1. Background
Breast cancer is the most common cancer and the second leading cause of death among women (1). Mammography is the primary method for examining changes in the breast tissue (2). It is recognized as the most common diagnostic method for detecting breast cancer in an early stage (3). Reduction of false positive results in cancer detection is important in therapeutic processes (4), as they can impose significant burdens on patients, including high cost, waste of time, and psychological stress (5). In developing and developed countries, the accuracy of detecting cancerous lesions is less than 50% and above 80%, respectively (6). This difference is partially due to the use of computer-aided detection (CAD) systems in developed countries.
With the advancement of medical equipment technologies, various approaches have been proposed for mammographic CAD systems (7). Since 2000, advances in diagnostic digital mammography have increased the accuracy of breast cancer detection and reduced the associated deaths (8). However, detection of suspected abnormalities is a difficult task, even for experienced radiologists. The small size of a lesion compared to the large size of a mammogram is an important dichotomy in image processing techniques for cancer detection (9). The reason for the large size of X-ray mammograms is the need for detection of very small calcification particles (10). Overall, identifying calcifications in mammograms is a subtle and time-consuming task, which causes eye strain and reduces the detection accuracy, resulting in the radiologists’ error over time (11).
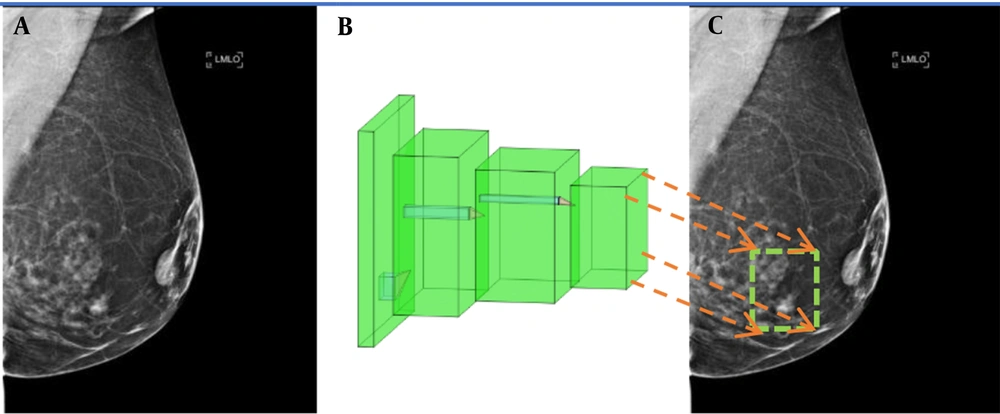
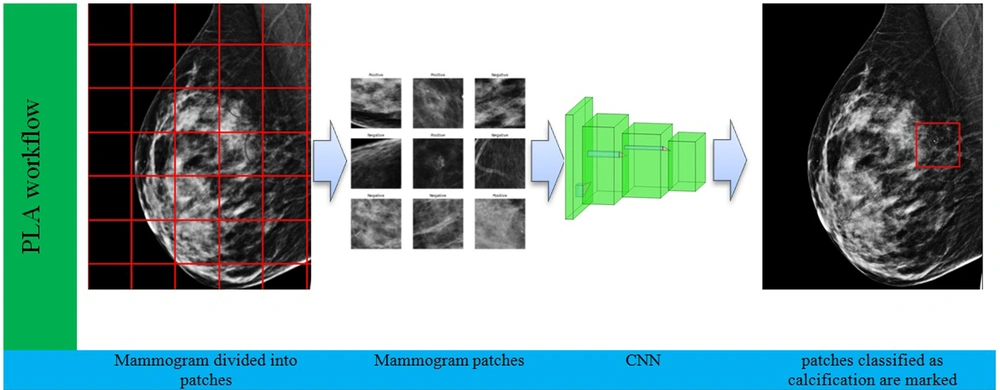
To reduce human errors, today, mammography CAD technologies are employed by radiologists to find suspected lesions. CAD systems can improve the rate of breast cancer diagnosis up to 20% (12). These systems often employ common deep learning-based region detection algorithms, including convolutional neural network (CNN)-based models, namely, region-based convolutional neural network (R CNN), Fast-R-CNN, Faster-R-CNN (12), and You Only Look Once (YOLO) algorithm (13). Figure 1 illustrates the conceptual workflow of region detection models, in which mammograms are fed into the network as input, and a box surrounding the lesion is marked at the output.
