1. Background
Diabetic foot infections (DFIs) are a frequent clinical problem. Diabetic extremity ulcers develop in almost 15% of people with diabetes and are a leading cause of hospital admission and amputation among this group of patients, while 85% of major leg amputations begin with a foot ulcer in the diabetic population (1, 2). Diabetic foot ulcers (DFU) are more prone to different bacterial infections that spread quickly, leading to irreversible tissue injury. Many microorganisms, alone or as part of a polymicrobial infection, can cause DFIs, of which non-spore forming, gram-positive cocci such as Enterococci are the most common bacteria (3).
Foot infections in individuals with diabetes are an increasingly common public health problem and are associated with mortality and morbidity. DFIs followed by amputations contribute significantly not only to the morbidity among diabetic persons, but are also associated with severe clinical dumps and remarkably increased mortality rates (2, 4).
Previous studies point toward gram-positive cocci, such as members of the Enterococcus genus, as the most common pathogens in DFI samples, contributing to the persistence or severity of the disease and leading to higher morbidity and mortality rates (5).
Over the past few decades, a major problem in treating diabetic foot infections has been the increased isolation rate of antimicrobial-resistant bacteria, especially methicillin-resistant Staphylococcus aureus (MRSA) and, to a lesser degree, glycopeptides-intermediate S. aureus (GISA), vancomycin-resistant Enterococci (VRE), extended-spectrum β-lactamases (ESBLs), or carbapenamase-producing, gram-negative bacilli. The isolation rates of these multidrug-resistant (MDR-resistance to at least three unrelated antibiotic classes) bacteria vary widely according to geographic areas and treatment centers. The potential presence of such MDR isolates emphasizes the importance of ideal sample collection for bacterial culture and antimicrobial susceptibility testing for the infected DFIs, as well as preventing the excessive antimicrobial use that drives this resistance (6, 7). The prevalence of Enterococci in DFIs has been increasing. This increased incidence might be due to prior antibiotic use (8, 9).
2. Objectives
The aims of this study were the isolation and characterization of Enterococci from diabetic patients with foot ulceration, confirmation of Enterococci and their genus, determination of the susceptibility profile of the isolates, and survey of the cross-resistance among Enterococcus isolates.
3. Methods
3.1. Patients
A total of 86 diabetic patients with foot ulcers were surveyed during 2012 - 2014 at Nemazee Hospital in Shiraz, Iran. The study population was defined as the total number of patients with type 1 diabetes mellitus (DM) and type 2 DM with foot ulcers (i.e., suspected infection according to physician decision) at initial visit and admission to hospital. Information regarding patients’ demographic and clinical features was gathered.
3.2. Isolation, Characterization, and Confirmation of Isolates
Swabs were collected from diabetic ulcers that were macroscopically examined and classified (10). Swabbing was performed on sloughy or inflamed tissue, as bacteria tend to present in greater number in such areas. From each patient, two swabs (for isolation of bacteria and wet mount microscopy) were obtained. The sterile cotton-tipped swab was moistened with sterile normal saline before sample collection. One of the prepared swabs was used for the isolation of bacteria. The other was used for wet mount microscopy. For the isolation of bacteria from collected specimens, the microbiological media used were blood, chocolate, and MacConkey agar, which were incubated for 16 - 18 hours at 35°C. Representative bacterial colonies recovered after incubation were sub-cultured on blood agar plates, which were incubated at 35°C in the presence of 5% CO2 for 24 hours (5).
The cultural characteristics of bacterial isolates on the agar plates were examined. The characterization and identification methods for the bacteria were carried out by standard procedures. Gram staining and cell morphology from air-dried, heat fixed smears were performed. The motility of the isolates was surveyed by hanging drop (HD) technique. Bacterial colonies were further characterized by different biochemical diagnostic tests, including catalase test, oxidase reaction, growth on bile esculine (BE) agar, growth in the presence of 6.5% NaCl, growth at 45°C, motility, pyrrolidonyl arylamidase (PYR), yellow pigment, and arginine dihydrolase (ADH). Final identification of different species of the Enterococcus genus was conducted by sugar fermentation tests (i.e., glucose, mannitol, sorbose, arabinose, sorbitol, raffinose, sucrose, and pyruvate).
