1. Background
Malta fever is an infectious disease presented in acute, subacute, and chronic clinical forms in people with symptoms ranging from fatigue, fever, headache, and myalgia to spondylitis, arthritis, and endocarditis. The disease is caused by the family of the Brucella bacteria, a small, gram-negative coccobacillus, which 10 species of this bacterium have been identified so far and classified according to biochemical and antigenic features of it and host's characteristics (1). Half of a million patients with brucellosis are reported annually (2). Additionally, sheep, goats, cattle, swine, and dogs are mainly affected by various Brucella species worldwide, causing socioeconomic losses due to disease-related problems in ruminants.
In some parts of the world, this pathogen is completely eradicated, and most of the affected people are from the Middle East, Mediterranean, Saudi Arabia, Africa, Central, and South America (3). Of these, according to a report from World Organization for animal health in 2003, Iran ranked fifth for the incidence of brucellosis in human, mostly resulted from B. melitensis and B. abortus (4). Considerable failure rates in the treatment without relapse highlight pursuing the evaluation of the control and prevention of the disease for its eradication (5). The control of this zoonotic disease is based on three principles, including the identification and elimination of infected animals, pasteurizing dairy products, and livestock vaccination (6). It seems that the best way for controlling the disease in endemic areas, including Iran, remains to identify and eliminate infected animals since both vaccination of livestock and avoidance of non-pasteurized dairy products are neglected due to the low level of public awareness.
The most commonly used method for detecting contamination in livestock is Rose Bengal Plate Test (RBPT), a serological test associated with limitations in determining the different strains of the disease (7). However, RBPT does not have 100% sensitivity and specificity due to the false-negative and false-positive reports resulting from blocking antibodies and the presence of antigenic analogues with other gram-negative bacteria (including Yersinia enterocolitis, E. coli, Salmonella and Vibrio cholera), respectively (8). One of the more recent methods for detecting infected cases is molecular methods. The results of which are not affected by the presence of interfering antibodies, on one side and on the other side, they can be widely used today due to the high sensitivity and specificity, time-saving, ability for determining the species and strains of the contamination agent (9).
2. Objectives
In the present study, screened RBPT positive sheep of Mianeh (Southwest of West Azarbaijan Province, Iran) were examined in terms of Brucellosis using Polymerase Chain Reaction (PCR) method for evaluation of the Brucella abortus Cell Surface Protein 31 (BCSP31) (10), since there was no information about the livestock brucellosis prevalence in this area. Moreover, the study aimed at estimating the prevalence of the common gene mutations involved in antibiotic resistance for fluoroquinolones (topoisomerase IV subunit C [parC] and DNA gyrase subunit A [gyrA]), rifampin (RNA polymerase β-subunit [rpoB]) (11), and streptomycin (aminoglycoside adenylyl-transferase-A1 [aadA1]) (12) to provide further information needed for choosing the most efficient and affordable treatment protocol. It is noteworthy to mention that these mutations result in the impairment of the binding site of antibiotics and affect the function of antibiotics as the main mechanism for resistance to antibiotic therapy.
3. Methods
3.1. Ethical Considerations and Preparation of Samples
In this study, 1,220 sheep aged more than 8 months from Mianeh and its suburb were selected randomly from May 2017 to April 2018, in which only 150 sheep were vaccinated against brucella spp. All procedures on animal were approved by the Ethics Committee of Mofid Children’s Hospital (IR.SBMU.RETECH.1396.1383). Under sterile and hygienic condition, 5 cc of blood was aseptically taken from jugular vein of each sheep and divided in tubes for serum preparation by centrifugation and also in tubes containing ethylene-diamine-tetra-acetic acid (EDTA) which were transferred and stored at -20ºC.
3.2. Serological Analysis
Agglutination-based RBPT was done through dispensing 30 μL of each serum sample on white glossy ceramic tile. After adding an equal volume of room temperature equilibrated RBT antigen, the content of each tile was mixed using applicators and the tile was left rocked on a rotator for 4 min (13). Eventually, any visible agglutination was regarded as a suspicious sample for Brucella species for further investigations using PCR.
