1. Background
Gastric cancer (GC) is the third principal cause of cancer-related death (1) and has a poor prognosis due to the lack of specific symptoms, leading to late diagnosis, as well as low survival rates (2, 3). Common chemotherapy drugs for the treatment of GC include paclitaxel (PTX), adriamycin (ADR), vincristine (VCR), 5-fluorouracil (5-FU), and platinum; however, chemoresistance is a major obstacle in treating GC (4). Therefore, understanding GC pathogenesis and finding new diagnostic and therapeutic biomarkers are necessary for the managing and targeted-therapy of GC.
Non-coding RNAs are classified into two groups: microRNAs (miRNAs, as a group of small ncRNAs, ~18 nt) and long non-coding RNAs (lncRNAs, > 200 nt) (5, 6). The recent group has been shown to play important roles in regulating gene transcription, post- transcriptional modifications, epigenetic regulation, and finally, protein translation (7). Various studies have shown that LncRNAs are involved in a variety of biological processes like chromatin regeneration, carcinogenesis, and cell differentiation (8). Furthermore, lncRNAs have been reported to be associated with tumor invasion, cellular proliferation, and metastasis in a variety of cancers (9-11). Studies have shown that several lncRNAs, such as HOXA11-AS, LINC00673, and XIST, facilitate GC progression by regulating β-catenin, LSD1, and miR-101 (12-14). On the other hand, linc00261 was shown to inhibit GC progression (15). Another lncRNA, AK023391, has also been reported to be involved in tumorigenesis and metastasis by activating the PI3K/Akt signaling pathway in GC (16). Also, investigating the profile of lncRNA expression revealed the upregulation of AK023391 in GC tissues and cell lines, indicating its important function in GC. Despite the fact that FISH analysis (using a tissue microarray) revealed no link between AK023391 expression and the clinicopathological features of 77 individuals with GC, Kaplan–Meier analysis revealed that elevated AK023391 expression was associated with poor survival in patients with GC. Furthermore, multivariate analysis indicated that AK023391 expression was an independent predictor of GC patients’ overall survival (OS) (16).
Forkhead box O3A (FOXO3a), also known as FOXO3 or forehead in rhabdomyosarcoma-like 1 (FKHRL1), is a member of the FOXO subfamily, which was initially discovered in the human placental cosmid. The FOXO3a gene is found on chromosome 6q21 (17) and regulates a wide range of cellular activities via controlling the expression and activity of effector genes. The activities and functions of FOXO3a are dependent on its subcellular location (18). The phosphorylation of FOXO3a leads to its translocation from the nucleus to the cytoplasm where it interacts with the 14-3-3 protein, preventing it from returning to the nucleus (19).
Forkhead box O3A is an important transcription factor that plays an essential role in tumor progression, angiogenesis, and cancer metastasis. This transcription factor is also important in GC progression by targeting the signaling pathway of PI3K/ protein kinase B (20). In addition, functional studies indicate that the upregulation of FOXO3a in GC tissues is associated with a poor prognosis in GC patients with stage II / III disease (21). More research is needed to identify an AK023391 LncRNA level correlation with FOXO3a in GC tissues.
2. Objectives
In the current study, the mRNA levels of the FOXO3a and AK023391 genes were investigated in GC tumor tissues. Also, the relationships of FOXO3a and AK023391 expressions with the clinicopathological features of GC were investigated.
3. Methods
3.1. Patients
In the current case-control study, 50 formalin-fixed paraffin-embedded (FFPE) GC tumor tissues and 50 normal FFPE tissues were obtained from Aramesh Lab, Tehran, Iran. Histopathological examination of the tissue specimens by a pathologist confirmed the diagnosis. All the participants were Iranians. Detailed clinicopathological parameters were gathered, including age, sex, Helicobacter pylori infection, and tumor grade, stage, and size. Non-Iranian patients, as well as the patients undergoing chemotherapy or radiotherapy were eliminated from the study. American Joint Committee guidelines were used to ascertain tumor stage (22). The Ethics Committee of the Islamic Azad University of Tehran Medical Sciences, Faculty of Pharmacy and Pharmaceutical Sciences, Tehran, Iran, approved the present study (IR.IAU.PS.REC.1399.034).
3.2. Selection of Genes
For selecting target genes, previously identified lncRNAs and their molecular epidemiologic data were carefully reviewed. Regarding certain criteria, AK02339, a lncRNA, and its target, FOXO3a, were finally selected because of their proposed association with cancer.
