1. Background
Extended-spectrum β-lactamases (ESBLs) are plasmid-mediated enzymes in bacteria that cause resistance to a wide range of β-lactam antibiotics. The CTX-M family has become the most prevalent type of ESBL worldwide, and has been detected mainly in Escherichia coli and Klebsiella species (1). CTX-M β-lactamases have been clustered into six groups based on their amino acid sequence similarities: CTX-M-1, -2, -8, -9, -25, and -45 (2). These enzymes hydrolyze cefotaxime better than ceftazidime, although CTX-M-15 (belonging to group 1) has been reported to have good activity against ceftazidime (3). Rapid emergence and dissemination of the CTX-M-15 enzyme has been widely reported, and it is currently the most widely distributed ESBL genotype in most parts of the world (4-6).
Large nosocomial outbreaks caused by CTX-M-producing Klebsiella pneumoniae have been reported worldwide (7, 8). Intensive use of extended-spectrum cephalosporins and cross- transmission have been associated with the emergence and dissemination of ESBL-producing K. pneumoniae (1). The ESBL genes responsible for resistance are mainly carried on mobile genetic elements, which can readily spread among bacteria (9). Class 1 integrons and transferable elements like conjugative plasmids or insertion sequences (IS) (e.g., ISEcp1) play an important role in the dissemination of antimicrobial resistance genes between species or genera (10). It has been reported that blaCTX-M genes are carried on the plasmids ranging from 7 to 430 kb, mainly on large transmissible plasmids (11). Due to the role of plasmids in the horizontal transfer of antimicrobial resistance genes, much attention has been paid to the identification and classification of bacterial plasmids. Carattoli et al. demonstrated a polymerase chain reaction (PCR)-based replicon typing (PBRT) scheme to detect 18 plasmid replicons commonly found among the Enterobacteriaceae (12). The blaCTX-M types have been shown to be associated with specific plasmid replicons, for example, blaCTX-M-9 to replicon HI2, blaCTX-M-15 to replicons FII, FIA, and FIB, and blaCTX-M-3 to replicon N. In addition, different genetic elements have been shown to be involved in the mobilization of blaCTX-M, such as insertion sequence ISEcp1, which is also responsible for the expression of genes encoding β-lactamase (11). The epidemiology of CTX-M-producing strains is complex, involving the dissemination of strains and the dissemination of mobile genetic elements bearing blaCTX-M genes. Outbreaks of clonal strains have been reported worldwide (2, 8). Genotypic methods are required to demonstrate clonal dissemination between K. pneumoniae isolates. Pulsed-field gel electrophoresis (PFGE) is the current gold standard method for bacterial typing (13).
2. Objectives
In our previous study, we investigated the prevalence of the blaCTX-M-1 group in 31 clinical K. pneumoniae isolates from samples of children between May and December 2011. Our results showed the presence of the blaCTX-M-1 group in 17 of the isolates (14). The present study was designed to characterize these 17 CTX-M-1 group-producing K. pneumoniae isolates and to investigate the epidemiological relationship between them using PFGE.
3. Methods
3.1. Bacterial Isolates
In our previous cross-sectional study (14), 31 K. pneumoniae isolates were identified from samples of children aged 0 - 12 years admitted to three hospitals in Tehran (Loghman-e-Hakim in the southwest, Imam Khomeini in the central region, and Milad in the northwest) between May and December 2011. Antibiotic susceptibility of the isolates was determined using the disk diffusion method. The presence of the blaCTX-M-1 group and class 1, 2, and 3 integrons was investigated through the use of PCR. The results showed a high level of antibiotic resistance among the isolates: 17 isolates were simultaneously resistant to amoxicillin-clavulanic acid, cefotaxime, ceftriaxone, aztreonam, and ceftazidime. All were susceptible to imipenem. Class 1 integron was detected in 8 isolates, whereas classes 2 and 3 were not detected. The blaCTX-M-1 group was detected in 17 isolates. These 17 CTX-M-1 group-producing K. pneumoniae isolates were included in the present study for further characterization and determination of clonal relationships.
