1. Background
Enterococcus faecalis is one of the most common types of group D streptococci or enterococci, which is present in the gastrointestinal tract as a normal flora (1). So far, many studies have investigated the role of Enterococcus in the development of nosocomial infections caused by antibiotic-resistant bacteria. For example, this bacterium has recently been identified as a major leading cause of nosocomial infections, mainly due to its ability to induce biofilms (2). Microbial biofilms are complex communities of surface-bound bacteria that cause irreversible binding of microorganisms to surfaces such as living tissues, increased antimicrobial resistance, and pathogenicity (3). Several virulence factors inducing biofilm formation have been identified in E. faecalis strains, including surface protein compounds (Esp, GelE, and Agg) (4). E. faecalis surface protein (Esp) is a cell wall protein contributing to bacterial adhesion, colonization of surfaces such as the urinary tract, and biofilm production (5). It has been shown that isolates without Esp are able to produce biofilm after obtaining the plasmid carrying the Esp gene (6). The presence of the Esp gene in E. faecalis strains isolated from urine samples and catheters with high antibiotic resistance may be associated with further bacterial colonization and the development of urinary tract infection (7). Esp is a high-molecular-weight surface protein consisting of 1873 amino acids as well as N-terminal, central nucleus, and C-terminal domains. The C-terminal domain contains a hydrophobic membrane-spanning region. Recently, it has been finding that the N-terminal domain of Esp contributes to the interactions with host cells, and the central domain of this protein plays an important role in the accumulation of bacteria and the camouflage of the abounded protein in the host immune system (8). The occurrence of pathogenicity factors such as biofilm formation and increased resistance of E. faecalis to existing antibiotics through the expression of Esp is one of the important problems in the treatment of nosocomial infections. Therefore, exploring new antibacterial agents and modern methods to defeat microbial resistance is essential and is considered as a priority in medical studies (9). Research has shown that various synthesized products containing the central ring of 1, 3, 4-oxadiazole can inhibit the growth of gram-negative bacteria in certain concentrations (10). Many studies have investigated the antibacterial properties of different 1, 3, 4-oxadiazole derivatives on E. faecalis strains in order to provide pharmaceutical structures against this bacterium (11-14). The use of computational methods to estimate the activity of different molecules as drug candidates accelerate the process of discovering new drugs. Today, computational and fast docking tools are essential for the rational design of drugs. These techniques are based on the atomic structures and the interactions between atoms. In these techniques, the ligand or drug is moved randomly or by special algorithms at a macromolecular level so that an ideal position is found to be attached to (15, 16). In this study, the effect of synthesized materials was initially investigated on the expression of biofilm-inducing Esp in E. faecalis isolates. The effect of synthesized derivatives on the bacteria present in the biofilm, especially in deeper layers, was comparable to the antibiotic treatment and phagocytic function of immune cells through inhibiting biofilm formation and regulating the development of degenerative environments. Just as it is important to identify genes involved in the biofilm formation and pathogenesis of this bacterium, it is equally important to recognize that environmental factors play a role in regulating the expression of these genes.
2. Objective
The present study aimed to investigate the effect of new 1, 3, 4-oxadiazole derivatives based on phenyl group on the expression of E. faecalis. Esp gene, which is effective in promoting the biofilm formation ability of drug-resistant specimens. Then, the binding mechanism of compounds affecting the expression of the desired gene to the active site of Esp was investigated, the molecular docking method was performed, and the results were analyzed.
3. Method
This research study was conducted in the microbiology laboratory of Islamic Azad University, Tehran Branch (code number: 10130553971010) in 2019. Starting materials, solvents, and culture environments were obtained from Merck, Germany, and were used without further filtration. Microbiological tests were performed using a Memmert- INC153T2T3 incubator.
