1. Background
Since recognizing bacteria, humans have been always looking for effective medications against infections caused by these microorganisms. In addition, bacteria have achieved effective mechanisms to eliminate antibiotic effects. Following the development of antibiotic-resistant organisms, nowadays, we have had several reports on widespread outbreaks in different wards of hospitals (1, 2). Development of drug resistance has been considered as a common situation. The treatment of this type of illness has caused many problems (3, 4). E. coli is ubiquitous and found in water, soil, plants and makes the normal flora of human and animal intestines.
This bacterium is the most important common microbial agent of urinary tract infections. It is the cause of many hospital opportunistic infections such as sepsis, gastric wounds, urethritis, and neonatal meningitis (5-7). This organism is resistant to beta-lactam antibiotics due to the acquisition of plasmids that encode broad-spectrum beta-lactamases. Based on these facts, the treatment of E. coli infection has been problematic (8).
Overuse and misuse of antibiotics along with factors such as long-term hospitalization of patients and the use of such devices such urinary and intravascular catheters has led to the formation of multi-drug resistant bacteria and increasing the pattern of drug resistance in the hospital (9, 10).
2. Objectives
With the increasing use of antibiotics and subsequent resistance development, we need to evaluate the patterns of resistance in all regions continuously. Therefore, this study aimed to evaluate the antibiotic resistance pattern of E. coli in patients referred to the hospital during the years 2016 - 2018.
3. Methods
3.1. Bacterial Isolates
In this cross-sectional study, 2250 specimens of outpatient and inpatients obtained from Gerash hospital in three years of 2016 - 2018 were studied. After collecting the specimens, they were cultured in blood agar and eosin methylene blue (EMB) agar media by standard loop and incubated in 37ºC, for 24 hours.
A sample was considered positive for urinary tract infection (UTI) in the light of the number of yielded colonies (105 cfu per milliliter) and the cytology of the urine through microscopic detection of bacteriuria and PMNs (≥ 8 leukocytes/mm3). However, lower colony counts associated with significant pyuria or low PMN count associated with significant colony counts was considered and analyzed in the light of the clinical picture and the patient’s immunological status (11). Also, the study excluded patients if they were pregnant, planned to become pregnant for woman, had used systemic antimicrobial agents within 14 days, had a chronic illness, or had a known anatomical or functional abnormality of the urinary tract. Afterwards the genus and species of isolates were determined based on standard methods and biochemical characteristics of isolates, such as oxidase test, nitrate revitalization, Simmon’s citrate, TSI reaction, urease, indole, movement, MR/VP and lysine decarboxylase and finally all E. coli isolates were confirmed using API20E kit (bioMerieux SA, Marcy l’Etoile, France).
3.2. Antimicrobial Susceptibility Testing
Among the 1642 positive samples, 1067 nonduplicate cases were diagnosed as E. coli infection. In this study, patterns of antimicrobial resistance were studied using the disk diffusion method. Isolates were cultured on the Mueller Hinton agar medium (Merck, Germany) for susceptibility test against several types of antibiotics with appropriate disks containing ceftriaxone (30 μg), gentamicin(10 μg), ciprofloxacin (5 μg), cefixime (30 μg), amikacin (30 μg), nitrofurantoin (300 μg), nalidixic acid (30 μg), and, trimethoprim-sulfamethoxazole (1.25/23.75 µg) antibiotics (MAST, Merseyside, UK), according to the recommendation of the Clinical Laboratory Standards Institute (CLSI) (12). The strain E. coli ATCC 25922 and P. aeruginosa ATCC 27853 were used as positive control.
3.3. Statistical Analysis
SPSS version 22 (IBM, Chicago, IL, USA) was used to analyze the data and a P value of < 0.05 considered to indicate statistical significance.
