1. Background
Chronic hepatitis C virus (HCV) infection affects about 3% of human population worldwide and has become an important leading factor to liver fibrosis, liver cirrhosis, and hepatocellular carcinoma (HCC) (1). About 30% to 40% of patients with chronic hepatitis C (CHC) develop iron overload (2-5), which increases both progression of liver fibrosis and risk of carcinogenesis (6-8). In experimental studies, expression of HCV core or nonstructural proteins has been associated with low hepcidin synthesis, which was not related to HCV genotypes (9, 10).
Hepcidin is synthesized mainly in hepatocytes and is an important regulator of iron homeostasis (11). Specific conjugation of hepcidin with ferroportin leads to the degradation of ferroportin molecule and inhibits free iron efflux from cells (12). Increase in serum and hepatic iron concentration or inflammation are the strongest inducers of hepcidin expression. Ischemia, iron deficiency, and stimulation of erythropoiesis are the most important inhibitors of hepcidin production (13). The lack of hepcidin results in retained activity of ferroportin and finally affects modulation of iron export from cells.
Hepcidin response to inflammation is mainly mediated by interleukin 6 (IL-6) which stimulates hepcidin transcription through signal transducer and activator of transcription 3 (STAT3) (14). Another pathway that activates transcription of the hepcidin gene in response to iron stimulus includes induction of bone morphogenetic protein (BMP) signaling with phosphorylation of intracellular Smad proteins. This pathway is dependent of hemojuvelin (HJV) as a co-receptor (15). The third possible pathway of modulation of hepcidin expression in response to blood iron levels is related to the role of high-Fe (HFE) protein, which is not clearly elucidated. It is suggested that HFE protein might be associated with proper signaling response, mediated with BMPs. Another hypothesis indicates the role of an iron-sensing complex, consisted of transferrin receptor 2 (TfR2) and HFE, which might modulate hepcidin expression (16).
The analysis of hepcidin expression in experimental models of HCV replication disclosed a possible involvement of the oxidative stress in the induction of iron accumulation (9, 10). Increase in hepcidin concentration is expected in infections because it is a peptide with antimicrobial activity and an element of the host innate antiviral immunity (17). In this context, hepcidin inhibition becomes prognostically unfavorable in CHC and might be associated with a poor response to the antiviral treatment. Observations made on cell lines are contradictory as there is an ambiguous opinion on the effect of iron on the intensity of HCV replication (18-21). Bartolomei et al. found that knock-down of hepcidin increased intracellular ferritin and inhibited HCV replication (22). On the other hand, Liu et al. proved that hepcidin exhibits antiviral activity against HCV (23). The use of hepcidin as a factor with the potency to modulate the efficacy of antiviral therapy seems to be an interesting issue.
2. Objectives
The aim of this case-control study was to assess hepcidin mRNA (HAMP gene) expression in liver tissue in terms of iron metabolism parameters, HFE gene mutations, and efficacy of pegylated interferon alpha and ribavirin therapy in patients with CHC.
3. Patients and Methods
The study was conducted according to the ethical guidelines of the Helsinki Declaration. It was approved by the Local Independent Bioethics Committee at the Medical University of Gdansk (NKEB 270/2010). Informed consent was obtained from all enrolled subjects.
3.1. Patients, Clinical Diagnosis, and Laboratory Assessments
A total of 50 consecutive patients with diagnosis of CHC or chronic hepatitis B (CHB), who was qualified for antiviral therapy in Department of Infectious Diseases, Medical University of Gdansk, were recruited. Patients with CHB were recruited as the control group for those with CHC. It was planned to recruit 50 patients, including 25 to 30 subjects with CHC. Only patients who underwent liver biopsy were recruited. According to the Polish National Health Service (NFZ) recommendations for antiviral therapy, after the confirmation of CHC or CHB, 31 patients with CHC and 19 patients with CHB were enrolled. Patients with chronic liver diseases other than HCV- or HBV-related diseases or those with HBV/HCV, HCV/HIV, or HBV/HIV coinfections were excluded. We also excluded patients with a history of drug or alcohol abuse (> 25 g/d alcohol intake). We analyzed liver function tests including activity of alanine aminotransferase (ALT) and aspartate aminotransferase (AST), gamma glutamyl transpeptidase (GGT), serum bilirubin concentration, and body iron content markers including iron and ferritin concentration as well as transferrin saturation. The biochemical serum tests were done by Hitachi 912 automatic biochemical analyzer (Roche Diagnostics, Basel, Switzerland) according to manufacturer’s instructions.
