1. Background
Metabolic syndrome (MeS) is described as a combination of clinical disorders that increase the risk of obesity (central adiposity), insulin resistance, glucose intolerance, dyslipidemia, non-alcoholic fatty liver disease and cardiovascular diseases including atherosclerosis, stroke and hypertension (1, 2). During the past decades the prevalence of metabolic syndrome is increasing dramatically worldwide, and is becoming an important health problem (3, 4). The etiology of MeS is unknown and is considered to be the result of interaction between genetic and environmental factors. Mitochondrial transcription factor A (TFAM) is involved in the maintenance of the mitochondrial genome. TFAM gene is mapped on chromosome 10q21.1. TFAM plays an important role in direct regulation of mitochondrial DNA (mtDNA) copy number, affecting transcription initiation and replication, which indicates that TFAM is essential for the maintenance of mtDNA (5). TFAM is a nuclear-encoded protein of 246 amino acids (25 kDa) with a mitochondrial targeting pre-sequence of 42 amino acids (6). TFAM was initially recognized as a transcriptional activator of mitochondrial DNA (mtDNA) but latter, it was found to be crucial for mtDNA replication (7). Consequently, TFAM is considered a key regulator of mtDNA transcription and replication. Also, there are several lines of evidence proposing that TFAM regulates the transcription and replication of mtDNA in vivo. First, disruption of the TFAM gene in mice causes major cellular dysfunction, embryonic lethality, and mitochondrial diabetes resulting from mtDNA depletion and loss of oxidative phosphorylation capacity (5, 8). Second, TFAM levels are responsive to the amounts of mtDNA in the cell, since it is present in low amount in rho-zero cells lacking mtDNA (9). Third, differences in mitochondrial transcriptional activity and mtDNA synthesis, correlate with the relative amounts of TFAM (10, 11). Mitochondrial DNA (mtDNA) content dropped in an age-dependent manner and may be one of the causal factors in age-related type 2 diabetes (12). It has been suggested that age-related alterations of oxidative stress may affect mtDNA replication via regulating TFAM activity (12). It has been suggested that TFAM promoter methylation might play a role in the pathogenesis of insulin resistance. There is little information regarding the effect of TFAM promoter methylation on MeS.
2. Objectives
In the current study, we aimed to evaluate the association between TFAM promoter methylation and metabolic syndrome in a sample of the Iranian population.
3. Patients and Methods
This case-control study was performed on 151 patients with and 149 without MeS. MeS was defined using the national cholesterol education program adult treatment panel III (NCEP ATP III) criteria (expert panel on Detection and Treatment of High Blood Cholesterol in Adults, 2001) as described previously (4, 13). Ethical approvals were obtained from Ethics Committee of Zahedan University of Medical Sciences, and informed consent was obtained from all individuals. data including weight, height, waist circumference, systolic and diastolic blood pressures; blood levels of glucose, triglycerides, total cholesterol, HDL cholesterol and LDL cholesterol were collected as described previously (4, 13). Blood samples were collected in EDTA-containing tubes and genomic DNA was extracted using salting out method as described previously (14).
3.1. TFAM Promoter Methylation
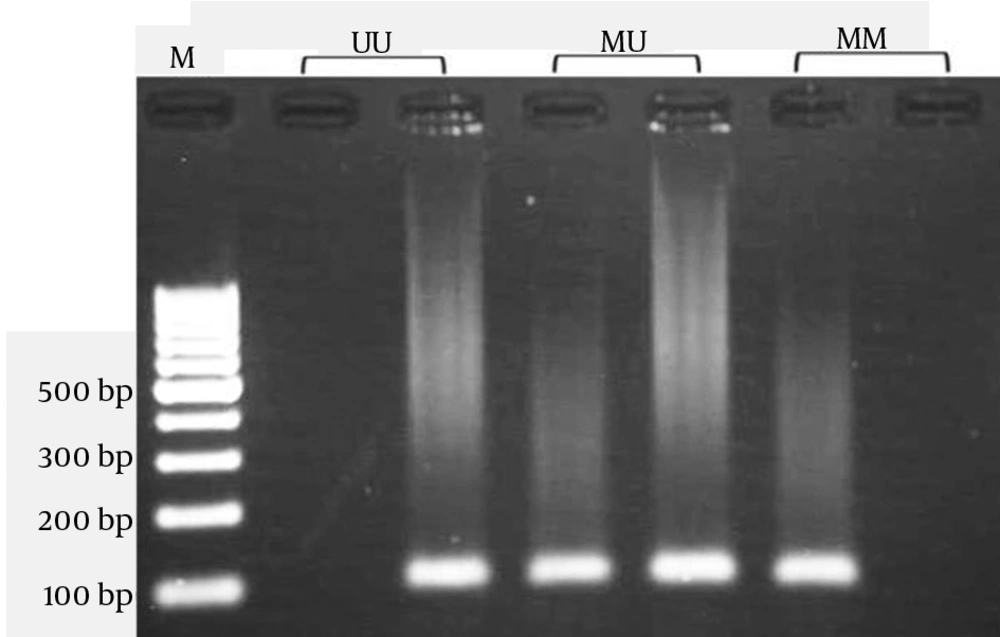
DNA methylation of CpG dinucleotides affects the expression of many genes (15). The 2378 bp long human mtTFA promoter contains 67 CpG dinucleotides, the so-called CpG islands. The CpG sites have been known to be methylation sites during genomic imprinting and gene regulation. The DNA samples were treated with sodium bisulfite, which converts unmethylated C to U. However, once methylation occurs at C residues, they will resist the treatment. Bisulfite treatment of DNA was done as described previously (16). We developed a nested methylation-specific PCR (nested MSP) method for detection the promoter methylation of TFAM that increased MSP sensitivity. First-stage PCR primers in the nested MSP recognized a bisulfite-treated template but did not discriminate between methylated and unmethylated alleles. In the second stage, two pairs of primers are used; one pair of primers is specific for an unmethylated template, and the other pair is specific for a methylated template. The forward and reverse primers for the first stage were 5ʹ-GTAAGTTGGAGGTTAGATTGAAAG -3ʹ and 5ʹ- ATAAAATCTACATTCCAACCCC-3, respectively, producing a 963 bp amplicon that was used as template for the second PCR stage. The second stage was done as described by Gemma et al. (17). Primer sequences used to amplify an unmethylated product were 5ʹ-TAATGGGTTTTATATAGATATATGG-3ʹ (sense) and 5ʹ-CAAAAATAATAACAAAAAAAACAAA-3ʹ (antisense), and primer sequences for the methylated reaction were 5ʹ-TTAATGGGTTTTATATAGATATACGG-3ʹ (sense) and 5ʹ-AAAAATAATAACGAAAAAAACGAA-3ʹ (antisense). The amplicon size was 102-bp.
