1. Background
Coronavirus disease 2019 (COVID-19) has infected hundreds of thousands of people around the world and resulted in high mortality rates. According to the World Health Organization (WHO), by November 5, 2022, the number of positive COVID-19 patients was more than 637 million, and the number of deaths exceeded 6.6 million (1). Besides direct COVID-19 infection, other aspects of life were also significantly affected during the pandemic. For example, travelling was restricted, businesses declined, and other illnesses, such as depression, obsessive-compulsive disorder, and obesity, became more common due to home quarantine.
With the onset of COVID-19, there were no effective vaccines available, causing major challenges in the management of this disease in many countries. Besides, the coronavirus was constantly evolving, making it more difficult to detect new variants. Therefore, wearing masks and social distancing, besides early diagnosis and isolation of the infected, were among the best strategies to manage and prevent the spread of this virus. Since the onset of the COVID-19 epidemic, reverse transcription-polymerase chain reaction (RT-PCR) assay has been used as a standard method for COVID-19 screening. However, a sensitivity of 60 - 70%, lack of diagnosis in early stages, time-consuming process, and need for special kits and equipped laboratories to perform the tests were among the significant limitations of RT-PCR at the time (1-3). These limitations required other technologies, such as computed tomography (CT), to be used for the diagnosis and severity assessment of COVID-19 patients.
There have been significant advances in medical imaging technologies in recent years, and today, they have become standard methods for diagnosing and quantifying various diseases (4-6). Ground-glass opacity (GGO), interlobular septal thickening, and consolidation are among the leading radiological patterns of COVID-19, which differentiate these patients from healthy individuals and other types of pneumonia (7). A study on 1014 patients in Wuhan, China, reported a sensitivity of over 97% for chest CT imaging to diagnose COVID-19 (3). Therefore, with the limitations of RT-PCR at the time of this study, chest CT scan could be considered a more effective, practical, and rapid method for diagnosing and assessing COVID-19, particularly in areas most affected by the epidemic (3). Moreover, the use of chest CT scan for COVID-19 screening has been advocated in previous studies, especially when the results of RT-PCR were negative (2).
The manual evaluation of three dimensional (3D) CT volume data is a tedious and time-consuming process, which is highly dependent on the expert’s clinical experience (8). However, development of artificial intelligence (AI)-based medical image analysis methods can help overcome the abovementioned challenges. As one of the subsets of AI, deep learning algorithms have been found to be effective in different medical fields, such as radiology, dermatology, ophthalmology, and pathology (9, 10).
Since the beginning of the COVID-19 epidemic, many efforts have been made to automatically analyze chest CT images to expedite and facilitate the proper diagnosis and management of COVID-19. The majority of the proposed methods fall into 2 general categories: Classification and segmentation (11). In classification, the goal is to assign a label to examples from the input domain. For the classification of COVID-19 cases, algorithms are trained on labeled CT images of healthy individuals and COVID-19 patients (in many cases, patients with other diseases) to learn predictive modeling (5, 12-14). Previously, we investigated the performance of various backbone architectures for classifying COVID-19 cases and found that the VGG19 architecture demonstrated the best performance (15).
On the other hand, in medical image segmentation, the goal is to label each pixel to determine the region of interest (ROI). VB-Net, U-Net, and other variants of U-Net are among the most common algorithms for medical image segmentation (5, 16-18). CT-based COVID-19 classification has shown to be more accurate when trained based on segmentation masks rather than binary (0 and 1) labels representing presence or absence of the lesion. The improved accuracy can be related to the complementary knowledge provided to the model using segmentation masks. Due to this, many studies have used deep learning segmentation techniques for the automated diagnosis of COVID-19 lesions and the subsequent interpretation of CT images (19, 20). Although most of these studies achieved high accuracy in detecting and segmenting COVID-19 lesions, their robustness was not comprehensively evaluated on multi-centric datasets (from different scanner makes and models and population geographies) due to the limited availability of COVID-19 datasets at the time. Hence, with the availability of more annotated datasets, it is essential to evaluate the generalizability of deep learning-based models.
Khan et al. (21) proposed a threshold-based segmentation method to quantify COVID-19-related pulmonary abnormalities. Their approach indicated a Dice similarity coefficient (DSC) of 46.28% for a combination of 2 COVID-19 CT datasets. However, the generalizability of their method on the datasets was not separately assessed. Fan et al. (22) introduced a lung infection segmentation network (Inf-Net) to automatically identify infected areas on CT images. They used a semi-supervised approach to compensate for the lack of data. However, they only had access to one labeled dataset, making it difficult to investigate the performance of the proposed model on unseen data acquired by different CT devices.