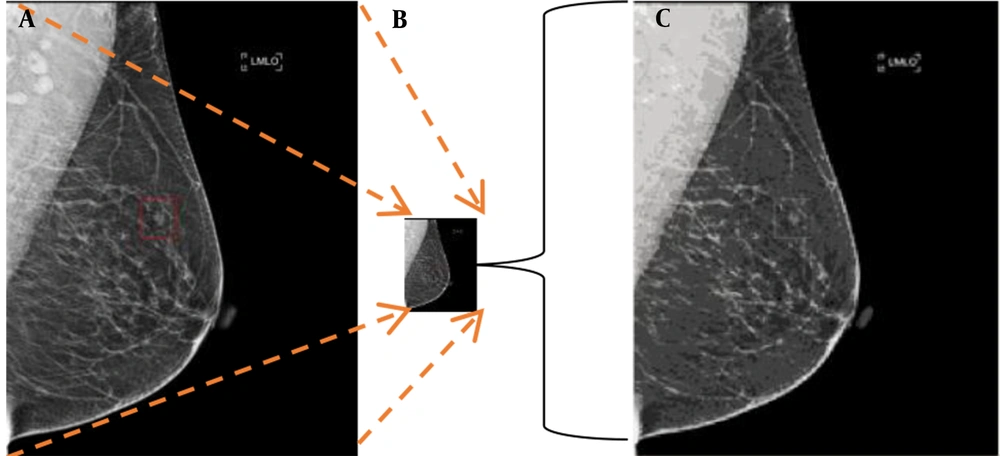
Common CNN structures, such as ResNet (14), VGG (15), and GoogleNet (16), which have been trained for object recognition on the ImageNet (17) database, are used in these region detection algorithms. The input size of these CNN structures is 224 × 224 pixels, whereas the size of X-ray mammograms is 2,560 × 3,328 (170 times larger than the CNN input size). Therefore, data preparation needs to be performed by downsampling to reduce the image dimensions. This resizing, however, results in the loss of informative pixels. However, empirically, only small patches in a mammogram contain lesions, and mammogram resizing blurs or eliminates them. An illustration of the negative impact of mammogram resizing on the CNN input is shown in Figure 2. To avoid this problem, this study proposed a novel alternative approach for detecting microcalcifications in mammograms.
2. Objectives
This study aimed to propose a CAD system for automated calcification detection in mammograms by training a model with an in-house dataset.
3. Patients and Methods
3.1. Dataset
A total of 815 mammograms were collected from 204 women (age: 51.43 ± 10.61 years), who were referred to Dr. Gity Imaging Center and Imam Khomeini Hospital Mammography Center during 2019 - 2020 (some participants were referred to these centers more than once). The left and right craniocaudal (CC) and mediolateral oblique (MLO) views were considered as separate images. Some women visited the centers more than once for screening, and for some women, only one breast or one view was collected. The mammography device was the Selenia Dimensions Mammography System (Hologic Inc., USA). This study was approved by the ethics board of Shahid Beheshti University of Medical Sciences, Tehran, Iran. Calcifications were annotated in the mammograms by specialists. The mammograms included CC and MLO views.
3.2. Patch Learning
In this study, each mammogram was divided into non-overlapping, fixed-size patches, which were used as the input to a CNN. By using this approach, the resolution of the lesion image was not deteriorated, thereby enhancing the detection accuracy. For each patch, the ground truth label was determined based on the annotations by an experienced radiologist. If a patch overlapped an annotated region, it was labelled as a calcification; otherwise, it was labelled as normal. Besides, training a CNN with patches as the input has the advantage of enlarging the training set, which improves the model training, especially considering the scarcity of mammogram datasets. We refer to the proposed approach as the patch learning approach (PLA). However, classification of malignant versus benign lesions is not within the scope of this study.