3.3. Susceptibility Testing
The bacterial isolates were subjected to antibiotic sensitivity testing on Muller-Hinton agar by the Kirbey-Bauer standard disc diffusion method (11). All inoculated plates were incubated for 16 - 18 hours in ambient air incubators at 35°C, and the results were recorded by measuring the zone of inhibition, according to the Clinical and Laboratory Standards Institute (11) protocols. Susceptibility of Enterococcus isolates was tested for clindamycin (CD, 2 μg), erythromycin (E, 15 μg), linezolid (LZD, 30 μg), penicillin G (PG, 10 μg), co-trimoxazole (TS, 1.25/23.75μg), rifampicin (RP, 5 μg), oxacillin (OX, 1 μg), ciprofloxacin (CIP, 5 μg), gentamicin 120 (GMH, 120 μg), ceftriaxone (CRO, 30 μg), cefixime (CFM, 5 μg), vancomycin (VA, 30 μg), gentamicin (GM, 10 μg), ampicillin (Ap, 10 μg), imipenem (IMP, 10 μg), cefepime (CPM, 30 μg), cefoxitin (FOX, 30 μg), chloramphenicol (C, 30 μg), cefotaxime (CTX, 30 μg), ceftizoxime (CZX, 30 μg), azithromycin (ATH, 15 μg). E. faecalis ATCC 29212 was used as quality control.
3.4. Detection of Vancomycin-Resistant Enterococcus (VRE) by BHI Agar Screen Plate
All Enterococcus isolates were examined for reduced vancomycin susceptibility by agar incorporation. Ten μL of a 0.5 McFarland bacterial suspension (final concentration = 106 CFU/mL) was spotted on the brain heart infusion (BHI) agar (Merck, Germany), containing 6μg/mL vancomycin, allowed to air dry for almost five minutes, and incubated at 35°C (11). Culture plates were examined at 24 and 48 hours of incubation for any discernible growth.
4. Results
In the current study, a total of 86 diabetic patients were investigated. Enterococcus spp. were isolated from 34 (39.5%) patients. The cases consisted of 20 males (59%) and 14 females (41%), aged between 28 - 85 years. In this study, 34 strains of Enterococcus were isolated from diabetic foot patients referred to Nemazee Hospital (Shiraz, Iran). Diabetic foot patients’ weights ranged from 45 to 100 kg, with the maximum number of cases in the weight group of more than 70 kg (n = 20, 59%) (Table 1). Nine (26.4%) patients had higher than general education. Twenty-five (73.5%) patients received antibiotic treatment on admission (24 patients received clindamycin and ciprofloxacin, 1 patient received cephalexin). Fifty (44.1%) patients had random blood sugar ranging between 130 - 300, and 19 (55.9%) had blood sugar of 300 - 450. Of the 34 patients, 15 (44.1%) had type 1 diabetes, 19 (55.9%) had type 2 diabetes (Table 2), and one patient (2.9%) died.
| Category | Results (%) |
|---|---|
| Gender | |
| Male | 20 (58.8) |
| Female | 14 (41.2) |
| Age, years | 28 - 85 |
| Weight, kg | 45 - 100 |
| < 75 | 1 (2.9) |
| 50 - 75 | 13 (38.3) |
| > 70 | 20 (58.8) |
| Education | |
| Higher than general education | 9 (26.4) |
| General education | 10 (29.4) |
| Lower than general education | 15 (44.2) |
Demographic Features of the Diabetic Foot Patients Infected with Enterococcus spp
| Category | Results (%) |
|---|---|
| Antibiotic treatment | |
| Clindamycin and ciprofloxacin | 24 (70.6) |
| Cephalexin | 1 (2.9) |
| No antibiotic | 9 (26.5) |
| Diabetic type | |
| Type 1 | 19 (55.8) |
| Type 2 | 15 (44.2) |
| Random blood sugar range | |
| 130 - 300 | 15 (44.2) |
| 300 - 450 | 19 (55.8) |
| Ulcer size | |
| > 4 mm | 24 (70.6) |
| < 4 mm | 10 (29.4) |
| Ulcer type | |
| Osteomyelitis | 6 (17.6) |
| Gangrene | 7 (20.6) |
| Cellulitis | 7 (20.6) |
| Neuroischemic ulcer | 6 (17.6) |