3.2. Characterization of the Genetic Support of the blaCTX-M-1 Group
The genetic environment surrounding the blaCTX-M-1 group was investigated by PCR mapping and sequencing of the amplicons (15). Specific primers for the insertion sequences ISEcp1 and IS26 (located upstream of blaCTX-M-1) and for the open reading frame orf477 (located downstream) were combined with blaCTX-M-1-specific primers (Table 1). Amplicons were then sequenced and their sequence similarity to other genes was searched using the basic local alignment search tool (BLAST) program.
| Target | Primer Name | Primer Sequence (5′ - 3′) | Amplification Conditions | Reference |
|---|---|---|---|---|
| blaCTX-M-1 group | M1F | GGTTAAAAAATCACTGCGTC | 1 cycle of 5 minutes at 94°C; 35 cycles of 1 minute at 94°C, 1 min at 54°C, 1 minute at 72°C; 1 cycle of 7 minutes at 72°C | (14) |
| M1R | TTGGTGACGATTTTAGCCGC | |||
| ISEcp1 | ISEcp1U1 | AAAAATGATTGAAAGGTGGT | 1 cycle of 5 minutes at 94°C; 35 cycles of 1 minute at 94°C, 1 minute at 52°C, 1 minute at 72°C; 1 cycle of 7 minutes at 72°C | (15) |
| IS26 | tnpA IS26 | AGCGGTAAATCGTGGAGTGA | 1 cycle of 5 minutes at 94°C; 35 cycles of 1 minute at 94°C, 1 minute at 58°C, 90s at 72°C; 1 cycle of 7 minutes at 72°C | (16) |
| Orf477 | ORF477R | CCAGGAACCACGGAGCTTAT | 1 cycle of 5 minutes at 94°C; 35 cycles of 1 minute at 94°C, 1 minute at 54°C, 1 minute at 72°C; 1 cycle of 7 minute at 72°C | The present study |
3.3. Plasmid Characterization
Plasmids were classified according to incompatibility groups by PCR using a previously described PBRT method (12). PBRT (five multiplex- and three simplex-PCRs) was carried out on total DNA using 18 primer pairs that recognize FIA, FIB, FII, FIC, Frep, HI1, HI2, I1, P, A/C, X, L/M, N, T, K, Y, B/O, and W replicons, which are the major plasmid incompatibility groups circulating among the Enterobacteriaceae (12). Positive controls were provided by Dr. Carattoli (Istituto Superiore di Sanita, Italy). The PCR conditions used were as follows: 5 minutes at 94°C; followed by 35 cycles of 1 minute at 94°C, 30 seconds at 60°C, and 1 minute at 72°C; and a final extension of 7 minutes at 72°C.
3.4. Conjugation and PCR Amplification of the blaCTX-M-1 Group From Transconjugants
To determine whether the blaCTX-M-1 group gene was carried on a conjugative plasmid, conjugation experiments were carried out using a broth-mating assay (17) with K. pneumoniae strains as donors and rifampicin-resistant E. coli A15R- (provided by Dr. Bedenic, Croatia) as the recipient strain. Cultures of recipient and donor strains grown in Luria-Bertani broth (Scharlau, Spain) were combined in a ratio of 1: 10 (donor to recipient) and incubated at 37°C for 16 hours. Samples (0.1 mL) of this mixture were spread onto the surfaces of Mueller-Hinton agar plates containing 300 µg/mL rifampicin (for the inhibition of donor strain growth) plus 3 µg/mL cefotaxime (to prevent the growth of E. coli recipients lacking blaCTX-M-1). Samples from donor and recipient strain were used as controls. The transconjugants growing on the selection plates were examined for acquisition of the blaCTX-M-1 group gene using PCR (Table 1).
3.5. Molecular Strain Typing
PFGE of XbaI-digested genomic DNA was performed according to the pulse net protocol from the centers for disease control and prevention (Atlanta, USA) (18) with minor modifications. Agarose plugs were digested with 10 U of restriction endonuclease XbaI (Jena Bioscience, Germany) at 37°C for 6 hours. Electrophoresis was performed on a 1% low melting agarose gel (Sigma, USA) using a CHEF-DR II apparatus (Bio-Rad, USA) under the following conditions: initial pulse time for 2.2 seconds and final time for 54.2 seconds at 6 V for 20 hours.
Banding patterns were compared using Gelcompar II software, version 4.0 (Applied Maths, Belgium). Salmonella Braenderup H9812 was used as the reference strain. A dendrogram was generated using the band-based Dice similarity coefficient and the unweighted pair group method with arithmetic mean (UPGMA). Clusters were created with a 1.5% band tolerance. The discriminatory power of the PFGE was assessed using Simpson's diversity index (19) via a free online tool, Comparing Partitions, available at http://Darwin.phyloviz.net/ComparingPartitions.