The proposed method aimed to detect and localize calcifications to help radiologists identify suspicious lesions. First, all images were divided into 224 × 224 patches. Subsequently, all patches containing calcifications were extracted from all mammograms, where 536 patches were obtained. It should be noted that if a calcification region spanned two or more adjacent patches, each of the adjacent patches was labelled as a calcification. Normal patches were extracted from all mammograms, which either contained or did not contain calcifications in other patches. Since the total number of normal patches was much higher than that of calcification patches, to achieve class size balance, a total of 536 normal patches were randomly selected from all available normal patches, and the rest of normal patches were discarded. It should be noted that an imbalance in the class size of the training set leads to classifier bias, which may degrade classification performance. Moreover, in this study, we used balanced class sizes in the test set to obtain significant performance results.
To construct the training and test sets, 100 patches, which were randomly selected from normal and calcification classes (a total of 200), were considered as the test set, while the rest of them (436 mammograms from each class) were used as the training set. The ground truth labels (based on the radiologist’s annotation) were used to train the CNN with the training set. On the test data, the CNN performed a binary classification to classify patches into normal and calcification groups. The classification accuracy was determined by comparing the CNN output with the ground truth labels.
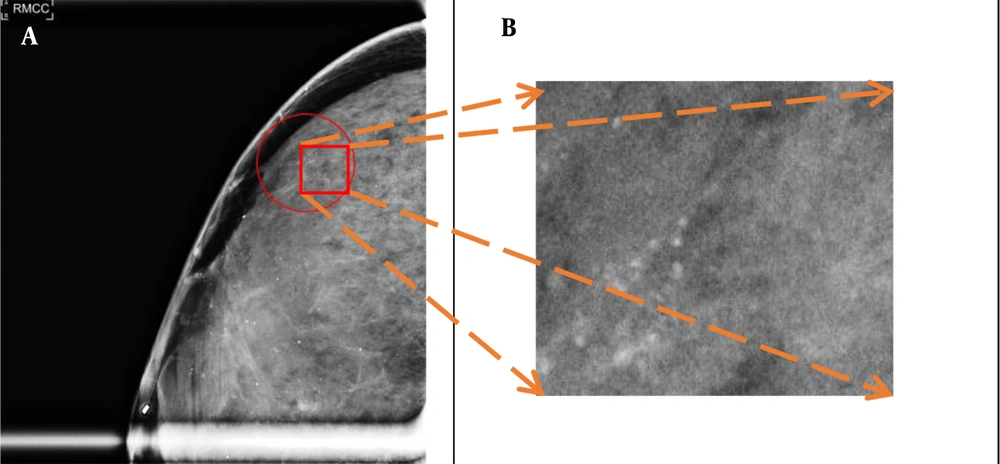
An annotated calcification in a mammogram is shown in Figure 3A, and a magnified image of the marked patch is displayed in Figure 3B. As shown in Figure 3, calcification patches often occupy less than 2% of the mammogram size; therefore, the standard procedure for resizing mammograms (reduction of image size by downsampling reduces the image resolution and results in information loss) blurs calcification spots and must be avoided.
3.3. CNN Structure
A ResNet50 model was used for the binary classification of patches with a size of 224 × 224 pixels. This model is a modern CNN, which has been trained on millions of images and achieved excellent performance in object recognition (14). Accordingly, in this study, it was selected as our pretrained network. The ResNet50 model was fine tuned with our mammogram patches by freezing the first 100 layers. The Adam optimization method, with a 0.0001 learning rate, was selected with a binary cross entropy loss function. A batch size of 32 with 120 epochs was used for training the CNN. Computation was conducted using a TensorFlow2 platform on a computer with Nvidia’ GeForce GTX 1080 GPU. The workflow of the proposed method for detecting calcifications in a new mammogram is presented in Figure 4.
The mammogram is first divided into patches with a size of 224 × 224 pixels. Next, each patch is separately fed into the convolutional neural network (CNN) for classification. The CNN classifies each patch into either normal or calcification. Finally, patches classified as calcification are marked.