Wang et al. (23) proposed a noise-robust learning framework based on a 2D convolutional neural network (CNN), combined with an adaptive self-ensembling framework for slice-by-slice image segmentation. Although they used a dataset, consisting of CT images acquired from 10 different hospitals, they randomly split the images into training, validation, and test sets and did not study the generalizability of their proposed method on unseen CT datasets. Shan et al. (24) developed a deep learning-based model, called VB-Net, to segment COVID-19-infected regions on CT scans. The developed VB-Net model was trained using a human-involved-model-iterations (HIMI) strategy on a dataset of 249 COVID-19 cases and validated on another dataset of 300 COVID-19 cases. The model yielded a DSC of 91.6%, and the average DSC between the two radiologists was 96.1%. The relatively similar values of DSC indicate the high accuracy of the deep learning-based model in quantifying COVID-19 lesions based on CT data. Nevertheless, the model was validated on a monocentric dataset, which might not represent the generalizability of deep learning systems on different patient populations.
Lastly, Müller et al. (1) implemented a standard 3D U-Net architecture through five-fold cross-validation on 20 CT scan volumes of COVID-19 patients. An average DSC of 76.1% was reported for this method. This study used a limited dataset of 20 cases and was tested on the same dataset it had been trained on; therefore, the generalizability of the model was not evaluated on an unseen dataset.
2. Objectives
The present study aimed to develop and evaluate a deep learning-based model for the automated segmentation of COVID-19 lung lesions using chest CT scans based on multi-centric data. Moreover, this study aimed to evaluate the effectiveness of lung segmentation as a preprocessing technique and to compare the performance of an AI-based model against radiologists. Finally, this study aimed to demonstrate the limited generalizability of deep learning-based models through domain shift and to propose a calibration strategy to improve their performance.
3. Patients and Methods
This section describes the image datasets and the proposed method used in the present study.
3.1. Image Datasets
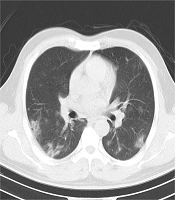
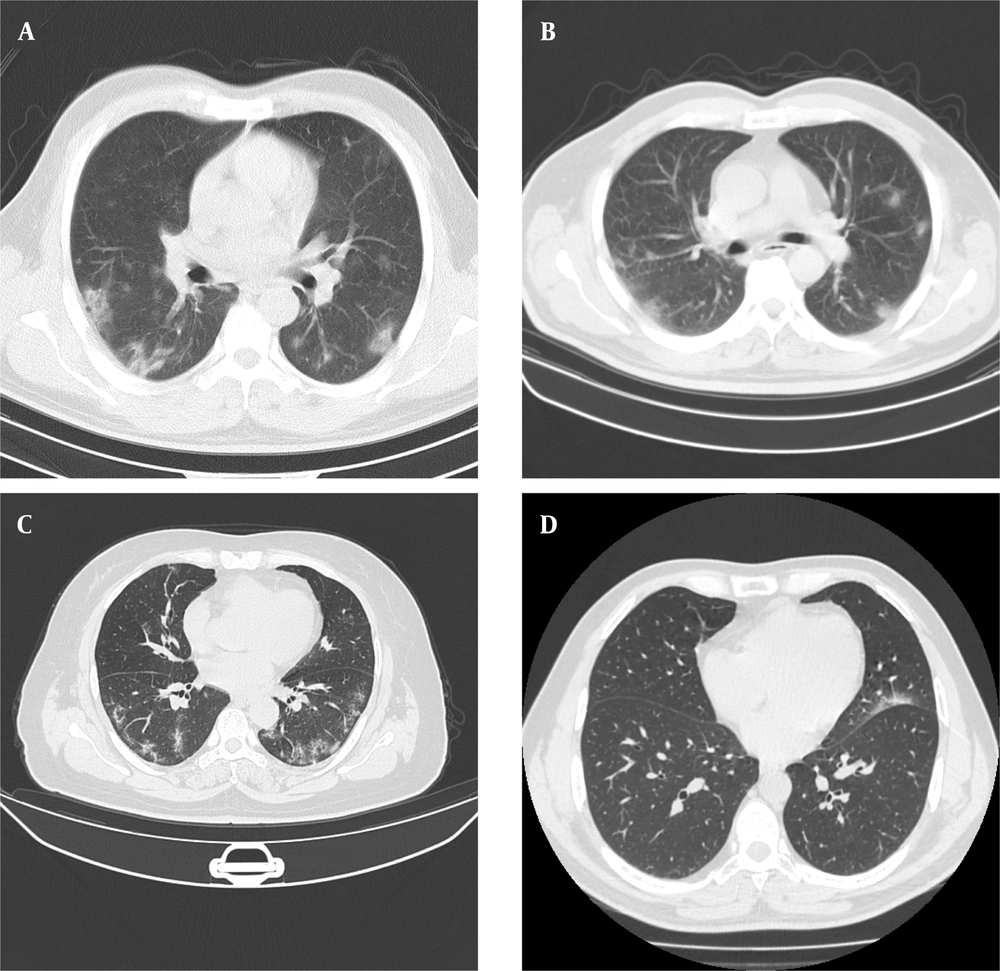
Four separate datasets (2 private and 2 public datasets) were used in this study. Appendix 1 summarizes the COVID-19 CT datasets of this study. Figure 1 presents some examples of CT images for each dataset.