The HCV and HBV infections were diagnosed based on detection in ELISA tests (Elecsys Anti-HCV Assay and HBsAg II Assay, respectively; Roche Diagnostics, Basel, Switzerland) and were confirmed by polymerase chain reaction (PCR) quantitative assays (COBAS TaqMan HCV Test v2.0; Roche Diagnostics, Basel, Switzerland) according to the manufacturer’s instructions. HCV genotyping was done by linear array assay for HCV genotyping (Roche Diagnostics, Basel, Switzerland) according to the manufacturer’s instructions. Finally, 19 out of 31 patients with CHC completed the antiviral therapy and among them, ten patients achieved sustained viral response. Patients received response-guided therapy with pegylated interferon and ribavirin according to European Association for the Study of the Liver (EASL) guidelines (24).
3.2. Histopathologic and Immunohistochemical Analysis
The liver specimens were preserved in 10% buffered formalin and routinely transferred to paraffin block. The hematoxylin and eosin, Masson’s trichrome for collagen, Gomori’s stain for reticulin, and Prussian blue for iron staining were done in all enrolled cases. Two independent pathologists, experienced in hepatopathology, assessed the inflammation activity and stages of fibrosis, iron deposits, and steatosis according to Scheuer score.
3.3. Analysis of the HFE Gene Polymorphism
Genomic DNA was extracted from peripheral blood leucocytes using a High Pure PCR Template Preparation Kit (Roche Diagnostics, Basel, Switzerland) according to the manufacturer’s instructions. Three variations in the nucleotide sequence of the HFE gene (C282Y, H63D, and S65C) were assessed by PCR and restriction fragment length polymorphism (RFLP) methods (5). The amplified PCR products were incubated for one hour at 37°C with the appropriate restriction enzymes, namely, SnaBI or RsaI (C282Y), BclI or MboI (H63D), and HinfI (S65C), analyzed in 2.5% agarose gel electrophoresis, and visualized with ethidium bromide staining.
3.4. Hepcidin mRNA Expression
Hepcidin mRNA (HAMP gene) expression was measured in fresh liver biopsy specimen after isolation of total RNA using RNeasy Mini Kit columns (Qiagen, Hilden, Germany). Only samples of A260/A280 ratio (index determining the purity of the genetic material) > 1.8 was used for the analysis. The quantification of gene’s expression was performed by real time PCR (RT-PCR). Reactions made in the LightCycler 2.0 system (Roche Applied Science, Mannheim, Germany) using two step quantitative RT-PCR by separately normalization through two stably expressed housekeeping genes, namely, beta-glucuronidase (*GUS) and phosphomannomutase I (*PMM1), for each sample as described previously (25, 26). First-strand cDNA was synthesized by reverse transcribing of 200 ng of total RNA in a final reaction volume of 40 µL using QuantiTect Reverse Transcription Kit (Qiagen, Hilden, Germany). PCR mixtures contained 1 μL of prediluted 1:2 cDNA, 1 μl of FastStart DNA Master SYBR Green I (Roche Applied Science, Mannheim, Germany), 4 mM of MgCl2, and 0.5 μM of forward and reverse primer in a final volume of 10 μL. In each assay, no template control was included. Triplicate reactions were performed in a LightCycler 2.0 system starting with polymerase activation step for ten minutes at 95°C, followed by 45 cycles of five seconds at 95°C and 15 seconds at 60°C for each primer pairs (HAMP F: 5’-GACCAGTGGCTCTGTTTTCC-3’, R: 5’-CAGGGCAGGTAGGTTCTACG-3’; GUS F: 5’-CCTGTGACCTTTGTGAGCAA-3’, R: 5’-AAACCCTGCAATCGTTTCTG-3’; and PMM1 F: 5’-AAGATCCGGGAGAAGTTCGT-3’, R: 5’-GGTCGGCAAAGATCTCAAAG-3’). The temperature transition rate was 20°C per second. Fluorescence data were acquired after each cycle. The absence of primer-dimers and unspecific products was verified after every run by melting curve analysis (65°C to 95°C) and agarose gel electrophoresis.
