1. Background
Escherichia coli is a gram-negative bacilli, which is able to colonize in humans (1). This bacterium is the most common cause of urinary tract infections (UTIs) in all age groups (2). Ability of E. coli in the pathogenesis is dependent on the virulent factors (2, 3). Several virulent factors including hemolysin (hlyA) and cytotoxic necrotizing factor type 1 (cnf1) are responsible for pathogenic potential of uropathogenic E. coli isolates (UPEC) (3, 4). These virulence factors are involved in damage to the tissues as well as bacterial dissemination (3, 4). Antibiotic resistance is another important factor in ability of E. coli in the pathogenesis and increased of morbidity and mortality of patients (5). At the moment, multi-drug resistant (MDR) and ESBLs producing isolates of E. coli have been reported worldwide and treatment options for MDR isolates is limited (5, 6). Several studies have been reported that acquisition of drug resistance mechanism may be related to loss of virulent factors (4, 6). Therefore, the aim of this study was determinate of correlation between antibiotic resistance and ESBLs production with presence of virulent genes hlyA and cnf1 in E. coli isolates collected from patients with a UTI in Kerman, Iran.
2. Methods
2.1. Sample Collection and Identification of Isolates
In a cross-sectional study, we collected 250 non-duplicate isolates of E. coli during June 2014 to March 2015 from patients with symptoms of a urinary tract infection who were admitted to the university-affiliated hospital in Kerman. Basic characteristics data such as gender and age were recorded. Bacterial isolates were identified by standard and biochemical methods including: oxidase test, lactose fermentation, growth on Triple Sugar Iron (TSI), Simmons’ Citrate Agar Media, and MRVP test (7) (All the culture media were from Merck, Co, Germany.).
2.2. Antibiotics Susceptibility of Isolates
The disk diffusion method was used for determination of antimicrobial susceptibility profile of isolates according to the Clinical and Laboratory Standard (CLSI) (8). The following antibiotics (Mast, UK) were tested: imipenem (IMI:10 μg), ceftazidime (CAZ: 30 μg), cefotaxime (CTX: 30 μg), amoxicillin/clavulanic acid (AUG: 20/10 μg), cefepime (CPM: 30 μg), Piperacillin/tazobactam (PIZ: 100/10 µg), nalidixic acid (NA: 30 μg), and ciprofloxacin (CIP: 5 μg). E. coli ATCC 25922 was used as control strain in antimicrobial susceptibility test (8).
2.3. Detection of ESBLs Producing Isolates
ESBLs producing isolates was detected by combined disk method with clavulanic acid according to CLSI guideline (8). The following antibiotics (Mast, UK) were used for ESBLs detection: CAZ (30 μg), CTX (30 μg), CPD (30 μg) alone as well as with 10 μg clavulanic acid. Klebsiella pneumonia ATCC 700603 (ESBLs positive) and E. coli ATCC 25922 (ESBLs negative) were used as control strains (8).