The computed tomography scans (CT) of patients infected with coronavirus disease 2019 (COVID-19) for the first (A) and second (B) private datasets, COVID19-P20 dataset (8) (C), and MosMedData dataset (25) (D). The image quality differences (e.g., spatial resolution, noise, and contrast) in chest CT scans suggest the importance of evaluating the generalizability of deep learning-based models.
3.2. Proposed Method
Figure 2 demonstrates a schematic representation of the workflow. The approach used for implementing the proposed algorithm can be divided into 3 main steps: (1) preprocessing step; (2) automated segmentation of COVID-19 lesions using a VGG16-UNet architecture (26); and (3) experimental setup.
3.2.1. Preprocessing
Data preprocessing is an integral part of medical image segmentation. In this study, image intensities were normalized to a scale of 0 to 1. All images were also resized to a size of 256 × 256 to reduce the computational burden. Besides, the effect of lung segmentation as a preprocessing stage was investigated. For this purpose, each lung was segmented (27), and the model was trained on CT images of lung segments. To evaluate the effectiveness of lung segmentation as a preprocessing stage, the VGG16-UNet model was evaluated with and without initial lung segmentation.
3.2.2. Segmentation
To segment COVID-19 lung lesions on CT slices, a deep learning approach based on the U-Net framework (17) was developed. A contracting path (on the left side) and an expansive path (on the right side) comprise the VGG16-UNet, similar to the regular U-Net. The contracting path is the VGG-16 structure with 5 convolutional blocks, consisting of 2 or 3 convolutional layers with a small 3 × 3 receptive field. The resolution was down-sampled at the end of each block using max-pooling layers, and the feature map was up-sampled by a factor of 2. The convolutional layers extracted higher-level features from the input image by moving from the first block to the last one. On the other hand, the expansive path attempted to restore images to their original dimensions. The resolution was up-sampled by a factor of 2 in each block, while the number of feature maps was reduced by a factor of 2. Two convolutional layers (3 × 3) followed each block after the concatenation stage, each with a batch normalization (BN) operation and a ReLU layer. Also, concatenation was performed with the corresponding feature maps from the contracting direction via skip connections.
To indicate the superiority of our proposed framework to other similar deep learning-based architectures, a comparison was made between U-Nets (17) and Link-Nets (28) with different encoder structures (26). Appendix 2 presents a comparison of U-Net and LinkNet architectures for semantic segmentation. The number of blocks was subject to change when using different architectures (e.g., VGGNets, EfficientNets, ResNets, and DenseNets). The U-Net and Link-Net architectures were completely similar in the encoder part. Nonetheless, 2 key components differentiate them from one another. First, in the U-Net architecture, the feature maps are concatenated in the contracting and expansive paths, while they are added together in the Link-Net architecture. Second, the decoder block in the U-Nets is composed of a 2 × 2 up-sampling layer and two 3 × 3 padded convolutional layers, followed by BN and ReLU layers. On the other hand, the decoder block in the Link-Nets consists of a 1 × 1 convolutional layer (reducing the number of channels by a factor of 4), a 2 × 2 up-sampling layer, a 3 × 3 padded convolutional layer, and a 1 × 1 convolutional layer (increasing the number of channels to the feature map size of the corresponding encoder block). A BN operation and a ReLU layer follow all the convolutional layers in the decoder part.