3.5. Statistical Analysis
Data analysis software STATISTICA version 10.0 (StatSoft Inc., Tulsa, OK, USA) was used to perform statistical analysis. Data were expressed as a mean ± standard error of means (SE) or median value. Standard error was used since the distributions of data were skewed. Analysis of differences between variations was done using Mann-Whitney U test and Spearman’s rank correlation test. The association between variables was estimated by univariate and multivariate linear regression. P value < 0.05 was considered statistically significant.
4. Results
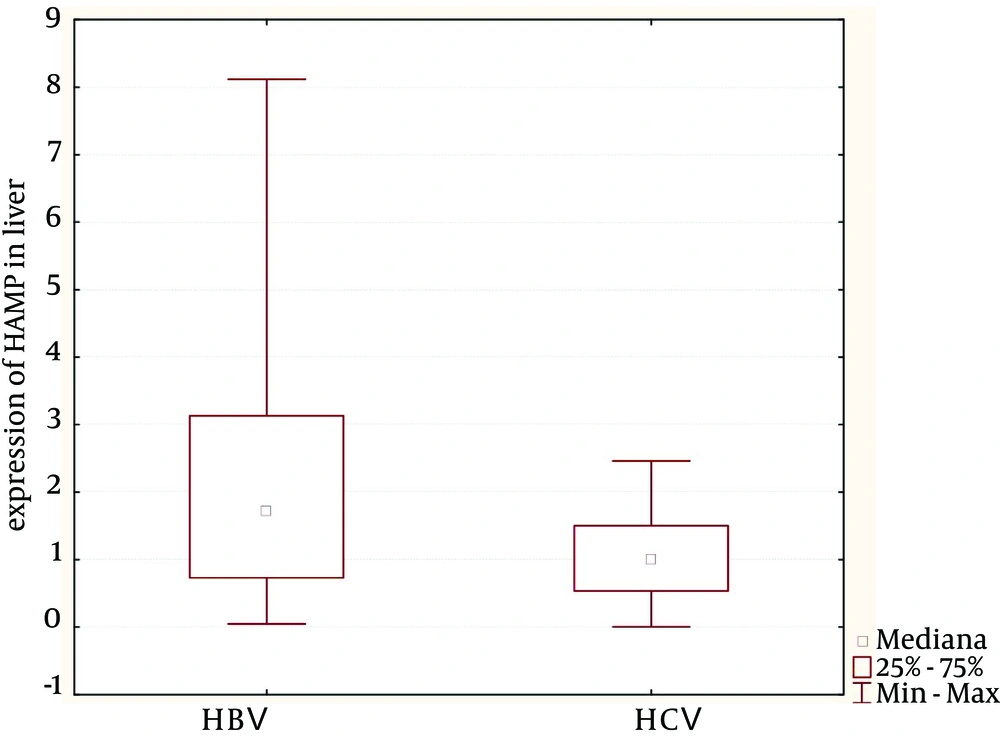
Patients with CHC were significantly older and had a higher GGT activity, more frequent iron deposits in hepatocytes, and lower mean HAMP gene expression in comparison to those with CHB (Table 1, Figure 1). Regarding genotype, 27 out of 31 patients with CHC were infected with genotype 1 HCV; mean viral load was 6.75 × 106 ± 4.05 × 106 IU/mL. Liver fibrosis and inflammation activity did not differ significantly between CHC and CHB (Table 1). In both groups, histopathologic inflammation activity was correlated with fibrosis. There was no association between inflammation activity and HAMP gene expression. More advanced fibrosis was significantly associated with higher values of transferrin saturation in those with CHC(P = 0.029). No correlation between serum indices of iron accumulation and staging of fibrosis was observed in CHB group. Iron deposits in hepatocytes were detected in minority of patients with CHC (7/31), and they were almost mild and correlated with inflammation activity (r = 0.39).