4. Results
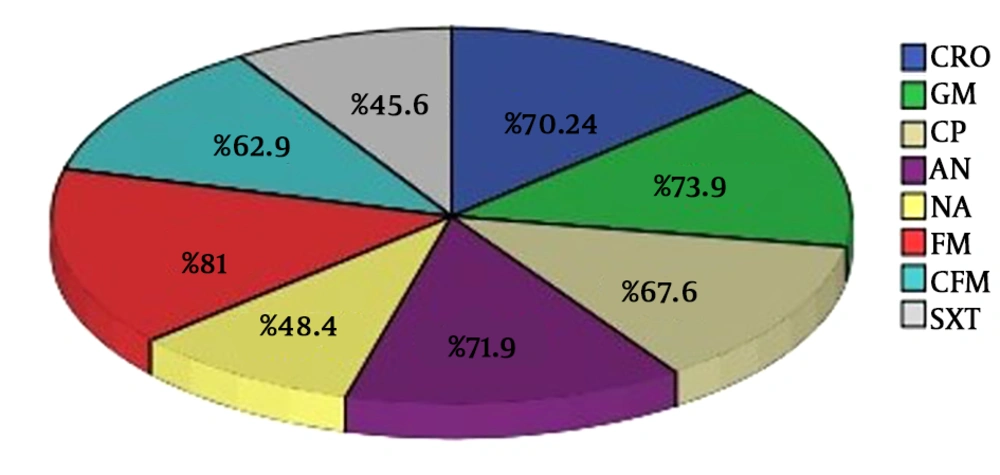
In this center, during the years of 2016 - 2018, 2250 samples were submitted for cultivation, 1642 positive cultivations were obtained, based on the bacteriology results, of which 1067 (65%) were from E. coli strain. Of the total positive cultures for E. coli, 241 samples were related to men (22.6%) and 826 samples belonged to women (77.4%). The age range of patients in this study was 1 - 100 years. Based on the findings, the highest sensitivity seen in antibiotics nitrofurantoin (81%), gentamicin (73.9%), amikacin (71.9%), ceftriaxone (70.24%), ciprofloxacin (67.6%), cefixime (62.9%), nalidixic acid (48.4%), trimethoprim-sulfamethoxazole (46.5%) (in that order) (Figure 1).
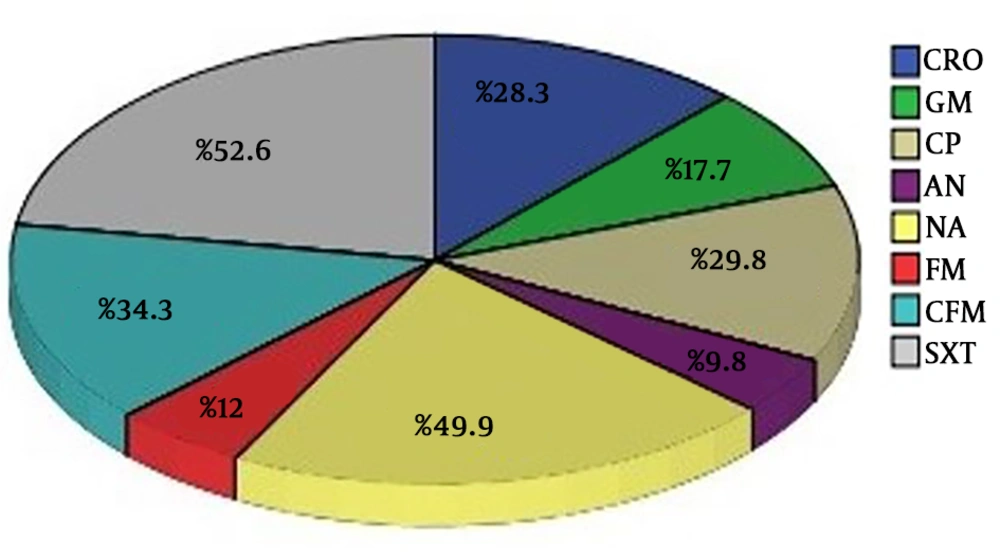
On the other hand, the maximum resistance rates were observed in trimethoprim-sulfamethoxazole (52.6%), nalidixic acid (49.4%), cefixime (34.3%), ciprofloxacin (29.8%), ceftriaxone (28.3%), gentamicin (17.7%), nitrofurantoin (12%) and amikacin antibiotics (9.8%) respectively (Figure 2).