2.4. Detection of hlyA and cnf1 Virulent Genes
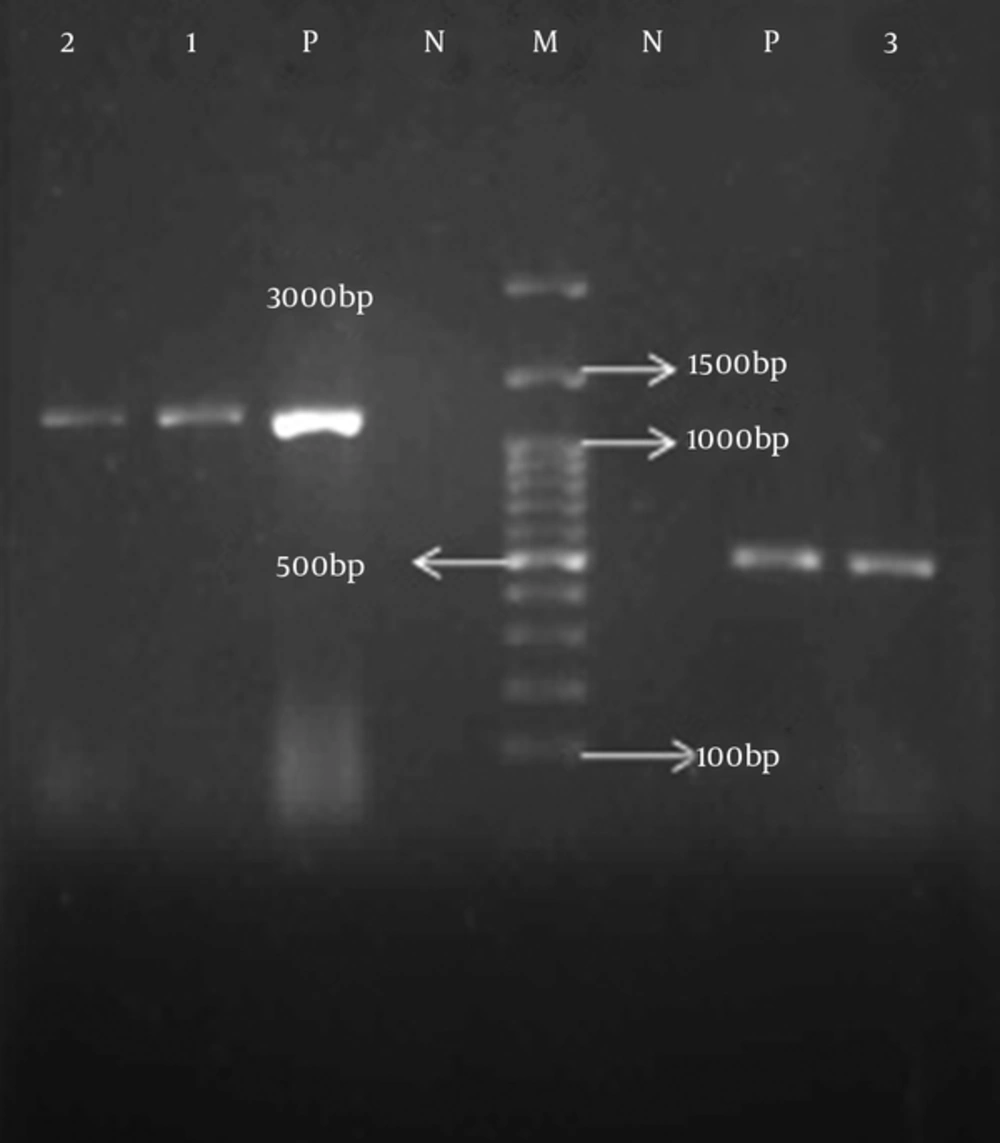
The total DNA was extracted by the boiling method (9). Virulent genes included hlyA and cnf1 detected by polymerase chain reaction (PCR). Specific oligonucleotide were: hlyA-F - 5’-AAC AAG GAT AAG CAC TGT TCT GGCT-3’ and hlyA-R -5’-ACC ATA TAA GCG GTC ATT CCC GTCA for hlyA-R gene (Product size: 1177 bp) (10) and cnf1-F - 5’-AAG ATG GAG TTT CCT ATG CAG GAG -3’ and cnf1-R -5’-CAT TCA GAG TCC TGC CCT CAT TAT T for cnf1 gene (Product size: 543 bp) (11). The PCR amplification was carried out in Biometra PCR Thermal Cycler by using PCR Master mix (Ampliqon Inc, Denmark) according to manufacture guideline, under the following conditions: initial denaturation for 5 minutes at 95°C followed by 30 cycles of 60 seconds at 95°C, and 45 seconds annealing at the specific melting temperature of each primer (62°C for hlyA and 62°C for cnf1). PCR assay was performed in a total volume of 15 μL containing: 0.5 µL of each primer (10 pM), 7.5 µL of PCR Master mix, 1 µL of DNA (20 ng) and 5.5 µL of H2O (DNase and RNase free water). PCR products were electrophoresis in 1% agarose gel.
2.5. Statistical Analysis
SPSS v.22 (SPSS Inc., Chicago, IL, USA) was used for data analysis. We used the Chi-squared test to assess the variables correlation between antibiotic resistance and ESBLs producing with presence of hlyA and cnf1 genes. P value < 0.05 was considered statistically significant.
3. Results
The age of the patients with a UTI ranged from 1 - 93 years. Among 250 UPEC isolates, 193 (77.2%) isolates were from females and 57 (22.8%) were from males. The rate of resistance to imipenem (IMI:10 μg), ceftazidime (CAZ: 30 μg), cefotaxime (CTX: 30 μg), amoxicillin/clavulanic acid (AUG: 20/10 μg), cefepime (CPM: 30 μg) and piperacillin/tazobactam (PIZ: 100/10 µg), nalidixic acid (NA: 30 μg) as well as ciprofloxacin (CIP: 5 μg) antibiotics were 4.8%, 46%, 47.2%, 61.6%, 46.8%, 24%, 68%, and 55.2%, respectively. In total, 44.8% of isolates were ESBLs positive. hlyA and cnf1 genes detected in 28.8% and 29.2% of isolates, respectively (Figure 1). Co-exist of hlyA and cnf1 genes observed in 23.2% of isolates. hlyA and cnf1 genes observed in 8% and 10% among of ESBLs positive isolates, respectively. Prevalence of NA, CIP, CTX, CAZ, PIT-sensitive isolates were over NA, CIP, CTX, CAZ, PIT-resistant isolates among the UPEC isolates with the cnf1 and hlyA (Table 1). Statistical analysis revealed significant correlation between susceptibility to NA, CIP, CTX, CAZ, PIT antibiotics with presence of hlyA and cnf1 in isolates and also significant correlation observed between ESBLs negative (non ESBLs) isolates with hlyA positive isolates (P value < 0.05) (Table 1). In our study, there was no significant correlation with age and gender of patients with presence of hlyA and cnf1 in isolates.