Univariate linear regression analysis revealed that in patients with CHC, HAMP gene expression was significantly dependent to ALT and AST activities as well as serum iron concentration (Table 2). On the contrary, HAMP gene expression in CHB group was significantly lower in older patients and in those with more advanced liver fibrosis. Absence of hereditary hemochromatosis-related HFE variant was correlated with higher values of HAMP expression in CHB(Table 2). Among the HFE gene mutations were detected in 12 patients with CHC and seven patients with CHB, with domination of known weak interaction with iron homeostasis (H63D) in all, except in one patient with CHC (C282Y/H63D). Their distribution was similar in both groups (Table 3). Serum iron and ferritin concentration and transferrin saturation were independent of HFE gene mutations presence. HAMP mRNA expression did not correlate with iron deposits in hepatocyte and HFE gene mutations in both groups.
Univariate linear regression analysis did not confirm significant association of HCV viral load with HAMP expression. Nevertheless, in the two-factor linear regression analysis, HCV viral load and serum iron concentration appeared to be independently associated with HAMP expression (P = 0.028 and P = 0.0003, respectively). Efficacy of assessed antiviral treatment was not significantly associated with HAMP gene expression in the studied group.
| Variable | CHC Group (n = 31) | CHB Group (n = 19) | P Value |
|---|---|---|---|
| Sex (male/female) | 11/20 | 12/7 | 0.06 |
| Age, y | 44 ± 2 | 37 ± 3 | 0.043 |
| ALT, µkat/L | 1.837 ± 0.23 | 1.67± 0.38 | 0.23 |
| GGTP, µkat/L | 1.27 ± 0.17 | 0.902± 0.20 | 0.049 |
| Bilirubin, µmol/L | 13.68± 1.54 | 15.39 ± 1.37 | 0.21 |
| Serum iron, µmol/L | 25.8 ± 2.1 | 23.2 ± 2.1 | 0.91 |
| Transferrin Saturation, % | 39 ± 4 | 36 ± 4 | 0.91 |
| Ferritin, pmol/L | 433.67± 101.12 | 685.34 ± 487.60 | 0.95 |
| HAMP Gene Expression | 1.0 ± 0.12 | 2.2 ± 0.47 | 0.039 |
| Inflammation Activity d | 2 | 2 | 0.47 |
| Liver Fibrosis d | 2 | 2 | 0.61 |
| Hepatocyte Steatosis d | 0 | 0 | 0.20 |
| Hepatocyte Iron Deposits, present/absent | 7/24 | 0/19 | 0.028 |
a Abbreviation; CHC, chronic hepatitis C; CHB, chronic hepatitis B; ALT, alanine aminotransferase; GGTP, gamma-glutamyl transpeptidase; and HAMP, hepcidin gene.
b Data are presented as mean ± SE.
c P value < 0.05 is considered as statistically significant.
d Liver histopathology data are presented as median value (score, 0-4).
| Variable | CHC Group (n = 31) | CHB Group (n = 19) | |||||||
|---|---|---|---|---|---|---|---|---|---|
| Univariate Analysis | Multivariate Analysis | Univariate Analysis | |||||||
| β Coefficient | SE | P Value | β Coefficient | SE | P Value | β Coefficient | SE | P Value | |
| Age, y | 0.10 | 0.18 | 0.6 | - | - | - | -0.46 | 0.21 | 0.047 |
| ALT, µkat/L | 0.43 | 0.17 | 0.014 | - | - | - | -0.24 | 0.23 | 0.32 |
| AST, µkat/L | 0.45 | 0.17 | 0.01 | - | - | - | -0.28 | 0.23 | 0.25 |
| Serum Iron, umol/L | 0.59 | 0.15 | 0.049 | 0.76 | 0.16 | 0.0003 | -0.26 | 0.25 | 0.32 |
| Absence of HH Alleles | -0.30 | 0.18 | 0.1 | - | - | - | 0.51 | 0.21 | 0.025 |
| Liver Fibrosis | 0.35 | 0.17 | 0.34 | - | - | - | -0.49 | 0.21 | 0.032 |
| HCV RNA, IU/L | -0.24 | 0.23 | 0.31 | -0.40 | 0.16 | 0.028 | ND | ND | ND |
a Abbreviation; HAMP, hepcidin gene; CHC, chronic hepatitis C; CHB, chronic hepatitis B; SE, standard error; ALT, alanine aminotransferase; AST, aspartate aminotransferase; HH, hereditary hemochromatosis; HCV, hepatitis C virus; and ND, not determined.