3.3. DNA Extraction, Amplification and Detection of PCR Products
The extraction of the DNA was done from whole blood of RBPT positive sheep samples using QIAamp DNA mini kit (QIAGEN, Venlo, Netherlands) according to the manufacturer’s instructions. Samples were subjected to PCR (Master-Mix from amplicon, Korea for all PCR procedures) using specific primers for BCSP31 (Table 1) as a genus specific gene for Brucella spp. An initial denaturation at 94ºC for 30 min followed by 30 cycles of 30 sec at94ºC, 1 min at 56ºC and 30 sec at 72ºC of annealing which ended with a final extension at 72ºC for 3 min were completed for each sample (SimpliAmp Themal cycler. Applied Biosystems by Life technology).
| Gene/Primer | Sequence (5’ - 3’) | Reference |
|---|---|---|
| BCSP31 | (14) | |
| BCSP31-F | AAGGGCAAGGTGGAAGATTT | |
| BCSP31-R | CCTCGTTCCAGAGAACCTTG | |
| aadA1 | (15) | |
| aadA1-F | GTGGATGGCGGCCTGAAGCC | |
| aadA1-R | ATTGCCCAGTCGGCAGCG | |
| gyrA | (16) | |
| WP-gyrA | GACATTGCGAGAGAAATTACAC | |
| Control-gyrA | GATGTTGGTTGCCATACCTACG | |
| MAMAgyrA83 | CGGGCCAGATACTGTGCT | |
| MAMAgyrA87 | CTTTGCTAGCAGGCGTACCGCG | |
| parC | (16) | |
| WP-parC | CGGAAAACGCCTACTTAAACTA | |
| Control-parC | GTGCCGTTAAGCAAAATGT | |
| MAMAparC80 | TCTCGGACAATACTTCGCTA | |
| MAMAparC84 | CTCCGCTACCAGGACTACC | |
| rpoB | (17) | |
| ROF | GTCGCCGCGATCAAGGA | |
| RIR | TGACCCGCGCGTACAC | |
| R516B | GCTGAGCCAATTCATGGA |
Abbreviations: F, forward; R, reverse.
Identification of a mutation in aadA1 resulting in Streptomycin resistance was investigated using specific primers (Table 1). aadA1 is a member of gene family responsible for resistance to aminoglycosides through the production of aminoglycoside adenylyl-transferase (AAD) which inactivates streptomycin and spectinomycin by adding an adenyl or other nucleotidyl groups (18). To this end, conventional PCR included a thermal profile as follows: an initial activation of PCR at 95ºC for 5 min, 35 cycles of a three-step annealing/elongation (45 sec at 95ºC, 45 sec at 68ºC and 45 sec at 72ºC) and a final extension at 72ºC for 5 min.
Pair of distinct MAMA (Mismatch Amplification Mutation Assay) primers (Table 1) along with related control and WP primers was used to evaluate BCSP31 positive samples for mutated gyrA and parC as target genes for antibiotic resistance in fluoroquinolones. In this regard, for MAMA-PCR, an initial denaturation at 94ºC for 5 min and 30 cycles annealing of 40 sec at 94ºC, 40 sec at 54ºC and 40 sec at 72ºC ending with a final extension at 72ºC for 5 min.
Moreover, multiplex allele-specific (MAS) PCR was used for the investigation of mutation in rpoB as important and common mutated genes responsible for Rifampin resistance. For detection of a mutation in 516 codon of rpoB, an internal forward primer (R516B) along with a pair of external primers was used (Table 1). For this purpose, an initial denaturation at 96ºC for 3 min followed by 30 cycles of 50 sec at 95ºC, 40 sec at 65ºC and 20 sec at 72ºC of annealing and a final extension at 72ºC for 3 min were completed for each sample.
Eventually, PCR products along with a DNA ladder (Korea) were run onto electrophoresis gel (1.5% agarose containing TBE buffer; 100 mMTris-HCl (pH 8), 90 mM boric acid, and 1 mM Na2EDTA) with voltages of 100 V for 90 min, simultaneously. Obtained bands were visualized under UV light using UVIdoc gel documentation systems (Uvitec, UK) and molecular weight of each PCR product was compared with ladder.