3.3. Gene Expression
A RNeasy DSP FFPE RNA extraction kit (Qiagen Co, Germany) was employed to extract total RNA from tumor samples. Using a PCR cycler (Rotor-Gene Q MDx; Qiagen GmbH), qPCR was performed using cDNA fragments as templates to amplify the lncRNA AK023391 and FOXO3A genes using SYBR® Premix Ex Taq™ (Takara Bio, Inc.), following the manufacturer’s protocol. The experimental protocol was performed as follows: i) Thermocycling conditions included an initial activation phase at 94°C for 30 seconds, 35 cycles at 94°C for five seconds, and 35 cycles at 60°C for 35 seconds, and ii) Melting curve analysis. Primer sequences were designed by GeneRunner software for both genes. Primer-BLAST (NCBI) was employed to evaluate the specificities of the primers designed (Table 1). The B2M gene was used as a normalizer endogenous gene, and the 2-ΔΔCt method was applied to analyze qPCR data.
| Primer Name | Sequence (5’ to 3’) | Primer Size | GC% | Tm °C |
|---|---|---|---|---|
| FOXO3a | ||||
| Forward | TCTACGAGTGGATGGTGCGTT | 21 | 52.37 | 61 |
| Revers | CGACTATGCAGTGACAGGTTGTG | 23 | 52.18 | 61 |
| AK023391 | ||||
| Forward | ACCCCCATCCTAAACCCTGTAAAAC | 25 | 62.53 | 60 |
| Revers | TGTGGATTTGCTCATACTGCCCTG | 24 | 63.21 | 60 |
| B2M | ||||
| Forward | TGCTGTCTCCATGTTTGATGTATTCT | 25 | 40 | 56.98 |
| Revers | TCTCTGCTCCCCACCTCTAAGT | 22 | 54 | 57.93 |
3.4. Statistical Analysis
Data analysis was done in SPSS 21 (IBM Corp., USA), and graphs were plotted in GraphPad Prism. The unpaired t-test was used to determine statistically significant differences in gene expression, and P < 0.05 was regarded as the significance level. The association between lncRNA and FOXO3a genes’ expression was assessed via spearman correlation.
4. Results
4.1. General Descriptive Statistics
In this study, 50 patients with the final diagnosis of GC were studied, of whom 41 (82 %) were males; 10 (20 %) were smokers, and the mean age of the patients was 60.18 ± 13.65 years. Alcohol consumption was negative in all patients, but 28 (56%) of them were positive for the H. pylori infection. Regarding the 50 healthy individuals (i.e., control), the mean age was 36.72 ± 14.86 years. The patients from whom the GC tumor tissues were harvested had received no treatment. Detailed demographic characteristics have been shown in Table 2.
| Variables | Patient, No. (%) | Controls, No. (%) |
|---|---|---|
| Gender | ||
| Male | 41 (82) | 39 (78) |
| Female | 9 (18) | 11 (22) |
| Smoking | ||
| No | 40 (80) | 38 (76) |
| Yes | 10 (20) | 12 (24) |
| Disease stage | ||
| I | 9 (18) | |
| II | 2 (4) | |
| III | 21 (42) | |
| IV | 18 (36) | |
| Disease grade | ||
| I | 7 (14) | |
| II | 18 (36) | |
| III | 25 (50) | |
| Tumor size | ||
| > 5 | 21 (42) | |
| < 5 | 29 (58) | |
| H. pylori infection | ||
| Positive | 28 (56) | |
| Negative | 22 (44) |
4.2. Gene Expression of LncRNA AK023391 and FOXO3A
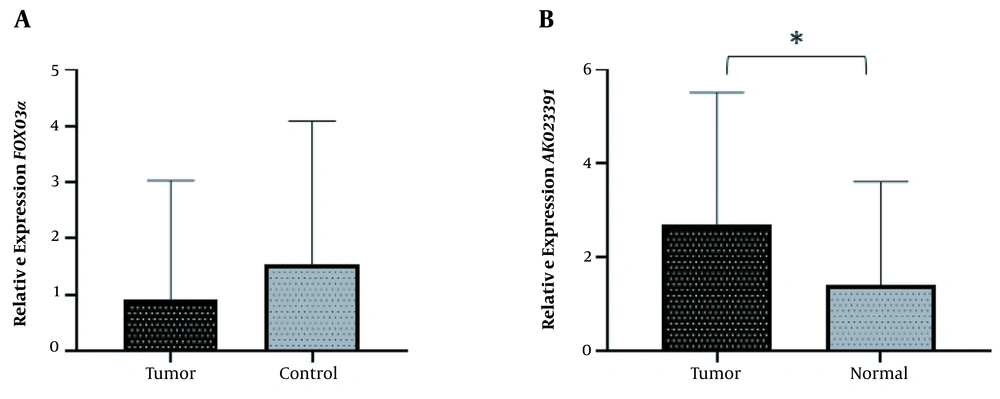
In order to investigate the role of AK023391 lncRNA and FOXO3a in GC, the expression levels of these genes were evaluated in GC tissues. No significant difference was observed in the transcription level of FOXO3a between GC and normal tissues (Mean difference = 0.9177, P = 0.2199) (Figure 1A). The expression of AK023391 was significantly up-regulated in GC tissues compared with normal tissues (1.284-fold, P = 0.0371) (Figure 1B).