Polymerase chain reaction (PCR) was performed using a commercially available PCR HotStart premix (AccuPower PCR HotStart PreMix; Bioneer Corp., Daejeon, Korea) according to the manufacturer's instructions. Briefly, 1μL modified DNA, 1μL each primer (10pmol/mL), and 17μL DNase-free water were added to AccuPower PCR PreMix. The PCR cycling conditions were as follow; 95˚C for 10 minutes, and 35 cycles containing a 95˚C for 30 seconds, 57˚C for 30 seconds, and 72˚C for 30 seconds with a final extension at 72˚C for 10 minutes. The PCR products were visualized on 2% agarose gel containing ethidium bromide and photograph was taken (Figure 1).
3.2. Statistical Analysis
The differences between the variables were assessed by Chi-square test or T-tests. The association between genotypes and metabolic syndrome was assessed by computing the odds ratio (OR) and 95% confidence intervals (95% CI) from logistic regression analyses. P value less than 0.05 were considered as significant. All statistical analyses were performed using the SPSS (Statistical Package for the Social Sciences) software version18 (SPSS Inc, Chicago, IL).
4. Results
A total of 300 subjects including 151 MeS patients (50 male, 101 female; mean age: 41.98 ± 14.65 years) and 149 subjects without MeS (45 male, 104 female; mean age: 43.53 ± 15.96 years) were recruited in the study. There were no significant differences between the groups regarding gender (P=0.621) and age (P=0.382). Neither the overall chi-square comparison of patients with and without MeS nor the logistic regression analysis showed any association between TFAM promoter methylation and MES (Table 1).
| TFAM Promoter Methylation | MeS | OR (95%CI) c | P Value | |
|---|---|---|---|---|
| Yes | No | |||
| MM | 82 (54.3) | 72 (48.3) | 1.00 | - |
| MU | 24 (15.9) | 38 (25.5) | 0.55 (0.30-1.01) | 0.070 |
| UU | 45 (29.8) | 39 (26.2) | 1.01 (0.59-1.73) | 0.944 |
| MU + UU | 69 (45.7) | 77 (51.7) | 0.79 (0.50-1.23) | 0.355 |
a Abbreviations: U, unmethylated; M, methylated.
b Data are presented as No. (%)
c Adjusted for sex and age
5. Discussion
Mitochondria are involved in the regulation of energy metabolism and their defects are correlated with aging and a variety of diseases including cardiovascular diseases, neurological disorders, myopathies, muscle weakness and cancer (18). In the present study, we investigated the possible association between TFAM gene promoter methylation and MeS in a sample of the Iranian population. Our findings revealed that TFAM promoter methylation was not associated with MeS. It has been reported that increased production of ROS in adipocytes with mitochondrial dysfunction involved in the down-regulation of GLUT4 and impaired insulin sensitivity (19). Gemma et al. (17) suggested that promoter TFAM methylation might play a role in the pathogenesis of insulin resistance. Sookoian et al. (20) evaluated whether promoter methylation (epigenetic factors) of peroxisome proliferator–activated receptor c coactivator 1 alpha (PPARGC1A) and TFAM in the liver are associated with peripheral insulin resistance. In non-alcoholic fatty liver disease (NAFLD) patients, they found that methylation levels of PPARGC1A promoter were correlated with homeostatic model assessment-insulin resistance (HOMA-IR) and plasma fasting insulin levels, while TFAM promoter methylation was inversely associated with fasting insulin. Metabolic syndrome is a combination of risk factors for cardiovascular disease (CVD) and type 2 diabetes mellitus (T2DM) (21). These factors include hyperglycemia, high blood pressure, dyslipidemia primarily characterized by increased levels of triglyceride and low HDL-cholesterol and obesity (particularly with abdominal localization) (4). The prevalence of MeS varies worldwide and depends in part on lifestyle, sex, age and ethnicity (4, 22). In conclusion, our findings showed no association between TFAM promoter hypermethylation and MeS. Further studies with are required to validate our findings in different ethnicities.