3.1. Preparation of Samples
In this study, 100 urine samples cultured on plates from patients with severe urinary tract infections were delivered by the research department of Shariati and Sarem hospitals in Tehran. In the microbiology laboratory of Islamic Azad University, Tehran Branch, 10 enterococcal strains were identified using biochemical methods. A pure culture of all isolated enterococcal strains was prepared in a blood agar medium, inoculated in brain-heart infusion broth containing 20% glycerol, and stored at -70°C. Biochemical tests including scoline hydrolysis, catalase, detection of pyrrolidonyl aminopeptidase, growth in the presence of Na 6.5 NaCl, motion test, pigment production, and casein test were performed.
3.2. DNA Extraction and Gene Identification
To extract bacterial DNA, several colonies of E. faecalis isolates were cultured in one to four milliliters of lysogeny broth (LB). The resulting precipitate was used to extract bacterial DNA using a DynaBioTM DNA extraction kit (CAT# KI0015). A PCR reaction was performed in a final volume of 20 μL containing 10 μL of Master Mix 2X, 1 μL of each primer (0.2 pmol), 4 μL of H2O, and 4 μL of template DNA (5 ng) using an Eppendorf Mastercycler Gradient. The timing program used to amplify the Esp gene was as follows: An initial degradation step at 95°C for 3 min followed by 35 cycles of degradation at 95°C for 30 sec, a binding step for the primers at 53°C for 30 min, and an extension at 60°C for 60 sec. A primer pair for Esp with a sequence (forward: 5'- GTGCGCGAAAATCAACACT-3', reverse: 5'-CATTTTTACGCATCCGGACTA-3') and product length of 100 bp was used; the design and synthesis of the primers were ordered to the South Korean Macrogen Company; also, a primer pair for 16s rRNA with a sequence (forward: 5'- TCAGCAGGGAAGAAGCGAAA-3', reverse: 5'- CCTACGAGCTCTTTACGCCC-3') and product length of 127 bp was used for control. Finally, PCR products were transferred to 1% agarose gel containing ethidium bromide, and after completing the electrophoresis step, they were visualized by UV-transilluminator. The marker used in this research was DNA Ladder Mix made by the SinaClone Company (Code: 2365) with a distance of 100 bp from 100 to 1300 bp.
3.3. Preparation of Suspension Derivatives
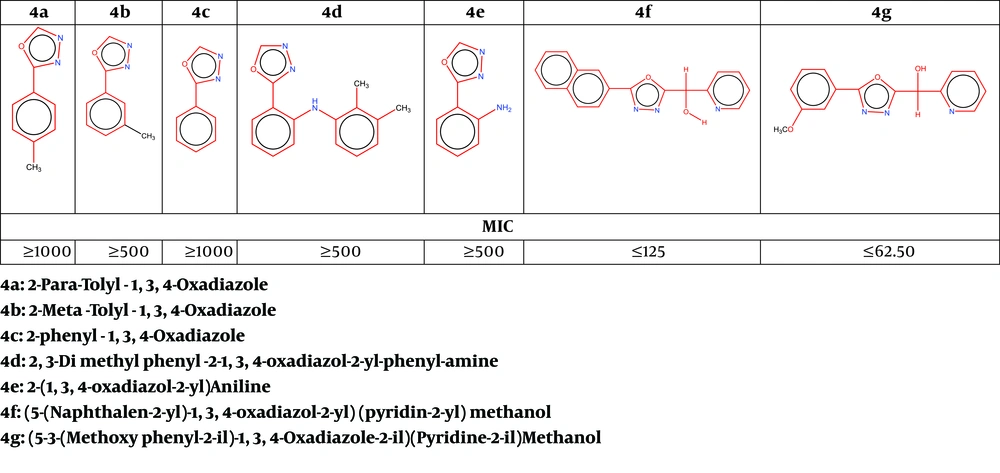
To prepare the stock solution, 10 g of each synthesized material was dissolved in 100 mL of dimethyl sulfoxide (99% DMSO) (Figure 1).
Structures of new 1, 3, 4-oxadiazole derivatives and their MIC results (10).