3.4. Statistical Analysis
Statistical analyses were performed using Statistical Package for the Social Sciences; SPSS 23.0 software (SPSS Inc., Chicago, USA). In this study, only qualitative variables were reported as relative frequency (percent).
4. Results
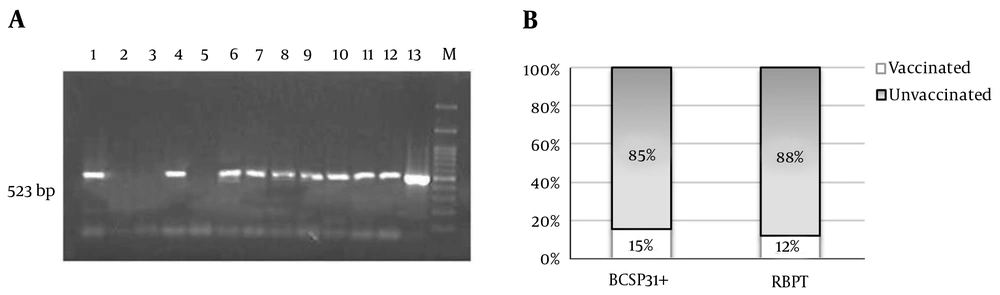
Of all 1,220 samples, 111 cases were reported positive for RBPT in which 13 cases were among those vaccinated against Brucella spp. Of these RBPT positive samples, 41% (n = 46) were confirmed to be positive for Brucella spp. using PCR for BCSP31 (Figure 1).
BCSP31: (A) Agarose gel electrophoresis of PCR products indicates positive Brucella spp. at 523 bp for 1,4, 6-12 and negative result for 2,3,5 (13 as positive control and M as ladder). (B) About 15% (n = 7) of BCSP31 positive samples were from vaccinated and 85% (n = 39) from unvaccinated sheep. Whereas, 13 out of 111 RBPT positive samples (12%) belonged to vaccinated sheep.
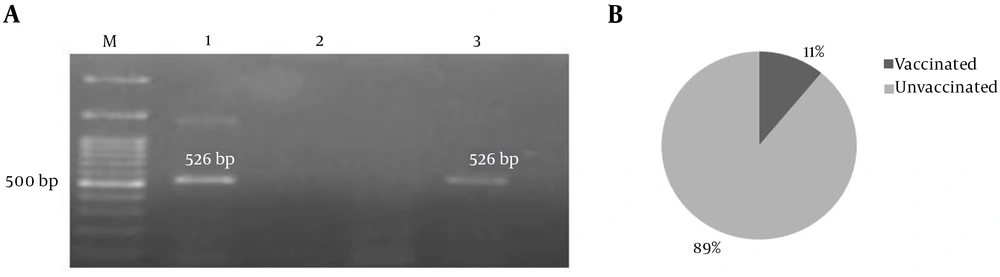
Of 46 BCSP31 positive samples, 9 (19.6%) were reported positive for aadA1 PCR product (526 bp) in which only one sample was from sheep vaccinated against Brucella spp. (Figure 2).
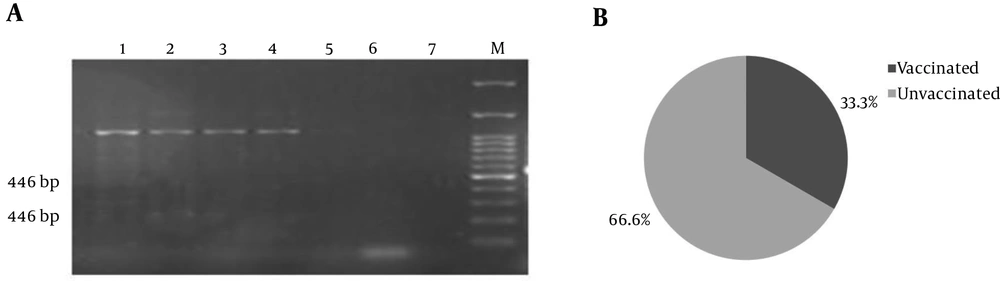
Only three of BCSP31 positive samples were shown to have a mutated parC (Figure 3). However, none of them were shown to contain a mutated gyrA because only 540 bp PCR products were visible on gel electrophoresis.
parC: (A) Duplication of two small gene fragments (446 bp and 238 bp) is carried out by these primers in wild strains (1 - 4 samples), while strains carrying mutations do not have 238 bp PCR products with MAMAparC primers (sample 5) compared to negative controls (6 and 7). (B) One out of three parC mutation-positive samples belonged to vaccinated sheep.