3.2.3. Experimental Setup
For a quantitative comparison of the tested architectures, a five-fold cross-validation was conducted on the first private dataset at the patient level. For this purpose, all the patients were randomly divided into 5 folds; each time, 4 folds were used for training, and 1 fold was used for testing. Next, 25% of patients from the training dataset were selected as the validation set to help with the optimization procedure and prevent overfitting.
To minimize the expert’s segmentation error in localizing COVID-19 lesions, the CT slices of patients, in which no area was specified as the infected region, were excluded from training. The main reason for this exclusion was the possibility of missing small lesions on CT slices by the radiologist during the annotation process. The trained model was tested on the second private dataset to analyze the generalizability of AI-based models. The mean and standard deviation (SD) of the 5 scores were calculated for analyses.
To further investigate the performance of the trained model, it was trained on 2 public datasets. First, the model was trained on the COVID19-P20 dataset, consisting of 20 COVID-19 cases (10 cases from the Coronacases Initiative resource and 10 cases from the Radiopaedia resource), using two-fold cross-validation. In each fold, half of cases from each group (Coronavirus or Radiopedia) were selected as the training set, and the rest were selected as the validation and test set. Second, the model was trained on a mixture of COVID19-P20 and MosMedData datasets, consisting of 70 cases, using five-fold cross-validation.
3.2.4. Statistical Analysis
In this study, the DSC, receiver operating characteristic (ROC) curve, area under the curve (AUC), and Pearson’s correlation coefficient (PCC) were measured to compare and investigate the results. Generally, the DSC computes the region-based similarity of the model segmentation result with the ground truth and is calculated as follows:
where P is the model output, and G is the mask of infectious areas specified by the expert. The DSC ranges from 0 to 1, with 1 representing the greatest similarity between the model output and the ground truth.
Additionally, for a binary classification problem, the ROC curve indicates the true positive rate (TPR) versus the false positive rate (FPR), which are measured as follows:
where TP, FP, TN, and FN denote the number of true positive, false positive, true negative, and false negative predictions, respectively. The optimal model would have a TPR of 1 and an FPR of 0. The model with the highest AUC was considered as the optimal model, as it would have a higher TPR for the same FPR.
Moreover, PCC was calculated to examine the performance of the AI model in quantifying chest CT severity scores. The PCC between the AI model and the radiologist was calculated in this study:
where cov (Z, Z´) is the covariance between the AI model and the radiologist, σZ is the SD of the AI model, and σZ’ is the SD of the radiologist. Statistical analysis was performed using Python (the np.corrcoef function for PCC calculation).
4. Results
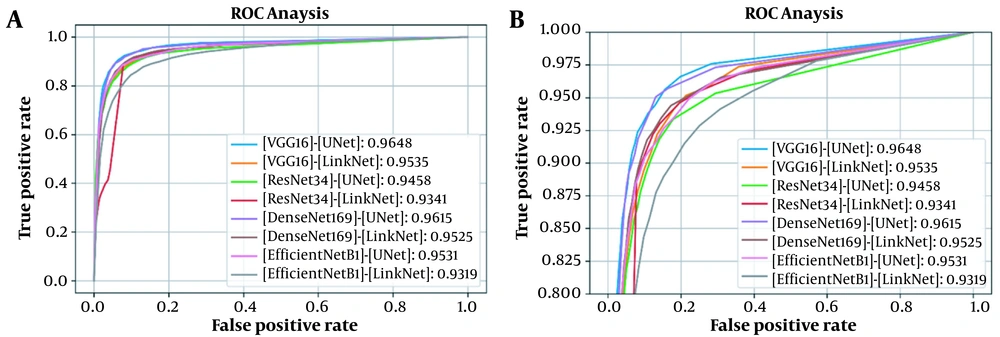
This section presents the results of analyses conducted in this study. First, the proposed architecture was compared against several similar models in terms of DSC, ROC curve, and AUC. Table 1 demonstrates the DSCs and AUCs, and Figure 3 represents the corresponding ROC curves for testing the model on the first dataset. The results showed the superior performance of the VGG16-UNet architecture to other architectures, with a DSC of 84.23% ± 1.73% and an AUC of 0.9648. The results were validated via five-fold cross-validation on the first dataset, including 297 cases (150 COVID-19 cases and 147 healthy individuals). Each of the five models, which was trained using five-fold cross-validation, was also tested on the second private dataset, containing 82 COVID-19 cases.