4. Results
4.1. Characterization of the blaCTX-M-1 Group Gene
Sequence analysis identified all blaCTX-M-1 group genes as blaCTX-M-15. The GenBank accession number for the blaCTX-M-15 gene is KC131462. PCR amplification using the specific primers reported in Table 1 indicated the presence of the insertion sequence ISEcp1 upstream of the blaCTX-M-15 gene in all the strains and the promoter region (-10 and -35) associated with this element. A 48-bp region was found between the right inverted repeat of ISEcp1 and the start codon of the blaCTX-M-15 gene. The orf477 gene was detected downstream of the blaCTX-M-15 gene in all strains, whereas IS26 was not found.
4.2. Plasmid Analysis
A total of eight different replicons, dominated by L/M, were detected in all isolates using PBRT (Table 2). Of the 17 strains tested, one strain was devoid of any replicons. Of the 18 replicons, 10 were not found, including the I1, HI2, X, FIA, FIB, W, A/C, P, Y, and T replicons. The IncL/M replicon was the most commonly detected replicon type (14/17 isolates, 82.3%), followed by HI1, FIC (4/17, each), IncK, FIIs (3/17, each), IncB/O (2/17), and Frep, N (1/17, each). In total, 8 of the 17 isolates (47%) contained multiple replicons. L/M + FIC were the most frequently recognized replicons in combination (4/17, 23.5%). The results of the conjugation experiment showed that blaCTX-M-15 was carried on conjugative plasmids in all of the K. pneumoniae strains. The PCR analysis performed on the transconjugants revealed the acquisition of the blaCTX-M-15 gene.
| Pulsotypes | Number of Isolate (s) | Replicon Types | Antimicrobial Resistancea | Class 1 Integronb |
|---|---|---|---|---|
| P46 | 1 | L/M | CTX, CRO, CAZ, AUG, ATM, TN, GM, CPM, AK | - |
| 1 | + | |||
| 1 | HI1, L/M | - | ||
| 3 | L/M | - | ||
| P54 | 1 | L/M, FIC, Frep, FIIs | CTX, CRO, CAZ, AUG, ATM, TN, TS, GM, CPM, AK | + |
| P50 | 1 | L/M | CTX, CRO, CAZ, AUG, ATM, TN, GM, CPM, AK | + |
| P5 | 1 | HI1, L/M, FIC | CTX, CRO, CAZ, AUG, ATM, TN, GM, CPM, AK | - |
| P49 | 1 | L/M | CTX, CRO, CAZ, AUG, ATM, TN, GM, CPM, AK | - |
| P47 | 1 | HI1 | CTX, CRO, CAZ, AUG, ATM, TN, GM, CPM, AK | - |
| P43 | 1 | HI1, N | CTX, CRO, CAZ, AUG, ATM, T, TS, CPM, FOX | + |
| P37 | 1 | L/M, B/O | CTX, CRO, CAZ, AUG, ATM, TN, GM, CPM, AK | - |
| P2 | 1 | L/M, FIC, B/O, K, FIIs | CTX, CRO, CAZ, AUG, ATM, TN, T, GM, CPM, AK | + |
| P14 | 1 | NT | CTX, CRO, CAZ, AUG, ATM, TN, CPM, AK | + |
| 1 | L/M,K | CTX, CRO, CAZ, AUG, ATM, TN, AK | + | |
| P1 | 1 | L/M,FIC,K,FIIs | CTX, CRO, CAZ, AUG, ATM, TN, GM, CPM, AK | + |
Abbreviations: ATM, aztreonam; CTX, cefotaxime; CRO, ceftriaxone; CAZ, ceftazidime; CPM, cefepime; FOX, cefoxitin; AUG, amoxicillin-clavulanic acid; T, tetracycline; TS, trimethoprim-sulphamethoxazole; TN, tobramycin; GM, gentamicin; AK, amikacin.
aThe resistance profiles excluded antimicrobial agents that exhibited intermediate resistance.
b+, positive; -, negative.