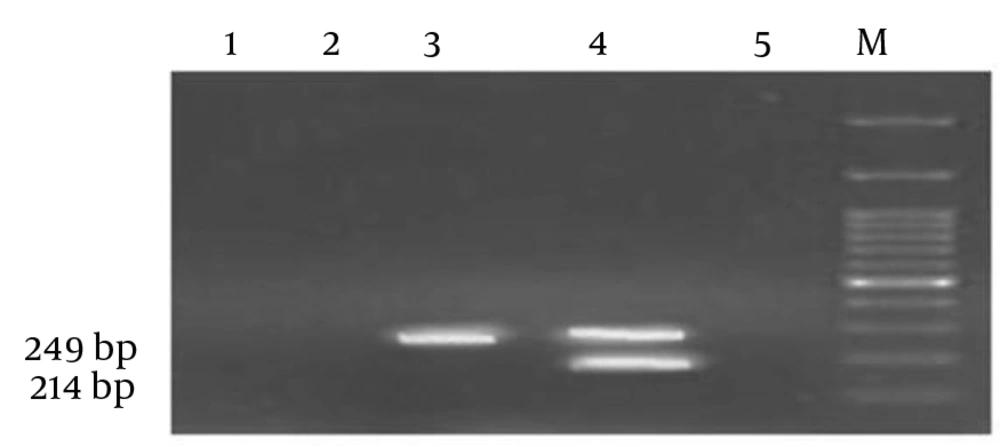
A BCSP31 positive sample obtained from a sheep vaccinated against Brucella spp. was shown to be positive for rpoB mutated strain resulting in resistance to rifampicin (Figure 4).
rpoB: Simultaneous presence of a 214 bp along with a 249 bp MAS-PCR product were regarded to be indicative of the wild-type strains (sample 4) where detecting a 249 bp MAS-PCR product were representative of rpoB mutated strain (sample 3) compared to negative controls (1,2 and 5) and loaded ladder (M).
It should be noted that results of internal control in all strains were positive for both genes, which confirms the accuracy of DNA extraction, as well as PCR and relative conditions.
5. Discussion
In the initial screening of this cross-sectional study, 111 RBPT positive samples out of 1,220 sheep blood were reported. Of these, 46 (41%) were detected to be BCSP31 positive using PCR method. The results of the investigations for mutations responsible for antibiotic resistance were indicative of 9 cases for aadA1, 3 for parC and one for rpoB positive.
Previously, similar studies have been conducted to determine the rate of Brucella spp. infection in livestock from different parts of Iran. Seroprevalence of brucellosis using RBPT, Wright standard tube agglutination and 2-mercaptoethanol (2-ME) agglutination tests were reported for 3% (n = 77) among 2,550 sheep investigated in Hamedan, Iran (19). Gharekhani et al. in abovementioned study also declared that seroprevalence rate is significantly (P < 0.05) lower in animals vaccinated against brucellosis which is in consistence with results of the current study (Figure 1B). Additionally, another study conducted on 740 blood samples of sheep from Sarab (East Azarbayjan province, Iran) using RBPT, serum agglutination test (SAT) and 2-ME test indicated that seroprevalence of brucellosis was 4.18% (n = 31) (20). But, the results of the RBPT in the current study were indicative of higher seroprevalence (approximately 9%) for brucellosis in sheep from Mianeh. Similarly, Shahbazi et al. reported an 11.12% (n = 397) seroprevalence rate for brucellosis in 3,570 sheep from Kermanshah province, Iran, using Wright, 2ME, and RBPT (21). Differences in the seroprevalence of brucellosis in various studies are due to various factors such as seasonal pattern of the disease occurrence (affecting the sampling time), vaccination pattern and livestock control, and most importantly the low levels of accuracy and sensitivity of the selected method for screening.