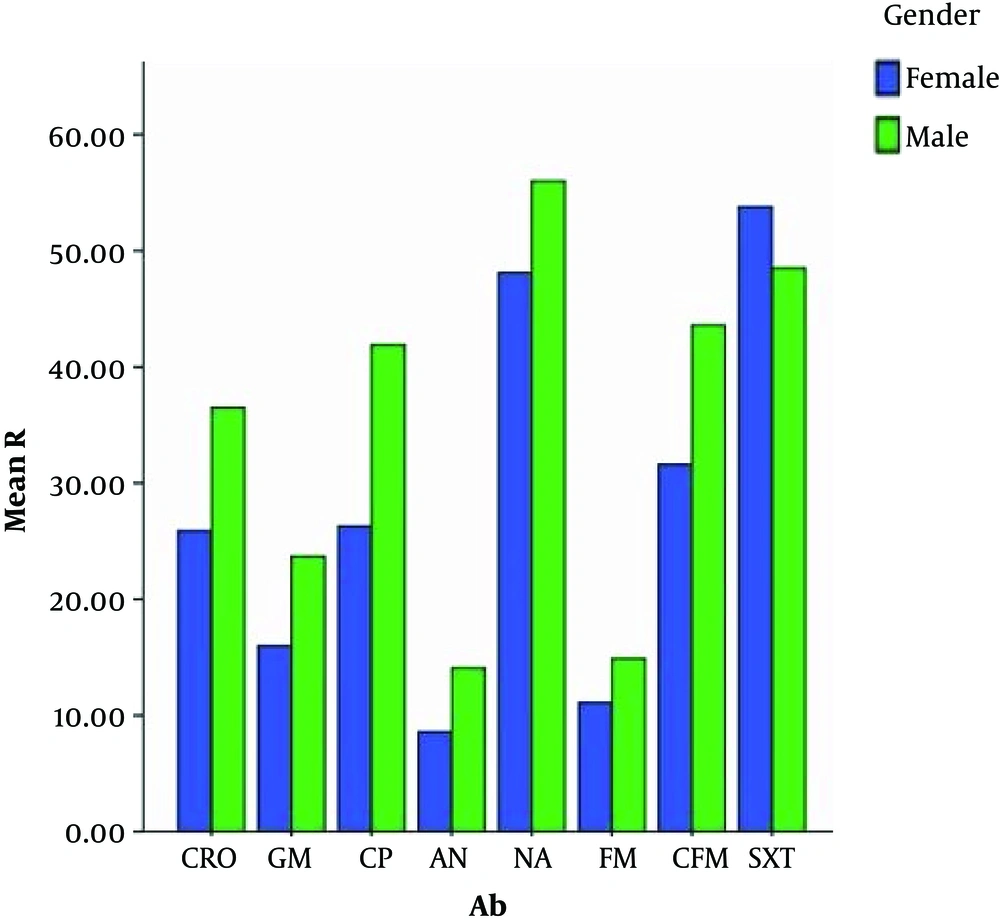
In the current study, via a chi-square test, a comparison was made between the percentage of resistance of different antibiotics among women and men. It was identified that the antibiotic resistance percentages of cefixime, ciprofloxacin, ceftriaxone, gentamicin, and amikacin had a significant difference between women and men. (P value < 0.05). Antibiotic resistance in men was higher than in women (Figure 3).
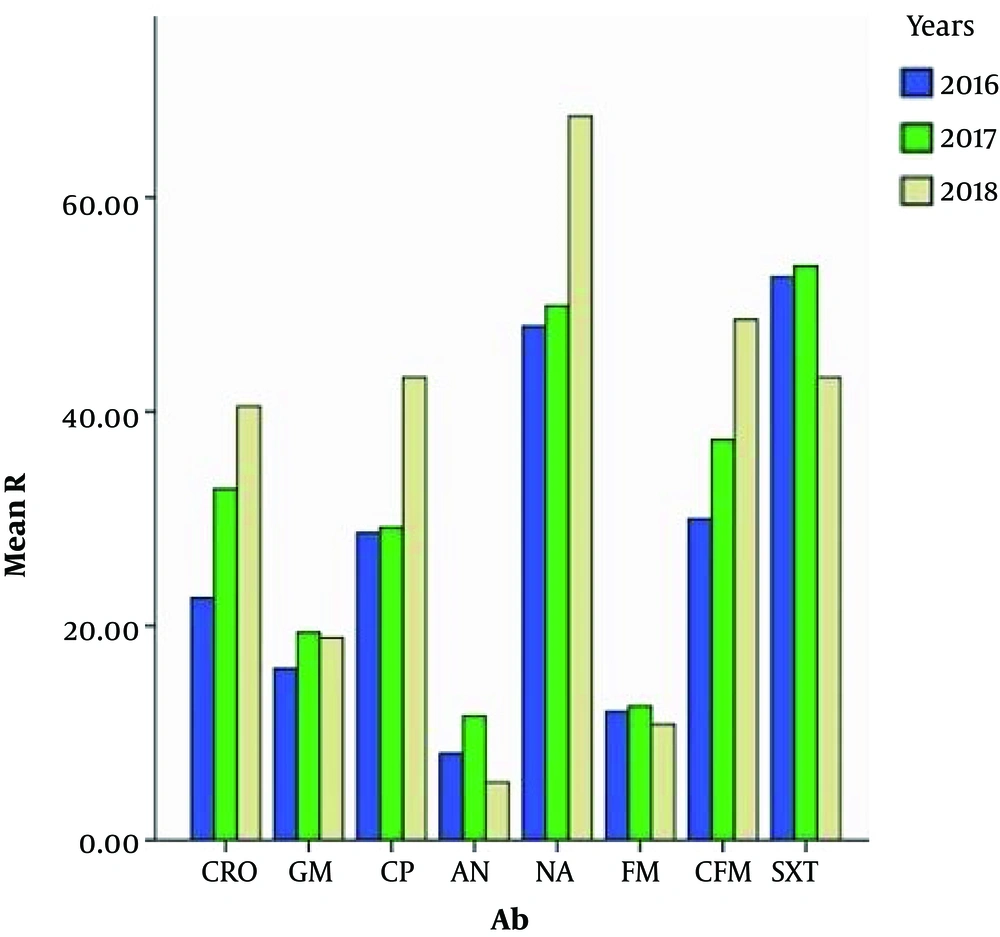
Also, through this test a significant relationship (P value < 0.05) was observed between the resistance of cefixime, ciprofloxacin, nalidixic acid, ceftriaxone, gentamicin and amikacin antibiotics and different years (2016 - 2018). Therefore, with increasing years, we observed an increase in resistance (Figure 4).