3.4. Determination of Minimum Inhibitory Concentration
Using the standard method recommended by the Clinical and Laboratory Standards Institute (CLSI) (17), the minimum inhibitory concentration (MIC) was applied to E. faecalis isolates in contact with synthesized derivatives. Based on this method, a 96-cell microplate with 12 rows was used, each containing 100 μL of sterile TSB (tryptic soy broth) medium. Then, 100 μL of the synthesized compound with a high dilution of 1000 μg/mL was added to the first row containing 100 μL of culture medium (TSB), and dilution continued until reaching a dilution of 62.50 μg/mL (1000 - 62.50 μg/mL). Afterwards, 100 μL of half-McFarlane microbial suspension was added to all wells. The microplates were then incubated at 37°C for a maximum of 24 hours (Figure 1).
3.5. RNA Extraction and cDNA Synthesis
Total RNA extraction was performed using a Cinnapure RNA extraction kit, and then the bacterial suspension containing Esp was treated with a dilution of the synthesized compound obtained from MIC (dilution after MIC). A DNase1 (Fermentase USA, Waltham) kit was used to remove possible RNA contaminants, extracted with genomic DNA, before cDNA synthesis. cDNA synthesis was performed using the AMV (avian myeloblastosis virus) reverse transcriptase enzyme (the product of Roche Company with a concentration of 25 U/μL). The extracted RNA was incubated at 65°C for 3 min. Then, reverse transcription (RT) was performed at 42°C for 60 min with 2 μL of random primer, 0.8 μL of AMV reverse transcriptase enzyme, 2 μL of dNTP (10 mM), and 2 μL AMV enzyme 10x buffer.
3.6. Esp Expression in the Presence of Derivatives
The polymerase chain reaction was performed in a final volume of 20 μL (Prime QMaster mix (2x) with cyber green (10 μL), forward primer (1 µL), reverse primer (1 µL), Rox dye (1 µL), DEPC water (5 µL), cDNA (2 µL)) using a Genet Bio kit (South Korea) (CAT NO: Q9210) in a Step One Plus Real-Time PCR system (Applied Biosystems, CA, USA). The 16R rRNA reference gene was used as the internal control in this study. Graph Pad Prism 8.4.3 software was used to calculate the relative expression of the gene, analyze data, and draw the relevant graphs. In order to obtain the ratio of the target gene expression to the reference gene, the data related to mRNA gene expression changes were analyzed by ΔCT = CT Esp – CT 16s rRNA, ΔΔCT = ΔCT Esp – ΔCT c.s and 2-ΔΔCT (-Log2) method based on efficiency.
3.7. Molecular Docking Studies
The initial structures of the compounds were plotted using ChemDraw 19.1 software. Chem3D software was used to optimize the spatial structures of the compounds. The compounds were optimized by the MM2 method. The Esp protein structure was obtained from the RCSB site with 6ORI code as PDB extension. Using AutoDockTools-1.5.6 software, both receptor files (6ORI) and ligand (derivatives) were converted to the required formats for the docking process. Receptor-ligand interactions were investigated using AutoDock Vina software and the best connection state was calculated. Receptor-ligand interactions were then observed using Discovery Studio 4.5 Client software. In order to better understand, the two-dimensional (2D) structure was used to present the results.
4. Results
4.1. Isolation and Identification of Strains from Delivered Samples
In this study, a total of 10 strains were isolated and identified as E. faecalis using diagnostic and confirmatory microbiological/biochemical tests.
