1. Background
Immunocompromised children with underlying diseases are at higher risk of hospital-acquired infection (1). These patients experience multiple hospitalization courses and receive antibiotics with likely effects on their gut microbiota composition. The richness of resistant variants of these bacteria in the intestine during hospitalization is mainly associated with extraintestinal infections and increased mortality risk.
The increased use of antibiotics has been reported in clinical settings during the COVID-19 pandemic. Among the resistant bacteria, carbapenem-resistant Enterobacteriaceae (CRE) is of primary concern. In these bacteria, Carbapenemases can hydrolyze all beta-lactam antibiotics, including carbapenems, as the last line of medication in severe nosocomial infections (2). Carbapenemase genes are encoded on transferable genetic elements, and their horizontal transfer makes them potentially responsible for outbreaks. The intestinal colonization of CRE is a risk factor for severe hospital infections and could serve as a reservoir for the transmission of these pathogens in clinical settings (2, 3). CRE is a major global concern leading to an increased mortality rate (4, 5). Carbapenemases are classified by their molecular structure (Ambler classification) in three groups (4, 6). Escherichia coli with NDM-1 is a human health-threatening pathogen, especially in vulnerable patients with underlying diseases, long-term hospitalization, and immune deficiencies (7, 8).
2. Objectives
The rates of fecal carriage of CRE vary across the world and patients with different health conditions. As a risk factor in HAIs, the fecal carriage of CRE is integral in vulnerable pediatric patients and accounts for comorbidity with SARS-CoV-2 and therapeutic regimens (9, 10). To find such a correlation in clinical settings, this study aimed to investigate the correlation between the intestinal carriages of CRE with the diversity of carbapenemase genes, SARS-CoV-infection status, and the types of antibiotics prescribed in immunocompromised children during hospitalization at oncology and hematology wards.
3. Methods
3.1. Patients and Samples
This study was conducted from August 21 to January 20 in Tehran, Iran. To this end, 102 stool samples were collected from immunocompromised pediatric patients admitted to the oncology and BMT wards of the hospital. The patients' information was obtained from questionnaires and hospital documents. All the stool samples were brought to the laboratory at least 2 hours after sampling. Selective cultures such as MacConkey agar (Pronadisa, CONDA, Spain) and biochemical tests such as TSI, IMViC, ureases, and lysine decarboxylase were used for Enterobacteriaceae characterization. Moreover, the COVID-19-related data of the patients were collected from the hospital documents.
3.2. Antibiotic Susceptibility Testing
The standard disk diffusion method was used to determine the sensitivity/resistance of Enterobacteriaceae isolates against imipenem (10 µg), meropenem (10 µg), amikacin (30 µg), gentamycin (10 µg), ceftazidime (30 µg), levofloxacin (5 µg), cefepime (30 µg), cefotaxime (30 µg), aztreonam (30 µg), and tigecycline (15 µg). Moreover, Muller Hinton agar (BD, USA), culturing 24 hours of isolates, diluting in 3 milliliters of physiological serum, and making standard McFarland were also used. The antimicrobial susceptibility was measured using the inhibition zone in the CLSI chart (11).
3.3. Detection of Carbapenemase Genes
Carbapenemase genes (blaOXA-48, blaNDM-1, blaKPC, and blaVIM) were assayed by PCR. Specific primers (SIGMA, Germany, and UK) were also used, and their sequences are presented in Table 1.
| Carbapenemase Genes | Base Pairs | Sequence (5 ˊ- 3') | TM°C | PCR Conditions | Reference |
|---|---|---|---|---|---|
| OXA-48 | 392 | F CCAAGCATTTTTACCCGCATCKACC | 65.5 | One cycle of initial denaturation at 95°C for 1 min, 30 cycles of denaturation at 95°C for 30 sec, Annealing at 55°C for 30 sec, Extension at 75°C for 1 min; and 1 cycle of final extension at 72 for 7 min. | (12) |
| R GYTTGACCATACGCTGRCTGCG | 61 | ||||
| NDM-1 | 129 | F CCCCGCCACACCAGTGACANCTC | 75.6 | One cycle of initial denaturation at 95°C for 1 min, 32 cycles of denaturation at 95°C for 30 sec, Annealing at 61°C for 30 sec, Extension at 7°C for 1 min; and 1 cycle of final extension at 72°C for 5 min. | (13) |
| RGTAGTGCTCAGTGTGGGCAT | 63 | ||||
| VIM | 390 | F GATGGTGTTTGGTCGCATA | 55.5 | One cycle of initial denaturation at 94°C for 10 min, 35 cycles of denaturation at 94°C for 30 sec, Annealing at 61°C for 40 sec, Extension at 72°C for 1 min; and 1 cycle of final extension at 72°C for 7 min. | (14) |
| R CGAATGCGCAGCACCAG | 57.2 | ||||
| KPC | 636 | F CTGTCTTGTCTCTCATGGCC | 60.5 | One cycle of initial denaturation at 94°C for 5 min, 32 cycles of denaturation at 94°C for 35 sec, Annealing at 62°C for 35 sec, Extension at 72°C for 32 sec; and 1 cycle of final extension at 72°C for 5 min. | (15) |
| R CCTCGCTGTGCTTGTCATCC | 62.5 |
4. Results
4.1. Patients
Among the participants, there were 54% (55/102) males and 46% (47/102) females, most of whom (43.13%, 44/102) were 1 - 5 years old. Moreover, 88% (90/102) of the patients had undergone treatment with broad-spectrum antibiotics such as carbapenems, cephalosporins, aminoglycosides, ciprofloxacin, and azithromycin. Table 2 presents the patients’ demographic information.