In this study, the relationship between antibiotic resistance and different age groups (1 - 100) was studied. According to the statistical analyses, a significant relationship was observed between the resistance of cefixime, ciprofloxacin, ceftriaxone, gentamicin and nalidixic acid antibiotics and the age of people, such that the resistance of cefixime, ciprofloxacin, ceftriaxone, gentamicin, and antibiotics was highest in the age group of 81 - 90 and resistance to nalidixic acid in the age group of 91 - 100 has 100 percent resistant (Table 1).
| Age | CRO | GM | CP | AN | NA | FM | CFM | SXT |
|---|---|---|---|---|---|---|---|---|
| 1 - 10 | 30.3 | 17 | 13.3 | 12.1 | 44.8 | 15.2 | 36.4 | 60 |
| 11 - 20 | 23 | 11.5 | 18 | 16.4 | 42.6 | 6.6 | 23 | 54.1 |
| 21 - 30 | 22.9 | 14.5 | 21.8 | 8.9 | 44.7 | 11.2 | 30.2 | 48.6 |
| 31 - 40 | 23.6 | 15.5 | 29.7 | 7.4 | 45.3 | 12.2 | 31.8 | 47.3 |
| 41 - 50 | 21.2 | 18.2 | 22 | 7.6 | 41.7 | 7.6 | 28 | 53 |
| 51 - 60 | 23.4 | 10.9 | 40.6 | 9.4 | 58.6 | 10.2 | 25 | 46.1 |
| 61 - 70 | 42 | 26.1 | 38.6 | 6.8 | 56.8 | 14.8 | 45.5 | 54.5 |
| 71 - 80 | 33.3 | 25.6 | 47.8 | 14.4 | 57.8 | 18.9 | 43.3 | 55.6 |
| 81 - 90 | 50 | 30.9 | 58.8 | 10.3 | 67.6 | 8.8 | 57.4 | 57.4 |
| 91 - 100 | 42.9 | 28.6 | 57.1 | 28.6 | 100 | 28.6 | 57.1 | 71.4 |
Antibiotic Resistance (%) by Age Group
5. Discussion
In this study, the most resistance is against trimethoprim-sulfamethoxazole antibiotics (52.6%) and the highest rate of sensitivity is related to nitrofurantoin (81%) which is consistent with the study of Mukherjee et al. and similar to the antimicrobial resistance of E. coli in the year 2018. In the Mukherjee study, the highest rate of sensitivity was related to nitrofurantoin (72.5%) (13). Resistance to antibiotics, among pathogenic bacteria, is a topic that has been considered today as a global problem. E. coli is one of the most important pathogens which shows resistance against common antibiotics (14, 15). In the study carried out in India by Sharma et al. (16) in order to evaluate the antimicrobial resistance of E. coli during 2012 - 2014, it was found that the highest rate of bacteria isolated from patients was E. coli with (67.66%). In this research, E. coli has shown the most rate of sensitivity to nitrofurantoin. Reduced sensitivity to nitrofurantoin is consonant with a recent study from 1.36 % in 2016 to 15.18 % in 2018.