4.3. Molecular Typing of Isolates by PFGE
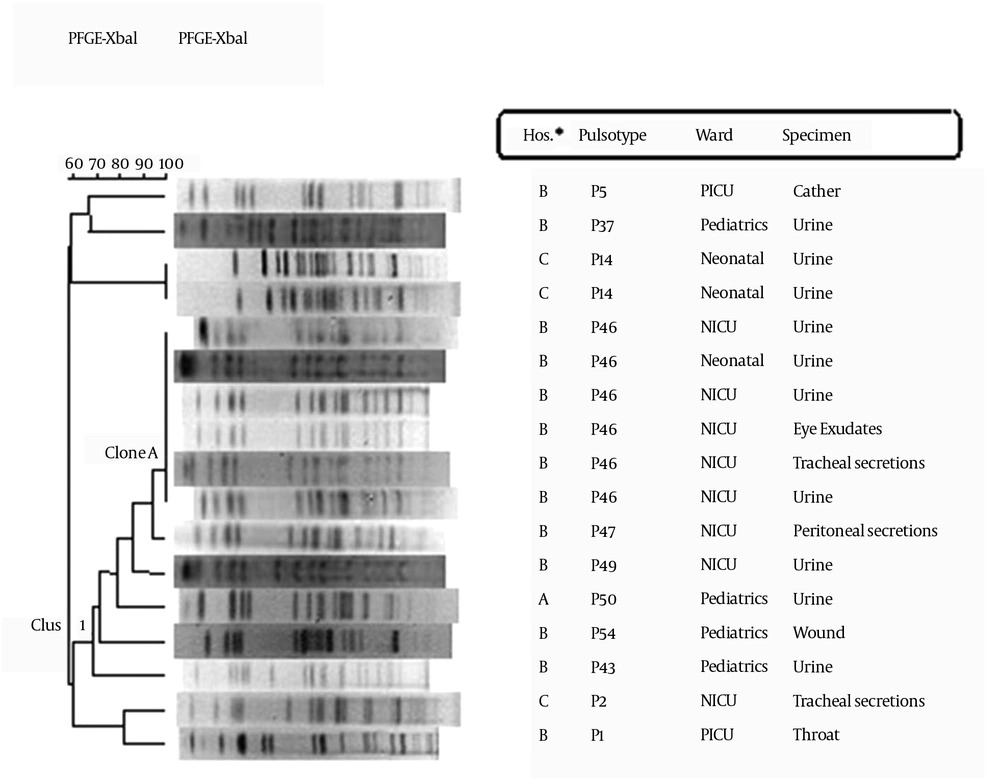
The PFGE analysis of the 17 blaCTX-M-15 positive isolates, identified 11 distinct PFGE profiles (pulsotypes). P46 was the most commonly found pulsotype [6/17, 35.2% of isolates]. Figure 1 shows the clustering of PFGE data. One major cluster (cluster I) was identified among the PFGE types. This cluster was composed of 11 isolates collected from hospitals A (Loghman) and B (Milad), and 6 pulsotypes were detected in the cluster (P43, P46, P47, P49, P50, and P54). Seven isolates from this cluster were found in samples collected from the neonatal intensive care unit (NICU), 3 isolates in samples from the Pediatric ward, and 1 isolate in samples from the Neonatal unit. Similar PFGE patterns (similarity above 80%) were observed among isolates in samples from the NICU and neonatal ward. One major clone (isolates with indistinguishable PFGE profiles) that was composed of 6 isolates (clone A) was identified in samples from the NICU and neonatal ward of hospital B (Milad). The discriminatory power of PFGE (Simpson's diversity index) was 0.882 with a 95% confidence interval (0.750 - 1.000).
The antibiotic resistance, presence of class 1 integron, and replicon content of each pulsotype are presented in Table 2. As shown, most isolates with the same PFGE type had similar antimicrobial resistance profiles. For example, six isolates with pulsotype P46 showed the same resistance profile.
5. Discussion
This study was conducted to characterize the 17 CTX-M-1 group ESBL-producing K. pneumoniae strains isolated in our previous study (14). Thirty-one K. pneumoniae isolates were identified in samples collected from children aged 0 - 12 years admitted to three hospitals in Tehran between May and December 2011. The blaCTX-M-1 group was detected in 17 isolates (14). These 17 CTX-M-1 group-producing isolates were included in the present study for further characterization and determination of clonal relationships.