Several studies examined various animal samples using PCR method to isolate positive samples for brucellosis more specifically (22-26). In a previous study, Mahzounieh et al. reported that 62.3% of seropositive samples showed negative PCR test results (27). This finding is almost consistent with the results of the present study, in which 59% of seropositive specimens for brucellosis did not show Brucella spp. specific BCSP31 amplification. This may be due to the cross-reaction of antibodies produced against other species of gram-negative bacteria present in the samples or activation of the immune response and the destruction auto-antigens (8). DNA extracted from the whole blood samples was used for PCR with specific primers. Although it does not provide the sensitivity and specificity for the isolation of the bacteria from clinical specimens, it provides more accurate results when compared to serological methods, especially in avoidance of false-positive results for vaccinated animals, which misleads the diagnosis of active infections (28, 29). As shown in Figure 1B, the number of specimens suspected for contamination with Brucella spp. was reduced from 13 cases detected by RBPT to 7 cases in the PCR investigation for BCSP31 gene.
Here, aadA1 confers spectinomycin and streptomycin resistance which has been shown that streptomycin is still effective for isolates of Brucella spp. (12, 30), but the results of this study may indicate that there is still concern about the emergence of resistance-causing mutations. Additionally, mutations in rpoB were also investigated to check the status of rifampin resistance as a substitution for streptomycin with milder side effects. In contrast, in other studies, including the study of Irajian et al. (30), although the fewer side effects with rifampin made it a good choice instead of streptomycin, the significant reason for the superiority of streptomycin was the drug resistance reported for rifampin in Iran (due to endemic tuberculosis), but in the current study, mutated rpoB as an indicator of resistance to rifampin was reported in only one case (Figure 4). Thus, more studies are needed to compare the effectiveness of these two regimens (DS and DR) in Iran along with the mutational investigations on isolated strains. Mutation detection in parC was reported in 3 samples (6.52%) in our study, and no mutation was detected for gyrA. In 2008, Valdezate et al. conducted a study on 62 isolated B. melitensis from patients over a period of 11 years to determine whether rifampicin and fluoroquinolone-resistance in B. melitensis were detectable using PCR (11). In the abovementioned study, one fluoroquinolone-resistant case with mutation in gyrA was reported (1.6%) in which no mutations were detected for parC and rpoB. It may be concluded that the lack of rpoB mutation associated with resistance to rifampin supports the selection of this antibiotic in the treatment of brucellosis and demonstrates the usefulness of PCR screening for resistant genotypes.
Currently, there is no definitive vaccination against brucellosis, and efforts are still ongoing to produce an effective vaccine. Vaccination can interfere with the results of screening tests of infected animals in different ways through the production of antibodies and DNA fragments causing false-positive for the detection of acute brucellosis. As shown in Figure 1B, 7 out of 46 BCSP31+ samples were obtained from vaccinated sheep, which only four of them were shown to be positive in terms of drug-resistant genes (rpoB, parC and aadA1). These four samples were regarded to be BCSP31+ as a result of vaccination but not resulted from the presence of active brucellosis. For three other BCSP31+ samples that were reported negative for the evaluated mutation carrier genes, more molecular studies are needed to prove that the positive PCR result for Brucella is whether due to the strains used in the vaccine. These findings may also show evidence of a failure in vaccination that can be estimated at least 1.3%, and the best scenario for the remaining 11 (7.3%) is to consider them as false-positives resulted from vaccination.
A previous study has shown that the most important risk factors for brucellosis in Mianeh are occupational exposure, food consumption and access to unpasteurized dairy products, the control of the disease in human is highly dependent on animal disease control and the need for completing the information in this area still exists (31). The results of this study emphasize, in line with the human studies, the higher prevalence of the disease in livestock of Mianeh, the effectiveness of vaccination in them, and as a consequence the requirement to increase the awareness of disease control. Unfortunately, the results of human studies have not been found for mutations responsible for drug resistance in patients with brucellosis in the Mianeh, and this study seems to be necessary to compare with livestock results in order to assess the policies for controlling the disease.
5.1. Advantages and Limitations of the Study
This study has revealed some important mutations of genes of antibiotic resistance in sheep brucellosis (Streptomycin, fluoroquinolones, Rifampin resistance). The limitations were about the number of specimens and failure to isolate the microorganisms from culture.