| Virulent Genes (n) | Antibiotic Agents and Rate of Resistance to Deferent Antibiotic, No. (%) | |||||||||
|---|---|---|---|---|---|---|---|---|---|---|
| NA | CIP | CTX | CAZ | CPM | IMI | PIZ | AUG | ESBLs | ||
| R, 170 (68) | R, 138 (55.2) | R, 118 (47.2) | R, 115 (46) | R, 117 (46.8) | R, 12 (4.8) | R, 60 (24) | R, 154 (61.6) | P, 112 (44.8) | N, 138 (55.2) | |
| hlyA(72) | 28 (16.4) | 25 (18.1) | 24 (20.3) | 22 (14.1) | 28 (23.9) | 5 (41.6) | 11 (18.3) | 27 (17.5) | 20 (17.8) | 52 (37.6) |
| cnf1(73) | 35 (20.5) | 28 (20.2) | 27 (22.8) | 25 (21.7) | 29 (24.7) | 5 (41.6) | 11 (18.3) | 28 (18.1) | 25 (22.3) | 48 (34.7) |
Abbreviations: AUG, Amoxicillin/Clavulanic; CAZ, Ceftazidime; CIP, Ciprofloxacin; CPM, Cefepime; CTX, Cefotaxime; IMI, Imipenem; N, Negative; NA, Nalidixic Acid; P, Positive; PIZ, Piperacillin/Tazobactam; R, Resistant.
4. Discussion
Virulence factors such as hlyA and cnf1 in UPEC isolates are involved in pathogenicity of this bacterium (2, 4, 6). The prevalence hemolysin (hlyA gene) and cytotoxic necrotizing factor type 1 (cnf1 gene) among UPEC isolates were 28.8% and 29.2%, respectively. The prevalence of hlyA and cnf1 in our study were similar to other studies in Iran. The prevalence of the hlyA gene was 23.3% in UPEC isolated from Iranian patient with UTI, in Jahrom Iran (12). In another study in Iran, prevalence of hlyA and cnf1 genes has been reported as 13.5% and 22.9%, respectively (13).
In agreement to other studies, prevalence of resistance to NA, CIP, CTX, CAZ, CPM, and AUG is high in our city. The rate of resistance to NA, CIP, CTX, CAZ, and AUG were reported about 53%, 51%, 69%, 72%, 77%, respectively, in Tehran, Iran (14). Furthermore, the rate of ESBLs producing isolates in our study in similar to other studies in Iran (14). In several studies in Iran, ESBLs isolates were reported in 20% to 89.8% in different regions in Iran (14, 15).
Correlations between virulent genes with antibiotic resistance in UPEC isolates have been investigated in many studies in different country of worldwide (4, 6, 16). In the present study, we observed a statistically significance between antimicrobial susceptibilities and presence of hlyA and cnf1 genes (P < 0.05).
In our study, we observed that the hlyA and cnf1 positive isolates were significantly associated with the sensitivity to NA, CIP, CTX, CAZ, and PIT as well as with the non-ESBL isolates. Harwalkar et al. reported that prevalence of hlyA and cnf1 in CIP-sensitive isolates is higher than in ciprofloxacin-resistant of UPEC isolates (4). Unlike Harwalkar et al., we observed an increased prevalence of the cnf1 gene in the non-ESBL isolates compared to the ESBL positive isolates; however, the difference was not statistically significant and the hlyA gene showed a significant association with the ESBL negative isolates (4).
Also, our study, similar to Velasco et al., showed that the resistance to nalidixic acid caused loss of virulence genes; therefore, resistance to nalidixic acid may lead to the loss of the ability to pathogenicity (17). In a study done by Arisoy et al., they showed an association between susceptibility to antibiotics and virulence factors in the E. coli isolates recovered from UTI patients (18). Similar to the present study, they reported that virulence gene hlyA was increased in sensitive isolates. Furthermore, Asadi et al. showed stronger associations between virulence factors and antibiotic resistance in UPEC isolates (12).
Finally, our findings in agreement with other studies suggested that resistance to NA, CIP, and ESBLs production may be associated with marked reduction in pathogenicity of UPEC isolates (4, 6, 17).
4.1. Conclusion
The study showed that, there are a many statistically significances between non-resistance and non-ESBLs with presence of hlyA and cnf1 genes. Our findings, similar to other studies, show that acquire of antibiotic resistance mechanism may be lead to loss of virulence factor.