4.2. Esp Molecular Identification
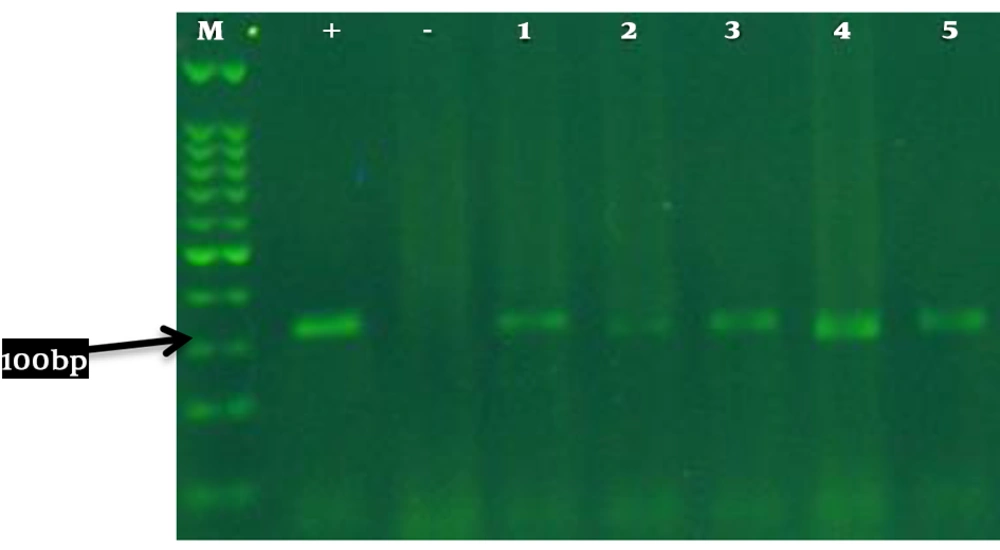
The amplification of Esp in bacterial isolates showed the presence of a 100 bp band in gel electrophoresis (Figure 2). Molecular identification of the Esp locus was performed for 5 isolates, and then the inhibitory activity of the synthesized compounds on biofilm formation and Esp expression was evaluated in these 5 isolates.
4.3. Esp Expression After Exposure to Inhibitors of Biofilm Formation
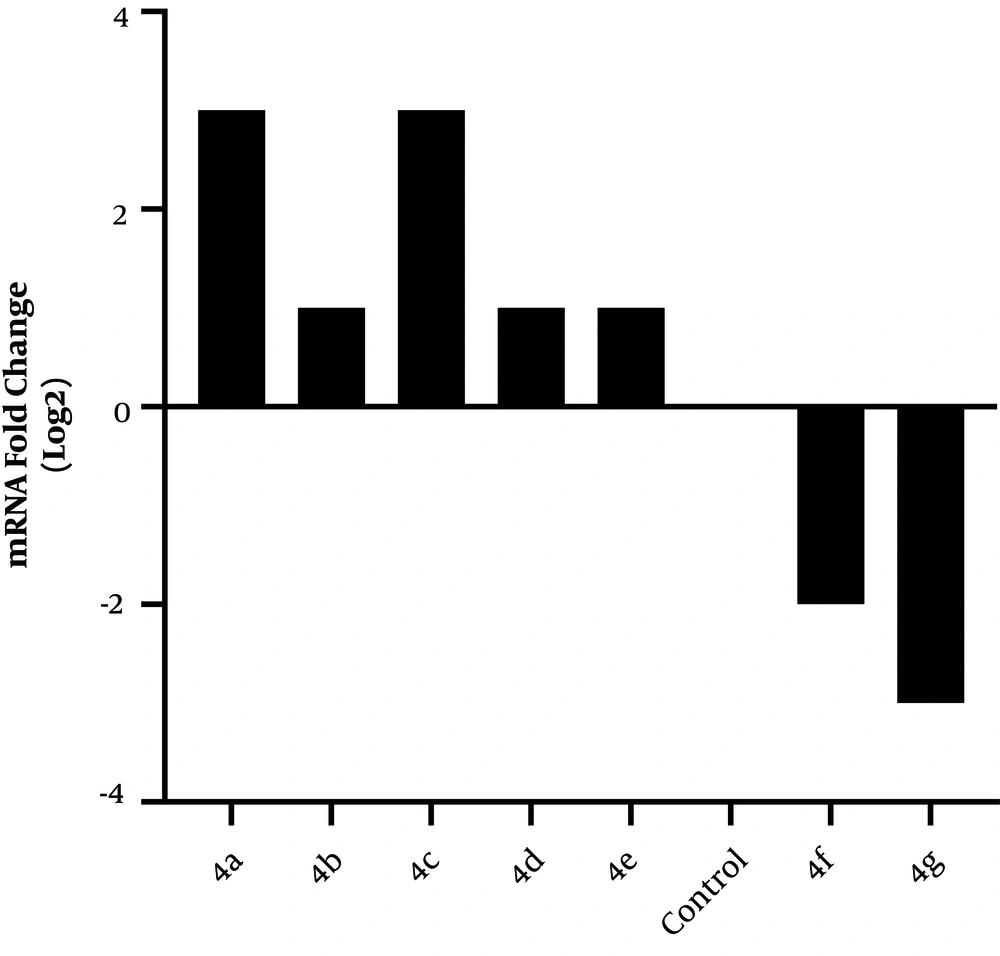
The relative expression of Esp was significantly reduced in the presence of derivatives, so that compounds 4f and 4g caused respectively a 2-fold and a 3-fold decrease in Esp expression compared to the control group (Figure 3).