| Model and encoder | Number of parameters (million) | Training time (s) | DSC (%) | AUC |
|---|---|---|---|---|
| U-Net | ||||
| VGG16 | 23.75 | 62.10 | 84.23 ± 1.73 | 0.9648 |
| ResNet34 | 24.45 | 42.10 | 82.68 ± 1.55 | 0.9458 |
| DenseNet169 | 19.51 | 82.13 | 83.76 ± 0.85 | 0.9615 |
| EfficientNetB1 | 12.64 | 81.44 | 82.58 ± 1.25 | 0.9531 |
| Link-Net | ||||
| VGG16 | 20.32 | 60.10 | 82.17 ± 1.91 | 0.9535 |
| ResNet34 | 21.63 | 36.60 | 82.13 ± 0.95 | 0.9341 |
| DenseNet169 | 15.61 | 77.13 | 77.32 ± 5.02 | 0.9525 |
| EfficientNetB1 | 8.55 | 76.25 | 80.41 ± 1.07 | 0.9319 |
Model Specifications, Dice Similarity Coefficients in Percentage (mean ± standard deviation), and Area Under the Receiver Operating Characteristic Curve for Private Dataset #1
According to Table 2, the VGG16-UNet model demonstrated superior performance to other architectures, with a DSC of 56.61% ± 1.48% on the (unseen) second private dataset. The proposed model was also trained and tested on 2 public datasets, that is, the COVID19-P20 dataset, followed by a combination of COVID19-P20 and MosMedData datasets. As shown in Table 2, the scores were consistent with previous results, making VGG16-UNet the optimal model in terms of performance, with DSCs of 60.10% ± 2.34% on the COVID-19-P20 dataset and 66.28% ± 2.80% on the combination of COVID-19-P20 and MosMedData datasets.
| Model and encoder | Private dataset #2 | COVID19-P20 | COVID19-P20+MosMedData |
|---|---|---|---|
| U-Net | |||
| VGG16 | 56.61 ± 1.48 | 60.10 ± 2.34 | 66.28 ± 2.80 |
| ResNet34 | 55.72 ± 1.42 | 53.35 ± 2.25 | 57.32 ± 7.56 |
| DenseNet169 | 55.09 ± 1.60 | 57.38 ± 3.04 | 65.08 ± 3.95 |
| EfficientNetB1 | 55.83 ± 1.30 | 53.16 ± 2.04 | 62.43 ± 2.47 |
| Link-Net | |||
| VGG16 | 54.46 ± 3.08 | 54.14 ± 4.05 | 61.82 ± 1.69 |
| ResNet34 | 54.25 ± 1.14 | 54.54 ± 0.90 | 57.48 ± 3.03 |
| DenseNet169 | 47.60 ± 10.56 | 54.22 ± 1.04 | 64.10 ± 5.08 |
| EfficientNetB1 | 53.24 ± 1.68 | 38.94 ± 6.14 | 49.29 ± 5.05 |
The Dice Similarity Coefficients in Percentage (mean ± standard deviation) for the Model Applications on the Private Dataset #2 and 2 Public Datasets (COVID19-P20 and COVID19-P20+MosMedData)
Second, the effect of lung segmentation as a preprocessing stage in training the COVID-19 lung lesion segmentation network was evaluated. The results suggested that having a lung segmentation preprocessing stage did not lead to improved performance on both datasets. For this purpose, a model proposed in a previous study (27), which was also trained on COVID-19 images, was employed. Table 3 presents the results of the VGG16-UNet model in terms of DSC. Considering the inferior performance of the model when using a preprocessing stage, no lung segmentation was performed for the following assessments.