4. Results
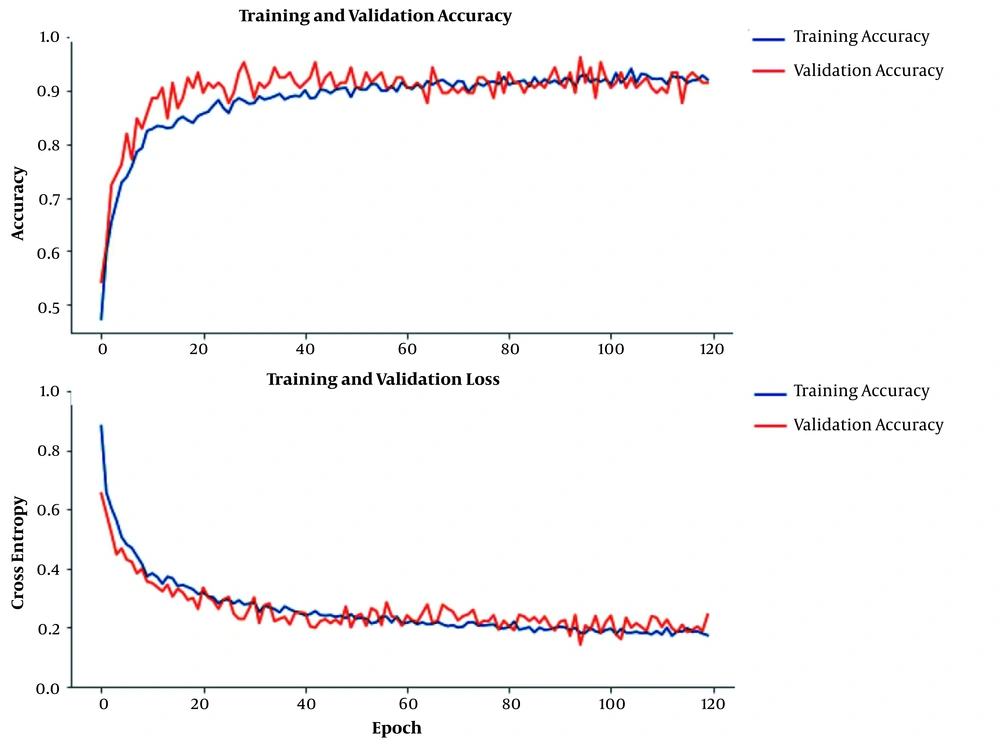
The plots of training and validation accuracy and loss during CNN training are shown in Figure 5 (5% of the training set was selected as the validation set). The close lines in the plots of training and validation indicate the CNN training efficacy. Our proposed PLA achieved an accuracy of 96.7% for the binary classification of 200 test patches. The sensitivity and specificity of PLA were 96.7% and 96.7%, respectively.
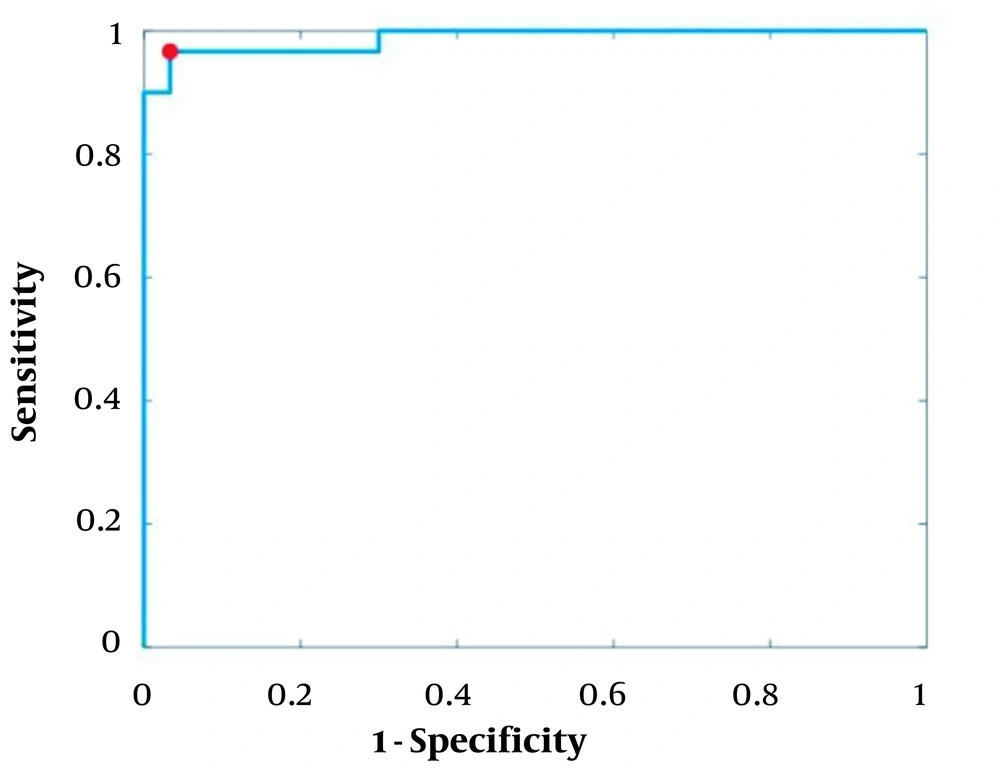
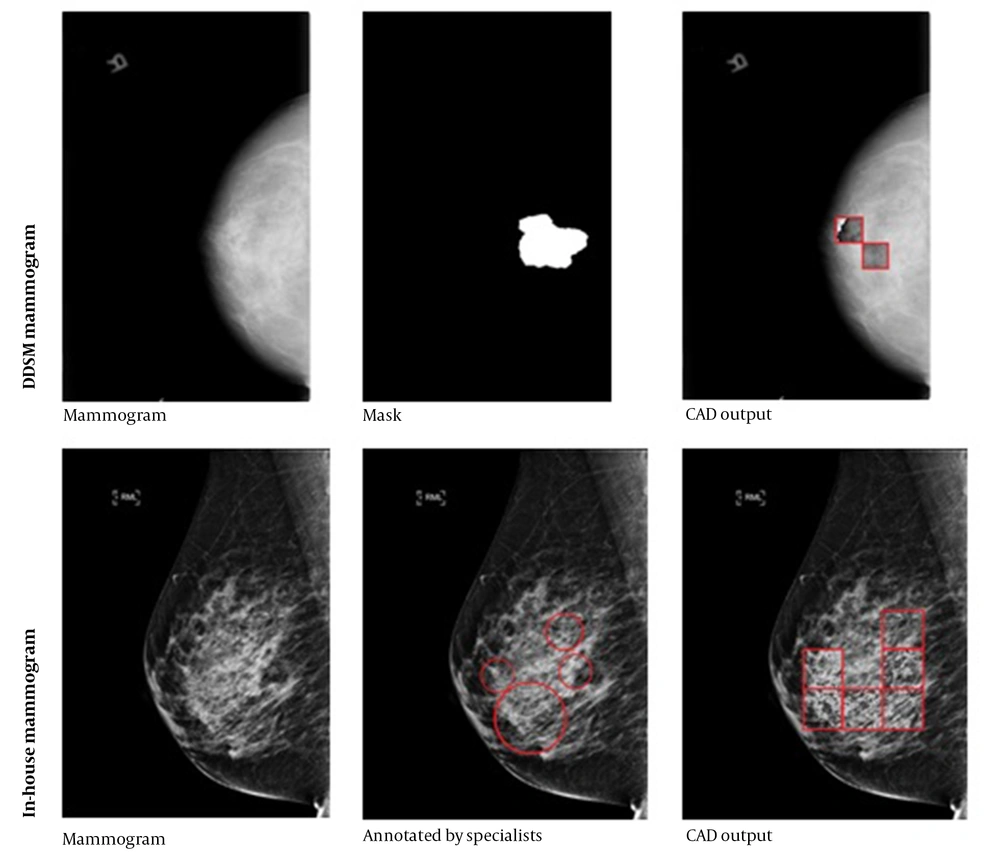
The receiver operating characteristic (ROC) curve is plotted in Figure 6, where the operating point is shown with a red dot. The area under the ROC curve (AUC) was 98.8, indicating the effective classification of the model. Generally, the operating point is selected depending on the preference, as there is a trade off between sensitivity and specificity; in other words, sensitivity can be increased at the expense of specificity, and vice versa. In this study, an operating point that maximized the classification accuracy was selected. This point was found as the intersection of the ROC curve with a line with a slope of 1 (balanced classes) and maximum y-intercept. Figure 7 indicates the performance of PLA on two representative test mammograms from the digital database for screening mammography (DDSM) dataset and our in house dataset, respectively.