4.3. Association of AK023391 LncRNA and FOXO3A Expression with Clinical Characteristics
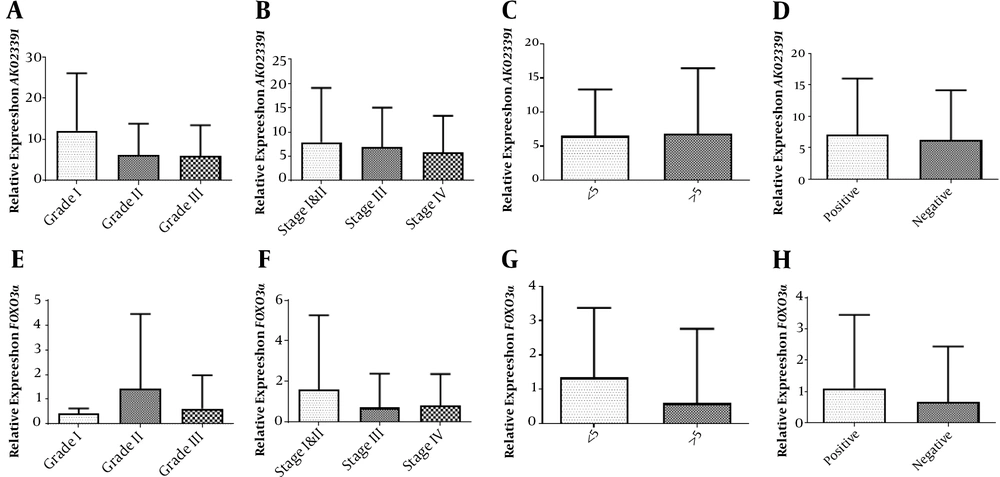
The relative expression of AK023391 demonstrated no statistically significant difference between the grad I vs. II (P = 0.2234), I vs. III (P = 0.1715), and II vs. III (P = 0.9426) disease (Figure 2A). No significant differences were identified in the transcription level of AK023391 between various disease stages [I&II vs. III (P = 0.8040), stage I&II vs. IV (P = 0.5725), and stage III vs. IV (P = 0.6544)] (Figure 2B). Moreover, no significant associations were observed between the transcription level of AK023391 and clinicopathological variables, including tumor size (< 5 & > 5 cm) and H. pylori infection (positive or negative) in GC patients (P = 0.9028 and P = 0.7205, respectively) (Figure 2C and D).
Association of the mRNA levels of the FOXO3a and AK023391 genes with clinicopathological features. (A) The transcription level of AK023391 in different grades of GC tissues, (B) The transcription level of AK023391 in different tumor stages, (C) The transcription level of AK023391 in tumors with different sizes (≤ 5 and > 5 cm), (D) The transcription level of AK023391 in patients with positive and negative H. pylori infection, (E) The relative expression of FOXO3a in different grades of GC tissues, (F) The transcription level of FOXO3a in tumors at different stages, (G) The transcription level of FOXO3a in tumors with different sizes (≤ 5 and > 5 cm), (H) The transcription level of FOXO3a in patients with positive or negative H. pylori infection.
In addition, statistical analysis revealed no significant association between FOXO3A expression and clinicopathological features or a significant difference in its expression comparing disease stage I vs. II (P = 0.5187), I vs. III (P = 0.7730), and II vs. III (P = 0.2395) (Figure 2E). There were no significant associations between the transcription level of FOXO3a and disease stage [I&II vs. III (P = 0.3663), I&II vs. IV (P = 0.4268), and III vs. IV (P = 0.8565), Figure 2F]. Also, the transcription level of FOXO3a was not associated with clinicopathological variables, including tumor size (< 5 & > 5 cm) and H. pylori infection (positive or negative) in GC patients (P = 0.2327 and P = 0.4939, respectively, Figure 2G and H).