4.4. Molecular Docking
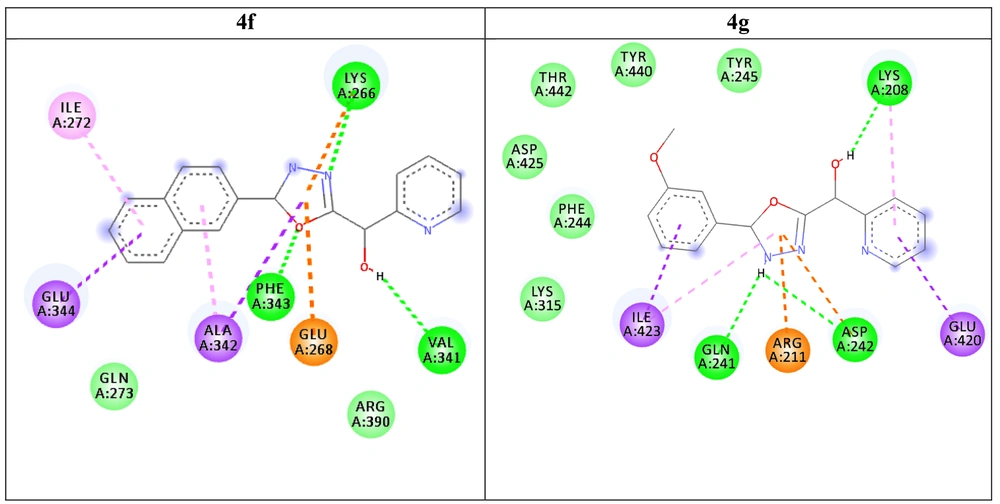
Initially, the CA2+ ligand was removed from the active site of the protein to validate the docking operation. After validation of the docking protocol, the three-dimensional structure of the synthesized compounds was docked into the active site of Esp. Data regarding the total energy of derivatives calculated by the MM2 method, ligand affinities, and amino acids involved in the formation of interactions as well as the lists of all ligand interactions, including hydrogen bonds and hydrophobic interactions are presented in Table 1. Moreover, the docking 2D results are shown in Figure 4.
