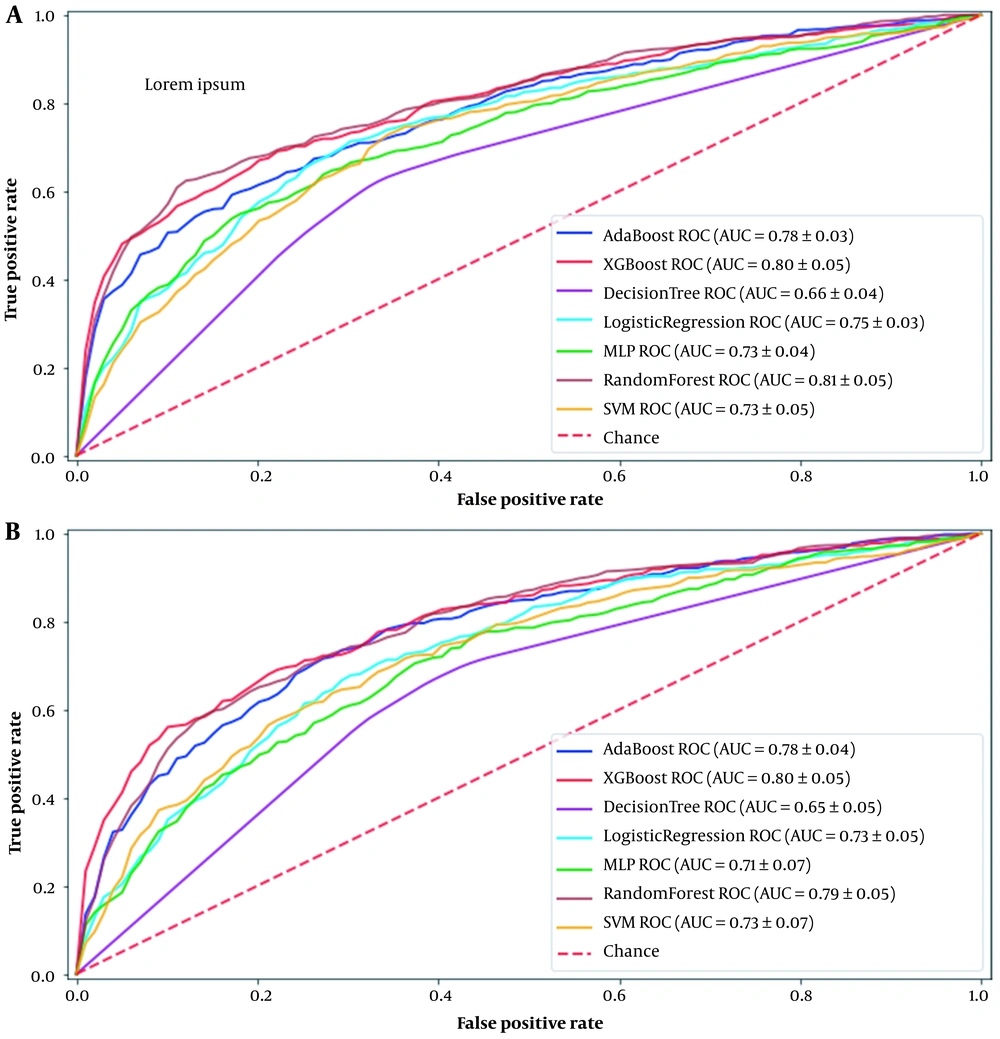
AUC comparisons of the different machine learning methods used in this study for the first and seventh days after cardiac surgery are shown in Figure 1. These curves are drawn for the best method for handling the missing values, which is iterative multiple imputations. Based on the results, the best machine learning models for predicting AKI in patients after cardiac surgery are RF and XGboost, with AUC 0.81 and 0.80 for the first day and 0.79 and 0.80 for the seventh day, respectively.
4.2. Interpretability of Machine Learning Models in AKI Prediction
After using the machine learning models in AKI prediction, we must pay attention to these models' interpretability. Despite their ability to predict, machine learning models do not explain to users how to make predictions. Due to the vital importance of predictions in medical applications, it is necessary to ensure their reliability for users. Therefore, this section investigates the best machine learning models' interpretability in AKI prediction after cardiac surgery using feature ranking, LIME, and Shapley methods.