| VGG16-UNet model | Dice similarity coefficient (%) | |
|---|---|---|
| Dataset #1 | Dataset #2 | |
| Without segmentation | 84.23 ± 1.73 | 56.61 ± 1.48 |
| With segmentation | 83.10 ± 1.39 | 54.90 ± 1.12 |
Evaluation of the Effect of Lung Segmentation on the Dice Similarity Coefficients in Percentage (mean ± standard deviation) for Private Datasets #1 and #2
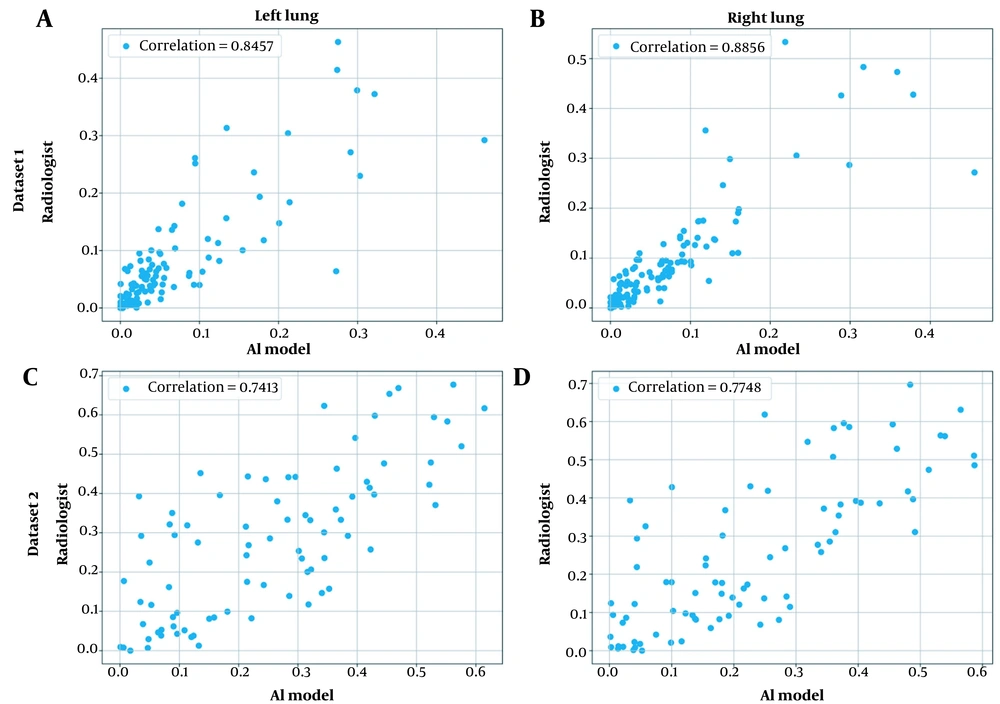
Third, chest CT severity scores were calculated by dividing the infected region by the overall region per lung segmented using the model developed in the literature (27). Appendix 3 presents the general procedure for calculating the chest CT severity score for a single CT scan. Figure 4 shows the scatter plot for the severity prediction of the AI model against a radiologist for both lungs and datasets.
Scatter plots for the correlation of chest computed tomography (CT) severity score predictions by the artificial intelligence (AI) model and the radiologist for both private datasets and lungs: A, The first dataset, the left lung; B, The first dataset, the right lung; C, The second dataset, the left lung; and D, The second dataset, the right lung.
Fourth, to reach a reasonable understanding of the practicality of deep learning-based models in real-world settings, the performance of the VGG16-UNet model was compared against the inter-rater agreement between the two radiologists on the (unseen) second private dataset. Table 4 compares the performance of the AI model against the average DSC between the two radiologists for 67 COVID-19 cases.
| Model | Dice similarity coefficient (%) |
|---|---|
| VGG16-UNet (threshold: 0.5) | 58.27% |
| VGG16-UNet (threshold: 0.15) | 63.48% |
| Radiologist | 64.01% |
Performance Comparison of the Models and the Radiologists on the Private Dataset #2 Based on the Dice Similarity Coefficients
Finally, the generalizability of the trained model was investigated by testing it on an unseen dataset. Next, a simple calibration strategy was proposed by changing the threshold value of the segmentation map for every new dataset, using a very limited number of cases. Table 5 presents the performance of the model, with and without calibration for the testing set of the second private dataset.
| Percentage of training data used for calibration (threshold with the highest Dice similarity coefficient) | 0% (0.5) | 10% (0.10) | 20% (0.05) | 30% (0.05) | 40% (0.10) | 50% (0.05) |
|---|---|---|---|---|---|---|
| Dice similarity coefficient (%) for the test set | 61.16% | 66.29% | 64.88% | 64.88% | 66.29% | 64.88% |
The Dice Similarity Coefficients in Percentage for 50% (testing set) of the Private Dataset #2
5. Discussion
5.1. Performance of Encoders
According to Table 1 and Table 2, in both U-Net and Link-Net models, the application of VGG16 encoder yielded the best performance. It can be concluded that the VGG16 encoder led to the extraction of richer features from the input image and consequently, a higher DSC. Comparison of the U-Net and Link-Net structures revealed the superior performance of the U-Net model using all 4 encoders, which could be explained by the greater number of parameters in the decoder part of the U-Net model. The higher number of parameters could improve the recovery of the lost spatial information in the encoder part.