| Ischemic ulcer | 4 (11.8) |
| Abscess | 4 (11.8) |
| Amputation | 14 (41.1) |
| Risk factor | |
| Vascular diseases | 16 (47) |
| Hypertension | 15 (44.1) |
| Retinopathy | 6 (17.6) |
| Osteomyelitis | 6 (17.6) |
| Neuropathy | 5 (14.7) |
| Nephropathy | 5 (14.7) |
| Isolated Enterococcus spp. | |
| E. faecalis | 17 (50) |
| E. faecium | 16 (47) |
| E. mundetti | 1 (2.9) |
| Vancomycin-resistant Enterococcus | 5 (14.7) |
Distribution of the Diabetic Patients Infected with Enterococcus spp. in Terms of the Antibiotic Treatment, Diabetic Type, Blood Sugar, Ulcer Size, Ulcer Type, Amputation, and Risk Factor (n = 34)
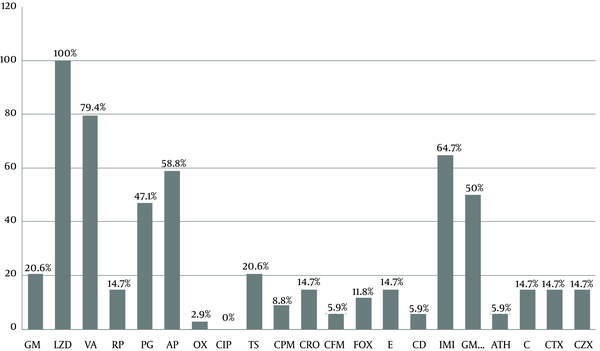
Enterococcus faecalis was the most common isolated Enterococcus spp. (50%). According to the in vitro antibiotic susceptibility testing, linezolid was the most effective antibiotic against Enterococcus isolates (all isolates [100%] were sensitive) and ciprofloxacin was the least effective drug (all isolates [100%] were resistant) (Figure 1). Susceptibility rates for vancomycin, imipenem, ampicillin, and gentamicin 120 were 79.4%, 64.7%, 58.8%, and 50%, respectively, and the susceptibility rate for erythromycin, rifampicin, ceftriaxone, chloramphenicol, cefotaxime, and ceftizoxime was 14.7% (Figure 1). Analysis of cross-resistance results revealed that more than 85% of the isolates were resistant to macrolides and 24 (70.5%) to chloramphenicol, clindamycin, erythromycin, azithromycin, and gentamicin (Table 3).
| Pattern | Antibiotic Resistance Patterns | Number (%) |
|---|---|---|
| A | E, ATH | 29 (85.3) |
| B | CPM, CTX, CZX, CFM, CRO | 28 (82.3) |
| C | E, ATH, CD, RP | 25 (73.5) |
| D | C, CD, E, ATH, GM | 24 (70.5) |
| E | GM, GM120 | 16 (47) |
| F | CIP, AP, GM, GM120 | 9 (26.5) |
| G | GM, GM 120, AP | 9 (26.5) |
| H | IMI, VA, CIP, AP | 6 (17.6) |
| I | VA, AP, GM | 5 (14.7) |
| J | VA, RP, IMI | 5 (14.7) |
| K | VA, AP, GM, GM120 | 4 (11.7) |
| L | VA, AP, GM, GM120, IMI | 4 (11.7) |
| M | CD, VA, RP, IMI, AP,GM, GM120 | 4 (11.7) |
Antibiotic Cross-Resistance Patterns of Enterococcus spp. Isolated from Diabetic Foot Patients (n = 34)z
5. Discussion
Most DFIs have a polymicrobial etiology, with enterococcal strains being part of the complex diabetic foot microbiota. Previous studies point toward the Enterococcus genus as one of the most common gram-positive pathogenic bacteria in DFI samples, contributing to the persistence or severity of the disease and leading to higher morbidity and mortality rates (5).
All DFI Enterococci present gelatinolytic, hemolytic, and biofilm forming (which contribute to the chronicity of infection) abilities. Since the screened virulence traits are considered among the most relevant for enterococcal pathogenicity mechanisms, often detected in clinical isolates and correlated with the persistence and severity of infection (5, 12). The choice of accurate antimicrobial depends on an accurate evaluation of sepsis severity, credible microbiologic data, and consideration of host factors, such as renal and vascular impairment (13). Lower extremity infections are a serious cause of morbidity and mortality in persons with diabetes mellitus (DM) (2). Microbiologically, diabetic foot infections are generally polymicrobial, but in this study, we focused on diabetic ulcers contaminated with Enterococcus. Enterococcus faecalis and Enterococcus faecium were the most common isolated Enterococcus species from diabetic foot infections (DFI) in the present study.