| Gene Mutations | Patients with CHC infection | Patients with CHB infection |
|---|---|---|
| C282Y/C282Y | 0 | 0 |
| C282Y/H63D | 1 | 0 |
| WT/C282Y | 1 | 1 |
| H63D/H63D | 2 | 1 |
| WT/H63D | 7 | 5 |
| S65C/S65C | 0 | 0 |
| WT/S65C | 0 | 0 |
| WT/WT | 20 | 12 |
a Abbreviation; HFE, hemochromatosis protein; CHC, chronic hepatitis C; CHB, chronic hepatitis B; and WT, wild type HFE gene.
5. Discussion
5.1. Hepcidin and Iron Overload in Chronic Hepatitis C
Different pathogenic conceptions are developed to explain iron overload in patients with CHC. Some authors find it as a surrogate of necroinflammation activity (27). Mild or moderate iron deposits in hepatocyte were present more frequently in patients with CHC than in those with CHB, despite the absence of histopathologic features of inflammation. Patients in both groups presented similar inflammatory activity and fibrosis stage, but not HAMP expression. The reduction of HAMP gene expression has been suggested to be the main reason of iron overload in CHC infection. Moreover, a few reports on HAMP expression in liver specimens or hepcidin concentrations in patients’ sera confirm its diminished synthesis in CHC (28-31).
Low expression of hepcidin in patients with CHC in our study was in agreement with some previous observations (28-30). Nevertheless, the presence and intensity of iron deposition in hepatocyte was independent of liver hepcidin expression, which was in contrast to findings of other authors. Fujita et al., Aoki et al., and Bergmann et al. reported that mild or moderate iron deposition in liver tissue or liver iron concentration correlated with hepcidin transcript levels (28, 29, 32). In our study, low hepcidin in patients with CHC , in comparison to those with CHB , was not associated with presence of iron deposits in hepatocytes. Furthermore, patients with CHC and the lowest values of HAMP expression did not accumulate iron in hepatocytes. These differences in the results of different studies might be explained by use of various methods to assess iron in the liver tissue including hepatic iron concentration, total iron score, and liver iron concentration (13).
The HAMP gene expression in CHC group depended on serum iron concentration. In other studies, association of hepcidin expression with serum iron parameters was varied. Fujita et al. reported that hepcidin was correlated with serum iron, ferritin, and transferrin saturation (28). Aoki et al. found positive correlation between hepcidin and only serum ferritin and the mean value of this protein exceeded the normal value (29). In contrary, only Abd Elmonen et al. found negative correlation of HAMP gene expression with parameters of iron stores, namely, serum ferritin and hepatic iron concentration, in Egyptian patients (30). The ethnic differences, which lead to functional changes of iron homeostasis regulatory genes or domination of other than HCV genotype 1, might explain above observations.
HAMP expression was not associated with ferritin concentration in the studied patients with CHC. Serum ferritin seems to be a good marker of tissue iron accumulation, which also did not correlate with HAMP expression in presented results. Patients with CHC who presented the lowest values of hepcidin expression did not have biochemical indicators of iron accumulation in serum. The significant association between serum iron and HAMP expression partially corresponded to the physiologic model of its induction by increasing extracellular iron content. Interaction between hepcidin and ferroportin is an element of a key system, which is responsible for maintenance of safe pool of extracellular iron and quickly and mainly responds to iron content in blood. However, regulation of hepcidin involves both circulating iron and liver iron. Lack of correlation between the HAMP expression of hepcidin and iron stores in studied patients with CHC could be explained by conducted studies on mouse models. Corradini et al. suggested differences in two ways of hepcidin regulation, depending on liver iron and serum transferrin saturation. It was proposed that liver iron stimulates bone morphogenetic 6 gene (BMP6) transcription and bone morphogenetic 6-hemojuvelin/SMAD (BMP6-HJV/SMAD) signaling pathway to upregulate hepcidin synthesis. In turn, circulating iron and increasing transferrin saturation induce activity of SMAD complex and promote hepcidin transcription without stimulation of BMP6 mRNA expression (33).