5.2. Lung Segmentation as a Preprocessing Step
The impact of lung segmentation on the overall performance of the model was analyzed in this study. As shown in Table 3, the application of lung segmentation as a preprocessing step decreased the performance of the VGG16-UNet model. This could be attributed to the inaccurate segmentation of the lungs, especially in the presence of a diseased lung. Appendix 4 shows accurate and inaccurate lung segmentations for a COVID-19 patient. The superior performance of discarding lung segmentation also indicates that the model could simultaneously learn the location of the lungs and COVID-19 lesion patterns.
5.3. Comparison of the Model with the Radiologist’s Performance
Validation of AI-based models in real-world settings via comparison against clinical experts’ performance is of monumental importance. In this study, a correlation analysis was conducted between the proposed model and a radiologist in terms of chest CT severity scores. The results showed a high correlation between the model and the radiologist’s opinion for quantifying the chest CT severity scores (0.89 and 0.85 on the first private dataset and 0.77 and 0.74 on the second private dataset for the right and left lungs, respectively).
Additionally, the performance of the model was compared against the average DSC between the two radiologists (i.e., inter-rater agreement), using 67 COVID-19 cases from the second private dataset. Based on the comparison of values presented in Table 4, the performance of the AI-based model was close to the inter-rater agreement, which demonstrated the accuracy of deep learning models in quantifying COVID-19 lung lesions on CT scans, even when they were not solely trained on that specific dataset.
Comparison of the performance of the proposed architecture against the radiologists’ performance indicated the advantage of time efficiency in AI-based models. While a manual assessment of CT scan volume by radiologists can take up to 15 minutes, our proposed model segmented each slice in 60 ms, resulting in COVID-19 segmentation in less than a minute. Therefore, an AI-based image analysis has the required speed to meet the high demand for image assessment during the COVID-19 pandemic, while facilitating accurate diagnoses (29).
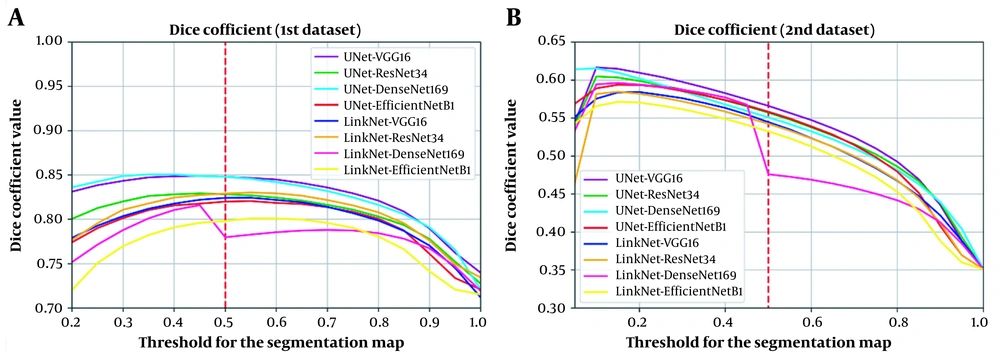
It is worth mentioning that in the present study, a threshold of 0.5 was used to classify each pixel on the output segmentation map into either a COVID-19 lesion (labeled as 1) or normal (labeled as 0); this threshold (0.5) was selected, because it is the best threshold for the validation part of the first dataset (Figure 5A). However, it is not certain if this finding applies to the unseen dataset (Figure 5B). The next section describes a simple calibration approach to improve performance on unseen datasets simply by changing the threshold of the segmentation map, using a very limited number of cases.
Analysis of the effect of threshold change in the segmentation map on the model performance suggests this effect on A, The validation set of the first private dataset; and B, The second private dataset (unseen dataset). The plot illustrates that variability in scanners and imaging protocols can negatively affect the accuracy of deep learning-based models. Therefore, use of a calibration procedure to determine the best threshold can improve performance.