| Variables | No. (%) (N = 102) |
|---|---|
| Gender | |
| Male | 55 (54) |
| Female | 47(46) |
| Age (y) | |
| ≤ 5 | 44 (43.13) |
| 6 - 10 | 41 (40.19) |
| ≥ 11 | 17 (16.66) |
| Ward | |
| BMT | 23 (22.5) |
| Oncology | 79 (77.5) |
| Underlying disease | |
| AML | 31 (30.3) |
| ALL | 13 (12.7) |
| Neuroblastoma | 10 (9.8) |
| Others | 48 (47.05) |
| Antimicrobial prophylaxis | |
| Broad spectrum antibiotics | 90 (88) |
| Non-broad spectrum antibiotics | 12 (12) |
| COVID-19 status | |
| Positive | 51 (50) |
| Negative | 51 (50) |
| Death | |
| Yes | 24 (23.5) |
| No | 78 (76.5) |
Abbreviations: AML, acute myeloid leukemia; ALL, acute lymphocytic leukemia.
4.2. CRE Frequency in Intestinal Samples Isolated from Immunocompromised Children
During the study period, 102 clinical Enterobacteriaceae isolates were collected, the frequencies of which was as follows: Escherichia spp. (59/102, 57.8%), Klebsiella spp. (34/102, 33.3%), Enterobacter spp. (5/102, 4.9%), Citrobacter spp. (2/102, 1.9%), and Serratia spp. (2/102, 1.9%). According to the antimicrobial susceptibility tests, the strains showed the highest and the lowest resistance rates against ceftazidime (74.5%, 76/102) and tigecycline (13.72%, 14/102), respectively. Table 3 shows antimicrobial-resistant patterns.
| Antibiotics | Escherichia spp. N = 59 | Klebsiella spp. N = 34 | Enterobacter spp. N = 5 | Citrobacter spp. N = 2 | Serratia spp. N = 2 |
|---|---|---|---|---|---|
| IMP (10 µg) | 16 (27.11) | 20 (58.82) | 2 (40) | 1 (50) | 2 (100) |
| MEM (10 µg) | 14 (23.72) | 20 (58.82) | 2 (40) | - | 2 (100) |
| AN (30 µg) | 11 (18.64) | 13 (38.23) | 2 (40) | - | 2 (100) |
| GM (10 µg) | 29 (49.15) | 19 (55.88) | 2 (40) | 1 (50) | 1 (50) |
| CAZ (30 µg) | 45 (76.27) | 25 (73.52) | 3 (60) | 1 (50) | 2 (100) |
| LVX (5 µg) | 25 (42.37) | 15 (44.11) | - | 1 (50) | 2 (100) |
| CTX (30 µg) | 45 (76.27) | 25 (73.52) | 2 (40) | 1 (50) | 2 (100) |
| CFM (30 µg) | 31 (52.54) | 19 (55.88) | 2 (40) | 1 (50) | 2 (100) |
| AZT (30 µg) | 41 (69.49) | 23 (67.64) | 3 (60) | 1 (50) | 2 (100) |
| TGC (15 µg) | 3 (5.08) | 8 (23.52) | 2 (40) | 1 (50) | - |
Abbreviations: IMP, imipenem; MEM, meropenem; AN, amikacin; GM, gentamycin; CAZ, ceftazidime; LVX, levofloxacin; CFM, cefepime; CTX, cefotaxime; AZT, aztreonam; TGC, tigecycline.
a Values are expressed as No. (%).