In the presented study, HAMP expression did not correlate with liver iron stores. Induction of hepcidin synthesis by BMP6-HJV/SMAD pathway, which is dependent of hepatocyte iron deposits, might be impaired in those with HCV infection. This hypothesis was developed by Eddowes et al. who suggested disruption of BMP/SMAD signaling pathway with reduction of HJV expression and suppression of hepcidin expression in the liver of patients with HCV infection (34).
5.2. Hepcidin and Liver Injury in Chronic Hepatitis C
The HAMP expression correlated with ALT activity in patients with CHC and increased in cases with more intense liver injury. Similar to Fujita et al. observations, intensity of histopathologic inflammation did not show any association with HAMP gene expression in the studied liver specimens (28). These findings might reflect differences in cytokine profile between lymphocytes in acute inflammatory reactions related to IL-6-dependent hepcidin induction and specific Th1-dependent immune cell response in CHC (28).
5.3. Hepcidin, Hepatitis C Virus Replication and Antiviral Treatment Efficacy in Chronic Hepatitis C
We confirmed other Polish observations that stated HCV genotype 1 dominates in Polish patients (35). In analyzed patients with CHC, HAMP expression in univariate analysis did not depend on viral load and genotype 1. Multivariate regression analysis revealed that HCV viremia and serum iron concentration were independently linked to HAMP expression. This association seems to be an indicator of a complex interaction between HCV replication and regulation of iron homeostasis that involved in both mechanisms of viral replication and host’s antiviral response. In our studied patients with CHC, there was no significant association of HAMP expression with an effect of treatment with pegylated interferon and ribavirin in any points of viremia control, i.e. early viral response, end of treatment response, and sustained viral response. This study was an initial assessment. Investigation in numerous study groups is warranted to clarify the association of HAMP expression with different models of antiviral response, e.g. null viral response, relapse, or even the issue of ineffective treatment due to the necessity of dose reduction due to serious adverse events.
Other authors did not compare liver HAMP expression to viral factors. In a study published by Fujita et al., the increase of hepcidin synthesis after the successful treatment with interferon and ribavirin in patients with CHC was confirmed but no correlation between HCV viral load and serum hepcidin level was found (28). This association of hepcidin with the efficacy of the antiviral treatment is interesting in the context of two issues. The first one relates to the possible influence of hepcidin on viral replication or immune response. The second one is associated with the regulatory effect of HCV proteins on hepcidin expression. Tai et al. found hepcidin as a cofactor of HCV replication in vitro. Hepcidin silencing in the full-length replicon model inhibited HCV replication (36). The same was also confirmed by Bartolomei et al. who observed inhibition of HCV replication in RNAi-mediated knock-down of hepcidin (22). On the contrary, Liu et al. found evidence for a direct antiviral activity of hepcidin against HCV replication. They recommended broadening studies on the therapeutic effect of hepcidin as an element of nonspecific immune response by postulating the suppressing effect of HCV on this regulatory peptide expression (23).
The association of hypoferremia, induced by increase of hepcidin, with significant decline in viral load during interferon therapy points to the possible involvement of hepcidin in the mechanism of antiviral response (37). This is contrary to Strnad’s results that did not confirm the role of hepcidin serum levels and its changes as predictive markers for the antiviral treatment response (38). The pathogenic role of hepcidin in terms of iron overload and course of CHC is still an open issue and needs further investigations in larger groups of patients. Based on our results, we rather suggest looking for possible therapeutic implications of decreased hepcidin, eg, muting the virus’s ability to block expression of hepcidin synthesis, which is the only factor that links hepcidin with iron overload in CHC.
In comparison with CHB, patients with CHC presented more severe liver disease with accompanying mild or moderate iron deposition in hepatocytes and decrease of HAMP expression in liver tissue. The presence of iron deposits in hepatocytes did not induce increase in HAMP. Despite the presence of active hepatitis markers, levels of HAMP expression were lower in CHC, which might be due to mechanisms that were specifically activated by HCV proteins. In this preliminary study, the evident association of decreased HAMP expression with the efficacy of treatment with pegylated interferon and ribavirin was not found. Further studies on the possible influence of hepcidin on viral replication and immune response or the regulatory effect of HCV proteins on HAMP expression are needed to assess the potency of hepcidin use in antiviral treatment.