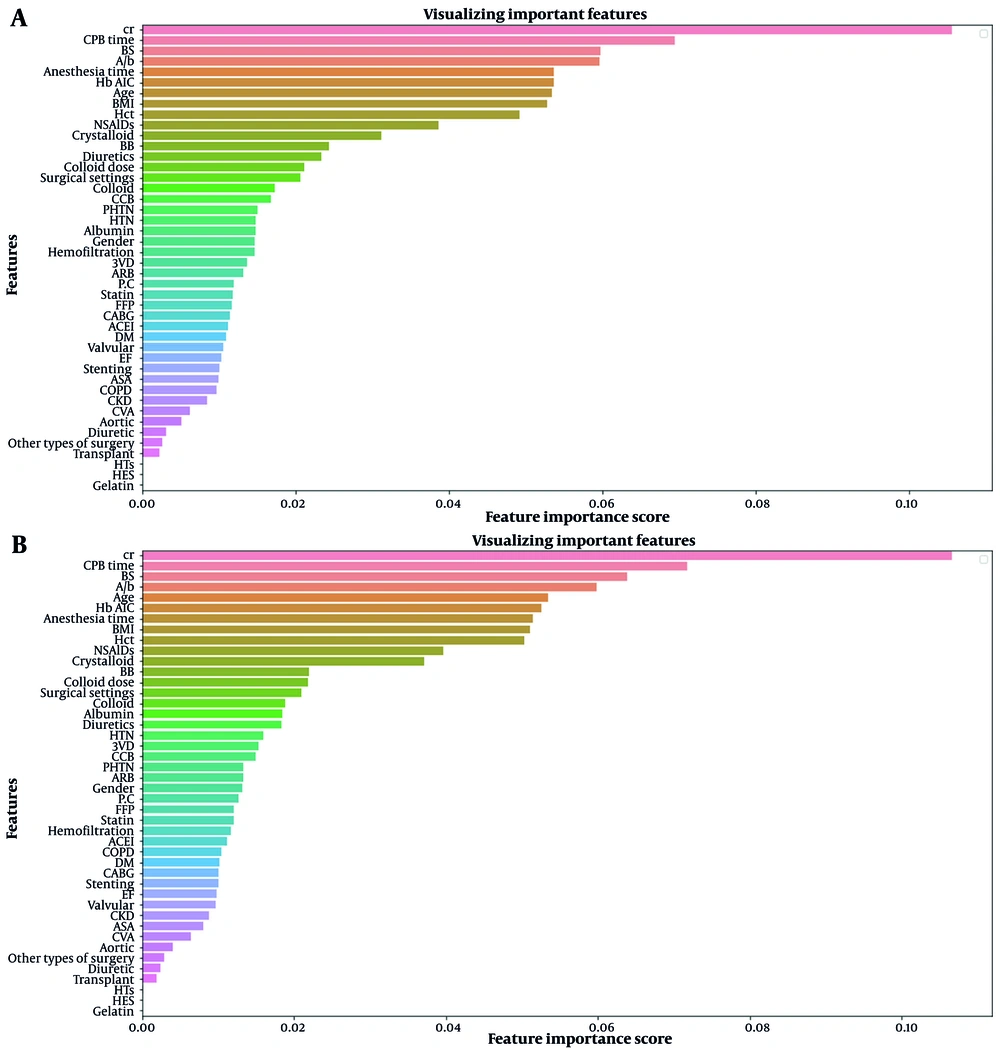
Figure 2 shows the importance ranking of features for AKI prediction on the first day after surgery using XGBoost and RF models. These charts illustrate which features influence the predictions made by these machine learning models. Appendix 3 in Supplementary File shows the same charts for the seventh day after surgery.