The results of antimicrobial susceptibility testing showed that linezolid is the most effective agent against Enterococcus spp. Third and fourth generation cephalosporins were ineffective against more than 82% of Enterococci isolates. According to CLSI recommendations, for Enterococcus spp., cephalosporins, aminoglycosides (except for high-level resistance screening), clindamycin, and trimethoprim-sulfamethoxazole may appear active in vitro, but are not effective clinically and should not be reported as susceptible (11).
Ciprofloxacin was the least effective drug against isolates; therefore, it should not be used empirically as a single agent. The data analysis indicates that antibiotics such as gentamicin, used extensively in the treatment of different infections caused by Enterococcus spp. in hospitals, were active only against about 20.6% of total Enterococcus species tested, but gentamicin 120 was effective against 50% of the isolates. In some studies, rifampicin exhibited good activity against Enterococcus species (14), but this was not the case in this study, in which only 18% of the isolates showed sensitivity. Approximately 85% of Enterococcus species were resistant to erythromycin, rifampicin, ceftriaxone, chloramphenicol, cefotaxime, and ceftizoxime, in contrast to studies of diabetic foot isolates in Saudi Arabia (15).
Given the alarming types of resistance (i.e., resistance to vancomycin) among Enterococcus spp. (16), our data showed that resistance to the vancomycin tested was found to be 20.6% among Enterococcus spp. As revealed, significant resistance to gentamicin 120 (50%) and imipenem (35.3%) were alarming. The analysis results of cross-resistance showed that 85.3% were resistant to macrolides. Four (11.4%) isolates were co-resistant to common antibiotics (including vancomycin, ampicillin, and gentamicin) used for the treatment of infections with Enterococcus.
Knowledge of the causative microorganisms (such as bacteria) in diabetic foot infections (DFI) and their antibimicrobial susceptibility profiles is essential for appropriate treatment and infection eradication. In patients with serious infections, the antibiotic therapy may have to be initiated empirically to prevent systemic invasion by infecting bacteria in a formerly debilitated patient while awaiting microbiology laboratory results (17).
In the present study, Enterococci were found in 39.5% of the patients, which is higher, compared to the report by Citron (39.5% versus 35.7%) (18). Our results for carbapenem (imipenem) resistance among Enterococcus spp. (35.3%) are not in agreement with some reports from Citron et al., which showed resistance to other carbapenems (ertapenem) to be 90% (18). Enterococcus spp. may show different response to members of the carbapenem class of antibiotics. Clinicians should consider the results of bacterial culture and susceptibility testing in the light of the clinical outcome of the infection for the empirical therapy regimen. Knowledge of the characteristics of infection, i.e., the type of bacteria commonly found and the clinical evidence of infection, can help in choosing an appropriate antibiotic, even if the culture reports are not available at the time of initiation of antibiotic therapy (19, 20).
In our study, 58.8%, 79.4%, and 47.1% of isolates were susceptible to ampicillin, vancomycin, and penicillin, respectively. In El-Tahawy (15), the Enterococci were fully sensitive to ampicillin and vancomycin, while 16% were resistant to penicillin. This may be due to factors such as the differences in treatment regimens used for infected patients in healthcare settings. Also, the majority of antibiotics are used in regional agricultural settings and food-producing animals; therefore, different resistance patterns can emerge and spread globally. In the current study, isolates with resistance to quinolones were seen, consistent with what was reported by Goldstein and colleagues (21).
It should also be noted that, in many cases, antimicrobial resistance is transmitted to the human population, hospitalized patients, and the hospital environment through other sources including animals, plant-based foods, fish, poultry, and other industries in which antibiotics are used for different purposes and may lead to emerging resistant strains of bacteria (22-25). The multidrug resistant (MDR) status attributed to the majority of the Enterococci continues to be highly relevant, especially in chronic severe Enterococci infections such as DFIs, since antimicrobial resistance often results in treatment failure. The presence of MDR diabetic foot ulcer Enterococci is of major importance, also due to the possibility of transmitting those MDRs to other bacteria sharing the same ecological niche, highly impairing the implementation of successful antibiotic therapy (5).
5.1. Conclusion
Isolation, identification, and antimicrobial susceptibility of pathogens can be helpful in optimizing antimicrobial use. Because these bacteria are often resistant to the prescribed antimicrobials, the physicians must decide if the superiority of clinical and laboratory evidence suggests they are invasive pathogens that require targeted antibiotic therapy. If the patient with DFI has not adequately responded to the empirical therapy regimen, treatment should be broadened to include all recovered microorganisms.