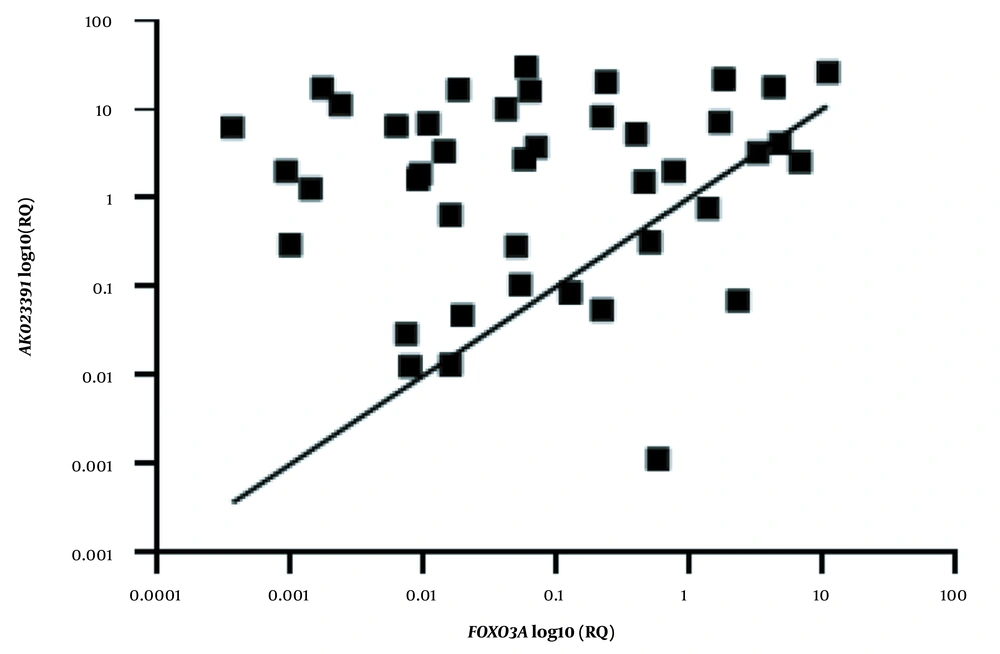
Linear regression analysis was performed to evaluate the correlation between the expression levels of the AK023391 lncRNA and FOXO3a genes in GC tumor tissues. The extent of mRNA expression is presented as log10 transformed values. The R and P values are presented for each analysis. RQ, relative quantification.
4.4. The Transcription Levels of AK023391 LncRNA and FOXO3a
Our results revealed that there was no significant correlation between the transcription levels of AK023391 and FOXO3a in GC tissues (R = 0.014; P = 0.34, Figure 3).
5. Discussion
It has been previously shown that lncRNAs contribute to GC development through several signaling pathways such as AKT and MAPK (23). Furthermore, interactions between lncRNAs and their target genes seem to affect pathogenic mechanisms in GC (24). Indeed, lncRNAs have a wide range of functions, including tumor growth suppression and oncogenesis regulation, and can act through a number of pathways (25). Our results indicated that AK023391 expression was significantly up-regulated in GC tissues compared with normal tissues. However, no significant association was found between AK023391 expression and clinicopathologic features. Similarly, Huang et al. reported that the expression level of AK023391 lncRNA was upregulated in GC tissues, which was positively associated with poor survival in GC patients (16). Likewise, AK023391 lncRNA has been reported to play a major role in cancer cell proliferation, development, and migration, as well as in tumor development and invasion by activating the PI3K/Akt signaling pathway (16). Previous studies have shown that AK023391 activates the PI3K/Akt pathway, resulting in GC tumorigenesis and invasion, accompanied by p53 downregulation, BCL-6, cyclinB1/G2, and c-myb elevated expressions, FOXO3a inactivation, and NF-κB induction; all of which contribute to GC progression. Thus, the AKT pathway is involved in GC tumorigenesis and metastasis (16, 26, 27).
However, in the present study, no significant correlation was observed between the expression levels of AK023391 lncRNA and FOXO3a. The results also showed no significant difference in FOXO3A expression between GC and normal tissues. In addition, FOXO3a expression was not associated with disease grade, disease stage, tumor size, and H. pylori infection. As a member of forkhead (FOX) transcription factors, FOXO3a regulates a variety of cellular functions such as cell death, proliferation, cell cycle progression, DNA damage, and tumor progression (17). Functional studies have shown that FOXO3a acts as a tumor suppressor in various cancers. Previous studies have shown that the down-regulation of FOXO3a was linked with a poor prognosis in glioma (28) and ovarian cancer (29) while its up-regulation correlated with a poor prognosis in triple-negative breast cancer (30), glioblastoma (31), hepatocellular carcinoma (32), and GC (22). By upregulating cathepsin L, FOXO3a overexpression facilitated GC tumor cells’ migration and invasion. The migration and invasion of GC cells were inhibited by knocking down FOXO3a, accompanied by cathepsin L downregulation (33). Furthermore, RUNX3 collaborates the complex of FoxO3a/FKHRL1 to induce apoptosis in GC cells by activating Bim promoter (34). Previous studies have also shown that AK023391 lncRNA and FOXO3a play different roles in different cancers. Further studies on the current topic are recommended.