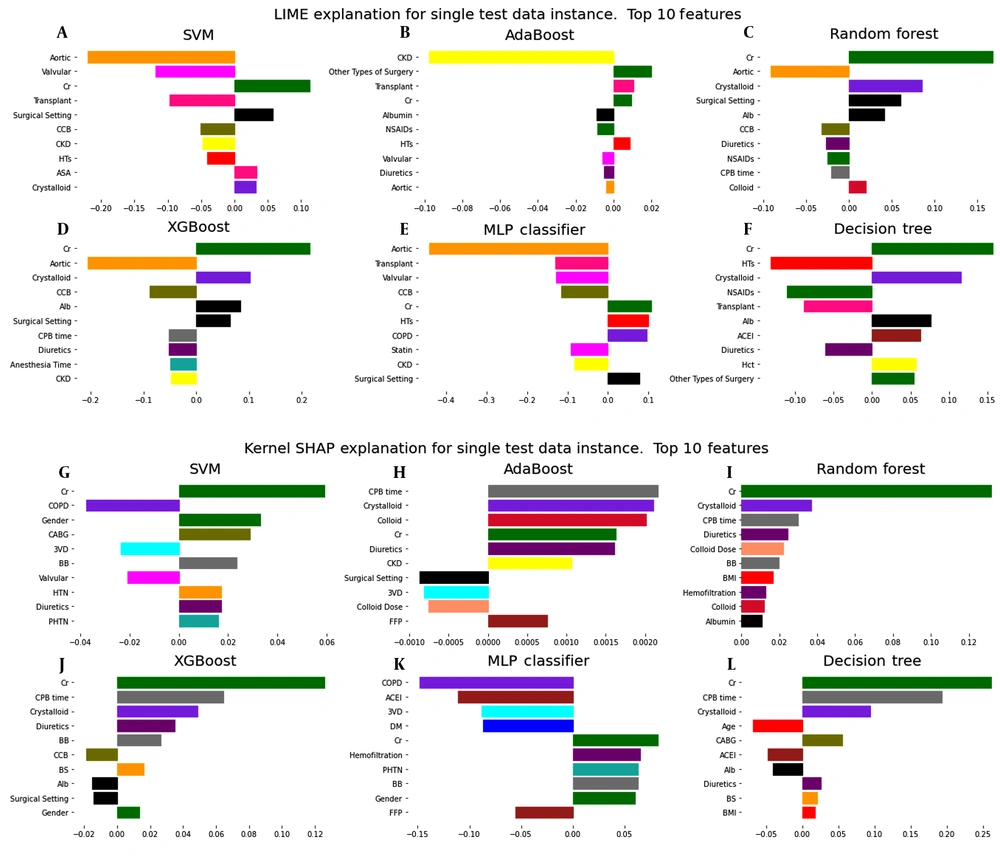
The models' interpretability in LIME and Shapley methods should be checked for each available sample. For a more accurate understanding of the interpretability of the models used to predict AKI on the first day after surgery, the data of a patient who was predicted to be TP were analyzed by LIME and Shapley methods. Figure 3 shows the results of these analyses for different machine learning models. Similar results are given for the seventh day after surgery in Appendix 4 in Supplementary File.
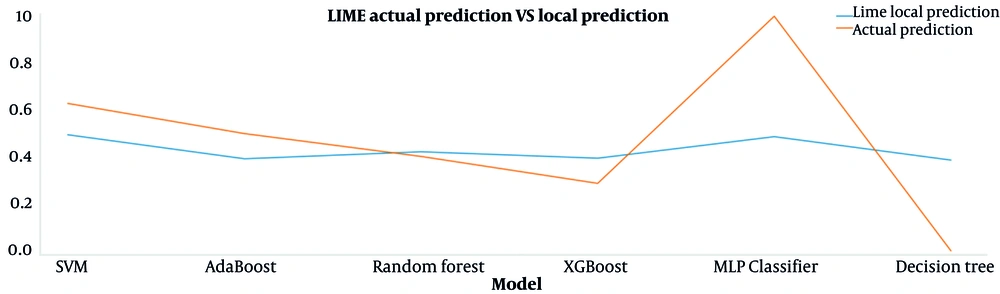
The results of the predictions made by the local LIME model and the main machine learning models for the interpreted data sample are presented in Figure 4. The proximity of each machine learning model's values indicates that LIME's interpretation of that model is trustworthy. Corresponding results for the seventh day after surgery are shown in Appendix 5 in Supplementary File. Figure 5 also shows the effect of features on the prediction of this TP data sample by the RF model, which is interpreted based on the Shapley method.
5. Discussion
AKI is a serious complication of cardiac surgery that can occur at a rate of 1 to 30%. Of these, AKI, which requires kidney replacement therapy, has an incidence of about 1 to 5% (22). Perioperative AKI is independently associated with an increase in short-term morbidity, treatment costs, and long-term mortality (23). In cardiac surgery patients, postoperative AKI is associated with an increase in ICU admission and the length of hospital stay. Also, the development of kidney disease is accompanied by high rates of gastrointestinal bleeding, respiratory infection, and sepsis. In patients undergoing CABG on a cardiopulmonary bypass, the incidence of renal failure is between 1 to 15%, with a mortality rate of 19%. The incidence of AKI cases requiring dialysis after CABG is about 2%, with a 23 to 88% mortality rate (24).
Kidney dysfunction in cardiac surgery patients is usually multifactorial. The most common cause is acute tubular necrosis which results from hypoxic damage to nephrons in the medullary region of the kidney due to hypotension, hypovolemia, or dehydration. Other common risk factors include preoperative renal disease with an elevated level of creatinine, type 1 diabetes mellitus, over 65 years of age, major vascular surgery, more than 3 hours of cardiopulmonary bypass, recent exposure to nephrotoxic agents such as dyes radiocontrast, bile pigments, aminoglycoside antibiotics and nonsteroidal anti-inflammatory drugs (NSAIDs) (24).