All of the CTX-M-1 group-producing K. pneumoniae harbored ESBL of CTX-M-15. The blaCTX-M-15 gene, first described in 2001 (20), is the most common ESBL reported in the world (4, 21, 22), including Iran. Reports have shown that this resistance gene is highly circulating in Tehran and the rest of Iran (6, 23-25). Among the ESBL-producing strains, the prevalence of blaCTX-M-15 reported in different studies in Iran varies from 62.6%, as reported by Feizabadi et al., (23) to 92.8%, as described by Alizadeh et al. (25). In several countries, such as Lebanon, India, France, Spain, the UK, Latin America, and the African countries, CTX-M-15 is now one of the most common CTX-M β-lactamases (5).
In order to comprehend how CTX-M-15 is disseminated, we performed a population analysis and gene characterization. XbaI-PFGE typing was performed in order to conduct the population analyses. Typing of isolates via PFGE grouped most of the isolates into one major cluster (Figure 1). Interestingly, this cluster was detected in two hospitals located more than 10 km away from each other. It would be worth investigating the possibility of a common source of infection, such as drinking water or food products, between the two hospitals and the possibility of inter- hospital transfer of patients. A major clone (clone A) was identified in the NICU and neonatal unit of hospital B (Milad). One possible explanation is patient-to-patient transmission between wards and the introduction of the clone into other wards through patient transfer. All the strains in this clone were simultaneously resistant to cefotaxime, ceftazidime, ceftriaxone, amoxicillin-clavulanic acid, aztreonam, tobramycin, gentamicin, cefepime, and amikacin. Antibiotic therapy with a carbapenem could be useful to control the infection. A multicountry study reported that carbapenems are the treatment of choice for infections caused by ESBL-producing K. pneumoniae (26). Co-resistance to several antimicrobial agents has often been described for CTX-M-producing isolates (27, 28). These arrays of resistance genes confer a multidrug resistance phenotype that can be transferred horizontally (28). Integrons are mobile genetic elements that can be located either on a chromo-some or a plasmid and can integrate antibiotic resistance genes. Eight of our 17 K. pneumoniae strains harbored class 1 integron, as previously described (14). The blaCTX-M-15 gene has been shown to be linked to mobile genetic elements such as insertion sequence ISEcp1 (2). We found ISEcp1 upstream of the blaCTX-M-15 gene in all strains. This insertion sequence is also able to provide promoter sequences enhancing the expression of the blaCTX-M gene (2). Another insertion sequence, IS26, was not found. The absence of amplification of the IS26-blaCTX-M-15 region in our studied strains might be due to the presence of a truncated IS26 sequence, as reported by Saladin et al. (15). In all the strains, the orf477 sequence was found downstream of the blaCTX-M-15 gene. A similar organization has been frequently found for blaCTX-M-15 in many studies (16, 29). Abbassi et al. reported the presence of ISEcp1 upstream and orf477 downstream of blaCTX-M-15 in K. pneumoniae isolates (2). Tayh et al. also found ISEcp1 and orf477 in the flanking sequences of blaCTX-M-15 in 16 broad-spectrum cephalosporin-resistant K. pneumoniae isolates collected between April and June 2013 from Palestinian hospitals in the Gaza Strip (30).
The conjugation studies revealed the transfer of plasmids carrying blaCTX-M-15 in all isolates. A high prevalence of the L/M plasmid replicon was seen in our strain collection (82.3%), although this replicon has been considered rare in CTX-M-15-expressing strains (31). Most studies have reported a dominance of IncF plasmid replicons in blaCTX-M-15-producing strains (11, 31-33). Due to several limitations, we were unable to determine the precise location of the blaCTX-M-15 gene and it was not clear which replicon type harbored this gene. The plasmids containing replicon L/M found in most of our strains may harbor the blaCTX-M-15 gene, although this point needs to be investigated further.
In conclusion, our results show that CTX-M-15 is the dominant CTX-M ESBL in our strain collection from children’s samples. We hypothesize that the clonal spread and genetic transit of mobile elements, such as ISEcp1, between unrelated strains are responsible for the dissemination of CTX-M-15 and support long term persistence in colonized hosts. Therefore, proper use of antibiotics with emphasis on the sensible prescription of antibiotics and suitable infection control measures are necessary to control the further spread of CTX-M-15-producing strains of K. pneumoniae.