| Derivative | Total Energy (K cal/mol) | Affinity | Amino Acids | ||||||
|---|---|---|---|---|---|---|---|---|---|
| Hydrophobic Bonds | Hydrogen Bonds | ||||||||
| Pi-cation- anion (Ionic Bond) | Pi-sigma (Covalent bond) | Pi-alkyl | |||||||
| 4f | 8.7341 | -9.2 | GLU, A:268 | ALA, A:342 | GLU, A:344 | ILE, A:272 | LYS, A:266 | VAL, A:341 | PHE, A:343 |
| 4g | 18.5052 | -7.6 | ARG, A:211 | GLU, A:420 | ILE, A:423 | - | LYS, A:208 | ASP, A:242 | GLN, A:241 |
Auto Docking
5. Discussion
The surface protein of E. faecalis (encrypted by chromosomal Esp) is involved in the primary binding and biofilm formation of this bacterium, and with the exacerbation of the disease, colonization is accompanied by bacterial stability in host tissues, especially in the urinary tract (18). Therefore, inhibiting biofilm formation is medically important (19). In this study, the in vitro and in silico ability of newly synthesized derivatives to inhibit biofilm formation induced by Esp was investigated. Biofilm formation is an important aspect of the pathogenicity of these bacteria induced byEsp; therefore, controlling these virulence factors could prevent severe infections caused by this bacterium. In the present study, molecular identification of the Esp locus was performed for 5 isolates, and then the inhibitory activity of the synthesized compounds on biofilm formation and Esp expression was investigated in these isolates (5 out of 10 isolates). Then, the interactions between compounds and surface proteins were designed and calculated through Auto Dock Vina software. The E. faecalis Esp gene has shown resistance to many synthetic and existing substances. Amini et al. (2019) investigated the inhibitory effect of curcumin nanoparticles on Esp expression associated with E. faecalis biofilm. They reported that according to the results of MIC and gene expression assay, nano-curcumin extract had no antimicrobial and inhibitory effect on biofilm formation and Esp expression in E. faecalis bacterial isolates (20). These results indicate the important role of Esp in biofilm formation in infections caused by this bacterium. The design and synthesis of new and alternative drugs are essential in today's world. In this regard, oxadiazole structures, especially 1, 3, 4-oxadiazole derivatives, are important among other compounds (21). They could be synthesized through a one-step process as a new step in the synthesis of drug structures. In this study, the relative expression of Esp in isolates treated with the synthesized materials significantly changed compared to the control isolates, so that compounds 4f and 4g caused significant decreases in Esp expression. Antibacterial, antiviral, and antifungal activities of 1, 3, 4-oxadiazole derivatives have been demonstrated in recent years through in vivo and in vitro studies. Due to the novelty of the synthesized derivatives used in this study which were synthesized for the first time by a one-step process, no study has evaluated the effects of these derivatives on Esp expression, and this was the first research to report the anti-biofilm properties of these compounds. In the present study, the results of MIC and Esp expression showed that our synthesized compounds had a very high inhibitory activity as well as a much higher antibacterial activity suitable for reducing Esp-mediated biofilm formation, especially compounds 4f (MIC 4f: ≤ 125, causing a 2-fold decrease in Esp expression than the control sample) and 4g (MIC 4g: ≤ 62.50, causing a 3-fold decrease in Esp expression than the control sample). These results are consistent with the results of other studies performed on gram-positive bacteria such as Staphylococcus aureus. Other compounds (4a, 4b, 4c, 4d, and 4e) alone did not significantly change Esp expression within the MIC range, and their maximum synergistic activity appeared to be in combination with other synthesized drug structures. The results of this study showed that compounds 4f and 4g could inhibit by binding to the active site of Esp. Based on the docking results, the inhibitory potential of the studied compounds differed, and the strongest binding with the highest inhibitory potential is related to the compound 4f. This compound with ΔG = -9.2 kcal/mol affinity had the most negative bond energy level among the two studied compounds. ΔGbind indicated the strength of the bond between the molecular compounds and the active site of the enzyme. In fact, ΔGbind is the sum of the two factors of intramolecular energy and torsional energy. The compound 4f can prevent the binding of the substrate to the active site by interacting with key amino acids of the active site of Esp. Table 1 and Figure 4 show all possible hydrogen bonds of this ligand with the active site. Based on the results of the present study, it can be concluded that this compound can inhibit Esp while interacting with important amino acids located in the active site of Esp. The combination of 4g with hydrogen and hydrophobic bonds can also be a good option to inhibit this protein. Due to high prevalence of Esp gene in drug-resistant E. faecalis clinical isolates in this research as well as in other studies, this finding could be considered as a diagnostic criterion in molecular identification of drug-resistant E. faecalis strains, and due to the biofilm formation ability of these bacteria, the synthesized derivatives could be used in designing and producing new therapeutic drugs.