Early detection of patients at high risk for AKI after cardiac surgery using risk scores can enable the anesthesiologist to apply early protective and therapeutic strategies to reduce AKI risk. Numerous risk scores have been developed to predict AKI, but there is still no guideline to recommend a predictive model (23). This study attempted to use ML techniques in predicting AKI after cardiac surgery. In this regard, ML methods were applied to this prediction. Evaluation of these methods was performed for two labels related to the first and seventh days after surgery, and the AUC of each method is reported in Tables 3 and 4. Based on the results, the best ML methods for classifying data are RF and XGBoost, with an AUC of around 0.8. RF and XGBoost are ensemble tree-based methods that usually show high efficiency in classification problems. Multiple imputations as a method of handling missing values have had a more significant impact on the output of the ML methods. However, there is not much effect on the RF and XGBoost results because of the ability of these methods to cope with the missing values. Also, using the combination of SMOTE and class weight methods for data oversampling gives the best results. In a study by Lee et al. (10), a similar attempt was made to evaluate machine learning methods to predict AKI for 2010 patients. In this study, the XGBoost method showed the highest performance in prediction.
Examining the interpretability of machine learning models is essential to ensure they work. In medical applications such as this study, the reliability of the model output is more critical than in other applications. What follows in the interpretability of models is how each of the features is involved in the prediction. The interpretability of models can be described in general and local terms. In general, we are looking to interpret the model based on the average of all the samples in the dataset. We have examined this in Figure 2 for both XGBoost and RF models. Based on this analysis, it can be generally said that the Cr (creatinine), CPB time (cardiopulmonary bypass time), BS (blood sugar), and Alb (albumin) features have the most significant impact on the predictions, respectively. However, the interpretability of a model in the local term examines how each feature affects a given sample. Therefore, to investigate the interpretability in the local term for black-box models such as XGBoost and RF, which have shown the best performance in the prediction, LIME (20) and Shapley (21) methods were used. Examining the results of these methods shows that for a particular patient predicted as a case with the risk of postoperative AKI, what features played a crucial role in this prediction?
Interpretation by the LIME method for a patient with postoperative AKI risk prediction shows that the Cr (creatinine) feature has the most significant positive effect on this prediction (see Figure 3A). Figure 4 compares the predicted values of the LIME local model and the main machine learning models for this patient. To trust the LIME interpretability, the predicted values for each primary model must be close to the corresponding values predicted by the LIME local model. In this plot, these predicted values for RF and XGBoost models are very close to each other, so it can be said that the interpretation obtained from the LIME method is reliable for this patient.
The Shapley method can also be used to interpret the machine learning models. Like LIME, this method examines models' interpretability based on individual samples. As shown in Figure 3B, the dominant feature with a positive role in prediction using RF and XGBoost is Cr (creatinine) for the same patient. In the force plot of Figure 5, the base value is 0.25. This value indicates the mean prediction of the test data. Features that force the prediction to move positively are displayed in red, and those that seek to predict negatively are shown in blue. Thus, the Cr (creatinine) feature largely makes the prediction positive.
Hence, the treatment team can first predict AKI incidence after cardiac surgery using patient information and then evaluate the prediction's outcome based on the model's interpretability for that patient. According to the importance of the determinant features, the treatment team can decide on the validity of the prediction.
One of the contributions of this study is the use of information from three different academic centers, which will help increase the validity of the results. Simultaneous use of retrospective and prospective data also improved the quality of existing data to provide high-quality and quantitative information suitable for machine learning models. Furthermore, most importantly, the use of interpretable machine learning methods makes it possible to assess the reliability of the methods appropriately. Limitations in the process of this study include inconsistent patient reports that increase the number of missing values. Also, the low incidence of AKI in stages 2 and 3 postoperatively among our patients led us to predict AKI regardless of its staging.
5.1. Conclusions
It can be concluded that using machine learning methods such as RF and XGBoost can predict AKI after cardiac surgery with promising efficiency. Interpretability of models can also help the treatment team ensure the validity of predictions. A reliable prediction of AKI incidence in patients can help the treatment team develop treatment strategies to prevent postoperative AKI. Preventing AKI can reduce treatment costs, length of hospital stay, and risk of death. In future work, we will optimize the parameters during surgery to reduce the risk of AKI in patients. In other words, we want to determine the anesthesia parameters during the surgery in such a way as to reduce the risk of AKI for the patient.