The highest rate resistance to antibiotics was recorded in Escherichia spp. to ceftazidime and cefotaxime (76.27%), Klebsiella spp. to ceftazidime and cefotaxime (73.5%), Enterobacter spp. to ceftazidime and aztreonam (60%), Citrobacter spp. to all antibiotics except for meropenem and amikacin (50%), and Serratia spp. to all antibiotics except for gentamycin and tigecycline (100%).
4.3. Frequency of Carbapenemase Genes in Intestinal CRE Isolated from Immunocompromised Children
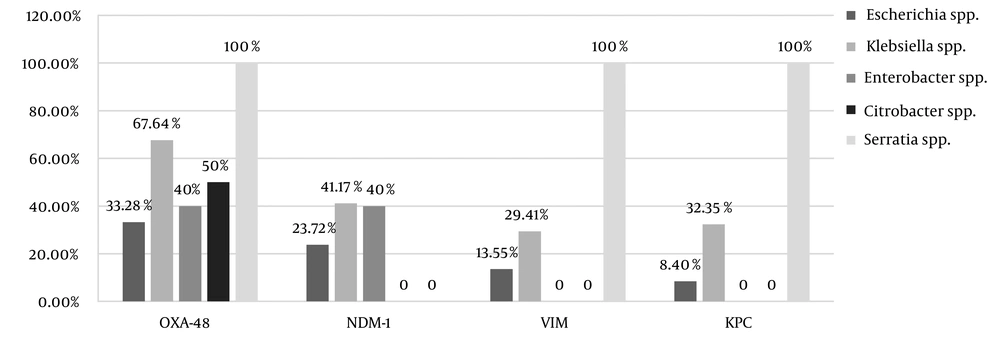
Carbapenem resistance was observed in 54% (55/102) of the strains, the frequencies of which was as follows: Escherichia spp. (42.37%, 25/59); Klebsiella spp. (73.52%, 25/34); Enterobacter spp. (40%, 2/5); Citrobacter spp. (50%, 1/2), and Serratia spp. (100%, 2/2). Out of the 55 CRE strains, the frequencies of carbapenemase were as follows blaOXA-48 (90.9%, 50/55), blaNDM-1 (54.54%, 30/55), blaVIM (36.36%, 20/55), and blaKPC (32.72%, 18/55). Figure 1 illustrates the frequency of the resistance genes among the CRE strains.
4.4. Intestinal CRE Carriage in Children with COVID-19 and Broad-Spectrum Antibiotic Usage
In this study, 50% (51/102) of the immunocompromised patients had laboratory-confirmed COVID-19. Of patients with intestinal CRE colonization54% (55/102), 65.45% (36/55) were COVID-19 positive after admission in the oncology and BMT wards. Out of patients with COVID-19 (50%, 51/102), 94.11% (48/51) underwent broad-spectrum antibiotics treatment. Table 4 presents the status of CRE carriers, COVID-19, and broad-spectrum antibiotic usage.
| CRE and Antibiotic Status | COVID-19 Positive; N = 51 | COVID-19 Negative; N = 51 | P-Value |
|---|---|---|---|
| CRE carriers and consumed broad-spectrum antibiotics | 29 (56.86) | 15 (29.41) | 0.027 |
| CRE carriers and not-consumed broad-spectrum antibiotics | 2 (3.9) | 4 (7.84) | |
| Not CRE carriers and consumed broad-spectrum antibiotics | 19 (37.25) | 27 (52.94) | |
| Not CRE carriers and not-consumed broad-spectrum antibiotics | 1 (1.9) | 5 (9.8) |
a Values are expressed as No. (%).
5. Discussion
The rate of hospital admissions in both infectious disease and pediatric intensive care unit (PICU) wards has been increasing because of the COVID-19pandemic. This had enhanced the likelihood of COVID-19 diffusion among other hospital wards (16). The description of broad-spectrum antibiotics such as carbapenems in these patients is associated with some side-effects, including the multiplication of carbapenemase genes, and the admission in different wards can also lead to the CRE outbreaks (17). The fecal carriage and intestinal colonization of CRE can raise the risks of secondary infections and increase mortality rates (18). CRE has revealed different drug resistance mechanisms, making the early detection and control of the CRE infections challenging (19).
Accordingly, we focused on the correlation between the intestinal carriage of CRE and the possible effects of carbapenemase genes on immunocompromised children during the COVID-19 pandemic.
The findings revealed that 65.45% of the immunocompromised pediatrics with the intestinal carriage of CRE were COVID-19 positive after hospitalization. Most of the patients were males (54%), and AML was the most frequent underlying disease among the patients. COVID-19 played an integral role in hospital prolongation and increasing death rates in intestinal CRE carriers.