5.4. Generalization Analysis and Calibration
The CT scans can vary depending on differences in CT scanners and imaging parameters. This variability manifests as different spatial resolutions, image contrasts, and noise levels, which can negatively impact the accuracy and consistency of deep learning-based models. However, there are several explanations for this performance decline. First, the performance decline from 84.23% ± 1.73% on the first dataset to 56.61% ± 1.48% on the second dataset could be partly explained by the presence of healthy subjects in the first (mixed) private dataset. The elimination of healthy individuals from the testing set resulted in a DSC of 74.35% ± 1.57% for COVID-19 cases. Second, further qualitative analysis of the second private dataset demonstrated its complexity, especially due to the presence of mild COVID-19 cases, leading to a significant decline in DSC when the model missed small lesions. The low inter-rater agreement between the two radiologists (DSC of 64.01% for 67 COVID-19 cases from the second dataset) provided further evidence of its complexity. Finally, domain shift is another key factor in performance decline due to differences in scanners and imaging protocols across different centers.
To minimize the performance decline, the effect of change in the threshold of the output segmentation map on both datasets was investigated. According to Figure 5, use of the same threshold for the two datasets would not necessarily lead to the best performance. Therefore, a calibration procedure can be employed to determine the best threshold for the unseen dataset. To demonstrate this possibility, we trained the best-performing model using the first private dataset and calibrated the segmentation map (output) of the model using data from the second private dataset.
For calibration, 0% (no calibration) to 50% of the cases from the second private dataset were used in 10% increments to study whether better calibration can be done with a greater number of cases. The other 50% of the data was fixed and set as the test set. For each experiment, output segmentation maps (of the model) were generated and thresholded in the range of 0 and 1, in 0.05 increments. Next, DSC was calculated and averaged across all training cases, and the best threshold was selected to be used on the test dataset. The DSCs reported in Table 5 are the average scores on the testing set. The results demonstrated that a very limited number of cases in a dataset (10%) could be used to calibrate the model when using an unseen dataset.
Other more advanced techniques that can mitigate variations in CT vendors and acquisition protocols include CNN-based image normalization and generative adversarial networks. One idea is to develop an adversarial neural network similar to a model proposed in the literature (30) as a harmonization block for transforming the CT data acquired from different scanners with various imaging protocols into a reference standard. Accordingly, the main network could conduct a more effective and uniform assessment of COVID-19 lesion segmentation. This suggestion could improve the performance of the main network for the unseen dataset, regardless of the scanner specifications and without applying a calibration procedure.
In conclusion, this study presented a deep learning-based approach to automatically detect and segment COVID-19 lung lesions using chest CT scans. It was found that a VGG16-UNet model performed better than other architectures and achieved a DSC of 84.23% ± 1.73% for a mixed dataset of healthy and COVID-19 subjects and a DSC of 56.61% ± 1.48% for the unseen dataset of COVID-19 patients. The performance gap was attributed to (1) the presence of healthy subjects in the first dataset, increasing the DSC from 74.35% ± 1.57% to 84.23% ± 1.73%; (2) complexity of the second private dataset due to the presence of mild COVID-19 cases; and (3) limited generalizability of the model due to domain shift because of variations in CT scanners and imaging protocols. The model was further assessed on 2 public datasets and achieved DSCs of 60.10% ± 2.34% and 66.28% ± 2.80% for the COVID19-P20 dataset and the combination of COVID19-P20 and MosMedData datasets. Moreover, the experiments suggested that lung segmentation was an ineffective preprocessing strategy for infection segmentation of COVID-19 cases. Also, the deep learning-based model showed good agreement with the radiologist’s performance. Finally, the generalizability of the model was evaluated in this study, and a simple calibration strategy was proposed, improving its performance by more than 5% based on DSC. Future research can focus on image harmonization techniques that can help mitigate unwanted variations in CT scans.
.jpg)
.jpg)
![A schematic representation of the workflow. It presents the pipeline for statistical analysis (Dice similarity coefficient [DSC] comparison and correlation coefficient calculation) and model comparison on both datasets. A schematic representation of the workflow. It presents the pipeline for statistical analysis (Dice similarity coefficient [DSC] comparison and correlation coefficient calculation) and model comparison on both datasets.](https://services.brieflands.com/cdn/serve/3170b/bba0cdcab2f09dc3f85a57839f82ba3f7640e185/iranjradiol-117992-i001-F2-preview.webp)