The results of Barour el al. (17), showed that among the 198 E. coli strains, elevated resistance rates were related to ampicillin (59.09%) and tetracycline (43.43%), and low resistance rates were observed for nalidixic acid (8.08%), ciprofloxacin (7.07%), kanamycin (6.56%), cefotaxime (4.54%), chloramphenicol (4.04%), nitrofurantoin (2.52%), cefoxitin (2.02%), gentamycin (1.01%), and however did not find colistin resistance isolates. Their results were accordance with our finding.
Sharma et al. (16) indicated that the rate of resistance of 3rd generation cephalosporins had been accompanied by an increase of 15% in the years of 2012 - 2014. In our study, we have used 3rd generation cephalosporins such as cefixime and ceftriaxone, which increased the resistance of both antibiotics over the years have been reported. The rate of the resistance of E. coli against trimethoprim-sulfamethoxazole in the new study has risen over the three years, and it was nearly 100 percent, while in our study the rate of resistance had been 52.6% in 2016, 53.6% in 2017 and 40.3% in 2018. Similar to our results, a study from Iran showed that among 100 E. coli isolates collected from different clinical specimens, the maximum resistance was related to amoxicillin (96%), ceftriaxone (78%), cefazolin (76%), co-trimoxazole (70%), cefotaxime (67%) and ceftazidime (66%) (18). The observation of drug resistance among E. coli strains in this study is alarming, and this could be attributed to the indiscriminate and extensive use of antibiotics, particularly beta-lactam antibiotics.
In another study performed by Sanchez et al. (19) in order to evaluate the E. coli antimicrobial resistance in 2002 - 2010, it turned out that ciprofloxacin resistance increased from 3.40 to 17.160 and trimethoprim-sulfamethoxazole resistance had increased from 17.9% to 24.26%. The ascending of resistance increase in the Sanchez study is consistent with the recent study. The incidence rate of resistance, an increasing trend in ceftriaxone, ciprofloxacin, nalidixic acid and cefixime and downtrend in nitrofurantoin, gentamicin, amikacin, and trimethoprim-sulfamethoxazole indicated that the resistance is contradictory with trimethoprim-sulfamethoxazole with the Sanchez study. In the study published in 2008 by Dallal et al. (20) of the 188 examined E. coli strains , the highest antimicrobial resistance respectively is related to cotrimoxazole (72%), nalidixic acid (69%), ciprofloxacin (99%) and gentamicin 31% which correspond with the rate of resistance in the current study that includes trimethoprim-sulfamethoxazole (52.6%), nalidixic acid (49.9%), ciprofloxacin (29.8%) and gentamicin (17.7%). But the rate of resistance in that was found in India in 2014 with the title: “E. coli antimicrobial resistance”, the highest resistance included ceftriaxone (71.4%), cotrimoxazole (64.2%), and the highest sensitivity is for amikacin (82.6%), nitrofurantoin (78.2%). The rate of ceftriaxone resistance contradicts with the recent study. Somashekara et al. (21) during a study in 2011 found that the highest rate of E. coli microbial resistance is related to nalidixic acid (78.8%), trimethoprim-sulfamethoxazole (75.2%), ciprofloxacin (57.8%) and amikacin (26%) antibiotics while in our study highest rate of resistance was firstly related to trimethoprim-sulfamethoxazole (52.6%) and after that nalidixic acid with 49.4%. Also through this study, the significant relationship (P value < 0.05) was observed between the resistance of cefixime, ciprofloxacin, nalidixic acid, ceftriaxone, gentamicin and amikacin antibiotics and different years (91 - 93). Therefore with increasing age, we have increase in the resistance. In this study, the relationship between antibiotic resistance and different age groups (1 - 100) was studied. According to the statistical analyses, a significant relationship was observed between the resistance of cefixime, ciprofloxacin, ceftriaxone, gentamicin and nalidixic acid antibiotics and the age of people, such that the resistance of cefixime, ciprofloxacin, ceftriaxone, gentamicin, and antibiotics was highest in the age group of 81 - 90 and resistance to nalidixic acid antibiotic in the age group of 91 - 100 has 100 percent resistant.
5.1. Conclusions
The results of this study showed that antibiotic resistance patterns exist in different regions, and the rate of this resistance is increasing continuously. This pattern is different from region to region due to the geographical disparity, district hygiene level, and the rate of arbitrary administration of antibiotics in different areas. The emersion of resistance to new antibiotics is a matter to be taken seriously, therefore, continuous evaluation of bacteriology, and correct and rapid diagnosis of antibiotic resistance, followed by identifying the correct line of treatment can make a significant contribution to preventing treatment failure, performing quicker treatment and saving time and cost to the patients. By recognition of resistance patterns in each area, we can prevent the resistant gene spread, and help physicians to cure patients more precisely.