Another study was conducted in the UK in 2022 to examine COVID-19 in immunocompromised children, and the findings indicated no mortality and the low risk of severe infections (1). The differences between this study and the present study might have underestimated the pivotal role of antibiotics administration and its association with CRE carriage and COVID-19.
Escherichia spp. was the most frequent strains isolated from immunocompromised children (57.8%) and also the most frequent carbapenem-resistant strain isolated from the intestinal CRE carriage of COVID-19 positive patients (68%). The second most frequent strain was Klebsiella spp. (33.3%) in 60% of COVID-19 positive CRE carriages. In a study carried out in 2021 in the USA, the frequency of CRE strains isolated from confirmed COVID-19 patients was 41.9%, and the most frequent species was Klebsiella pneumonia (90.3%) (17). The differences between the percentages of the CRE isolates could be attributed to differences in the samples as the target samples in the former study were feces; however, the latter one worked on blood and respiratory samples. Moreover, the frequency of Escherichia spp. was higher than the other kinds of samples.
Among intestinal CRE isolated from the intestinal track of the immunocompromised children, OXA-48 had the highest prevalence in this study. The first case of blaOXA-48 detection in Iran was reported by Azimi et al., who reported blaOXA-48 in 96% of imipenem-resistant K. pneumonia isolated from the immunodeficient patients (20, 21). According to a systematic review by Nasiri et al., blaOXA48 gene was the most common cause of carbapenem resistance in K. pneumoniae and E. coli in Iran (22). Pérez-Blanco et al. founded the higher frequency of OXA-48 in K. pneumoniae compared to other species, including K. pneumonia (93.6%), Escherichia coli (2.3%), Enterobacter spp. (1.7%), both K. pneumoniae and E. coli (0.5%), and Raoultella spp. or Citrobacter spp. (1.8%) (23). The high prevalence of OXA-48 among CRE in many studies can prove that carbapenem strains harboring OXA-48 has been boosting in Iran and becoming the endemic carbapenemase, thereby highlighting the significance of the strength detection and prevention control of CRE (22).
The detection of NDM-1 in K. pneumonia was first reported in 2013 in Iran (24). According to a study in 2019, NDM-1 was detected among 100% of the CRE isolates in Brazil, suggesting the high prevalence of NDM-1 in this country (25). In our study, NDM-1 was detected in 54.54% of the CRE strains. According to the data about harboring NDM-1 plasmid, a carrier of several other resistant genes, we can conclude that the spread of this challenging carbapenemase is in a medium range in Iran. On the other hand, the findings of this study suggest that NDM-1 has been increasing across the country in recent years. The high prevalence of NDM-1 in bacteria in hospitals can be a major challenge in treating and controlling infectious diseases (24).
The emergence of VIM-producing K. pneumoniae in Iran is also a concern. VIM-producing bacteria constitute the prevalent multidrug-resistant population of K. pneumoniae in Iran (26). In a study in 2019, 73% of Enterobacteriaceae isolates, especially Enterobacter spp., collected from clinical and environmental samples, were VIM carriers (27). In the present study, we reported 36.36% of VIM carriage among CRE.
The correlation between CRE intestinal carriage with COVID-19 and administrational antibiotics was significant in this study. To the best of our knowledge, no similar study has addressed this issue.
Note that most of the patients who were CRE colonized had previous hospital admission and had consumed broad-spectrum antibiotics, especially ciprofloxacin and carbapenems. In the present study, the most frequently used prophylaxis antibiotics in these two wards were meropenem, vancomycin, and amikacin. It can be assumed that immune deficiency in patients admitted to the BMT and oncology wards is a remarkable risk factor for upcoming infections such as COVID-19 and that the circle of these infections finally would end in prolonged hospitalization period, which itself is one of the most dangerous hospital-acquired infection-inducing risk factors. Moreover, this process could lead to the spread of the CRE strains by the carriers during admission in different hospital wards. In this study, there was no control group of inpatient children; hence, further studies are required to detect the relationship between CRE colonization with previous hospital admissions, prophylaxis antibiotics diet, and COVID-19.
5.1. Conclusion
Regardless of the COVID-19 pandemic, prolonged hospitalization and antibiotic prescription are main risk factors associated with the CRE intestinal colonization in immunocompromised children. The high prevalence of blaOXA-48 in these isolates, especially in Escherichia spp., can raise therapeutic challenges and concerns about systemic infections in these patients. The intestinal carriage of KPC could increase the risk of secondary infection treatment failure in immunocompromised and COVID-19 positive patients. The co-occurrence of NDM-1, OXA-48, and VIM-producing Enterobacteriaceae is a concern and need further consideration to control their spread rate and highlight the significance of maximizing the infection control measures.