The performance of the patch learning approach (PLA) on two representative test mammograms from the digital database for screening mammography (DDSM) dataset (first row) and our in house dataset (second row). The middle column shows the annotations by specialists, and the third column shows the marked patches by the proposed computer-aided detection (CAD) system.
By dividing the test set into three age ranges, including 31 - 45 years, 46 - 60 years, and 61 - 75 years, the model performance was calculated for each age range, as listed in Table 1. Based on the results, performance was approximately 97% and similar for all age groups.
| Age group (y) | Population | Classification accuracy |
|---|---|---|
| 31 - 45 | 58 | 96.4 |
| 46 - 60 | 79 | 96.6 |
| 61 - 75 | 63 | 97.3 |
5. Discussion
Daily reading of numerous mammograms, the majority of which are often normal, is a tedious task and may cause fatigue in radiologists, which in turn increases the risk of missing abnormalities. Therefore, there is a great demand for automated detection and localization of suspicious lesions, as it can avoid missing these lesions. This study proposed a method for automated detection; however, determining the malignancy of lesions is outside the scope of this study. In our future study, we will investigate a fully automated diagnosis system for determining the malignancy of detected lesions.
Recently, deep learning has been applied in medical imaging studies, such as mammography (18). Before the introduction of deep learning, other types of machine learning (ML) methods (19) were common for detecting lesions in mammograms. These ML methods require feature engineering, which is difficult and time consuming. Feature engineering refers to the process of designing and extracting relevant and useful representations from raw data. These features need to be designed by human experts. Previous studies on ML-based detection of lesions in mammograms have used features, such as wavelet (20), curvelet (20), Fourier transform (21), and edge gradient analysis (22). These features have been also used as the input to a classifier to detect or classify lesions (23). However, because it is difficult to find perfect features, the performance of ML methods is often inferior to that of deep learning methods. In deep learning, the network automatically learns useful features so that there is no need for feature engineering. Therefore, deep learning methods can be applied directly in mammograms without any preprocessing.
The CNNs are the most commonly used deep learning frameworks in mammography CAD systems, which have shown promising performance in detecting cancerous lesions. However, large datasets are needed to train CNNs, and formation of large specialist annotated mammogram datasets is expensive and time consuming. Suspected lesions in mammograms often occupy less than 2% of image pixels. Therefore, the bulk of a mammogram does not contain useful information for training a deep learning model. In previous studies, such as a study by Agarwal et al. (24), the whole image was fed into CNN models, which increased the training time substantially, while most of the data was not informative. As a solution, the proposed PLA divides the image into fixed-size patches, and only suspected patches, along with the same number of normal patches, were fed into the CNN for training. This strategy substantially reduced the training time, as only informative patches, which comprise a small percentage of each mammogram, were used for training the CNN.
The proposed PLA also has the advantage of being adaptable to various sizes of mammograms, as it operates on fixed-size patches of images. This allows the PLA to be trained with one dataset and tested by another with a different mammogram size. Table 2 lists the performance (AUC) of deep learning-based CAD systems for detecting calcifications in previous studies compared to our proposed PLA; the models and datasets used in these studies are also demonstrated. Our method outperformed these previous approaches; however, it should be noted that a direct comparison is not possible due to differences in datasets. The higher performance of our system may be attributed to the patch learning algorithm.
Abbreviations: AUC, area under the ROC curve; Mini-MIAS, Mini-Mammographic Image Analysis Society; CBIS-DDSM, curated breast imaging subset of the digital database for screening mammography; INbreast, full-field digital mammographic database; MIAS, Mammographic Image Analysis Society.
In conclusion, the results of this study highlighted the efficacy of our PLA. Future studies are suggested to focus on the application of this approach for detecting both masses and calcifications in mammograms.