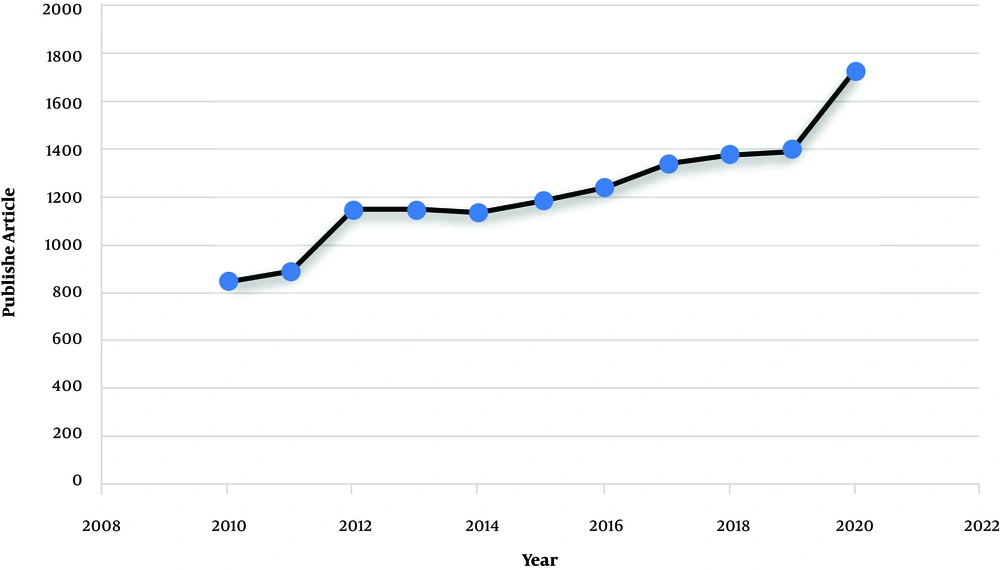
Non-obstructive azoospermia (NOA) (MedGen UID: 866757) is the most severe form of male infertility affecting around 1% of the male population and 10% of infertile men (1). The disease phenotype is included with the absence of any measurable level of sperm in the semen without any reproductive tract obstruction (2). Around 25% of non-obstructive azoospermia cases are due to the Y chromosome microdeletions (YCMs), Karyotype abnormalities, and missense mutations in genes involved in reproductive function. The remaining cases are categorized as idiopathic and show multifactorial inheritance (3). The illness also presents as consequences of endocrine and chronic diseases that interrupt the hypothalamic-pituitary axis, like diabetes, hypogonadism, and low levels of sex steroid hormone (4). Pathological conditions that affect the sperm transport through the vas deferens, such as varicocele, cancer, maldescended testes, and urological infections, may also be associated with NOA (5). In recent years, the application of advanced genetic analysis technologies, like deep sequencing, has considerably increased our knowledge about NOA (6). A quick search on datasets, like PubMed and Google Scholar, shows the increased interest in studying the genetic cause of male infertility. As indicated in Figure 1, searching using the keyword, such as “Non-obstructive azoospermia” indicates that there has been a big tendency to use this subject from 2010 to 2020. Accordingly, in this commentary, we provide an insight into an important development in NOA and discuss the fundamental concepts of NOA regarding its etiology, diagnosis, and management. In fact, we hope to draw a perspective on NOA for those who are interested in NOA. We first discuss all the documented genetic causes of the disease and then, we focus on the diagnostic and therapeutic methods and shed light on the importance of genetic testing in NOA.
Genetic Causes of Non-obstructive Male Infertility: Around 40% of patients diagnosed with NOA are currently classified as idiopathic and may be due to unknown genetic aberrations. The YYCMs, X-linked and autosomal monogenic factors, and cytogenetic defects, such as chromosomal translocations, are the well-described factors to cause or increase the risk of NOA in men (3). In the next part of this commentary, we explain these factors in detail.
Cytogenetic Defects: In comparison with obstructive form, the prevalence of aneuploidy in the karyotype of NOA patients is ten times greater and contains the X chromosome frequently. As estimated, NOA was diagnosed in around 95% of Klinefelter cases (47, XXY). Of note, focal spermatogenesis has been reported for mosaicism of this syndrome (47, XXY/46, XY) (7). Therefore, there is a possibility of fertility in mosaic patients using assisted reproductive technology (ART), such as testicular sperm extraction (TESE) techniques. Translocation between X and Y or between sex chromosomes and autosomes has also been contributed to the pathogenesis of NOA (8).
Autosomal Monogenic Factors: The contribution of several autosomal genes has been documented to germline development and spermatogenesis. Thus, it is not surprising if these genes show a positive correlation with male infertility. These genes include BAX, BCL16, c-kit, ATM, HSP70.2, RAD6B, MDHC7, CREM, DNA11/12, CFTR, SPATA16, AURKC, CATSPER1, GNRHR, MTHFR, SYCP3, SOX9, WT1, and NR5A1 genes (9). Around 60 - 90% of congenital bilateral absence of the vas deferens (CBAVD) patients harbor the mutations in the CFTR gene, particularly F508del mutation in 70% of CBAVD patients. This mutation is a 3-bp deletion, resulting in the loss of phenylalanine at amino acid residue 508 of the CFTR polypeptide (10). The mutations of the CFTR gene have been frequently reported in NOA, showing the probable role of this protein in spermatogenesis (11).
X-chromosome Monogenic Factors: Mutations in two X-linked genes, USP26 (Ubiquitin Specific Peptidase 26) and SOX3 (SRY-Box Transcription Factor 3) have been reported for some types of NOA. Several mutations have been reported for these two genes, which result in the severe impairment of spermatogenesis and hypogonadism (3). The USP26 gene encodes a deubiquitinating enzyme, which is actively involved in the regulation of the cell cycle, maintenance of the chromosomal structure, and gene silencing (12). The SOX3 gene is specifically expressed in developing testicular and neural tissues and encodes the SOX3 transcriptional activator, playing an important role in the regulation of embryonic development and in the determination of cell fate (13). An X-linked gene androgen receptor (AR) is presently considered for genetic analysis and counseling in the diagnosis of NOA. The gene encodes a transcription factor of the steroid hormone regulating the expression of androgen-responsive genes upon binding to the hormone ligand. Androgens are steroid hormones to mediate male sex development and spermatogenesis through binding to AR (1). AR is a highly polymorphic gene due to the presence of two variable number tandem repeats (VNTRs), (CAG)n (encoding for glycine), and (CGN) n (encoding for glutamine) in exon 1 of the gene. The length of repeats is associated with impaired sperm production and male infertility (1, 14).
Y Chromosome Microdeletions: Around 5 - 10% of NOA cases are positive for YCMs. Regarding the repetitive structure of the Y chromosome, susceptibility of this chromosome increases for non-allelic homologous recombination (NAHR) between two allele regions of DNA with high sequence similarity, the process, which can eventually be ended with YCM (15). Microdeletions of six key regulatory genes of spermatogenesis, including USP9Y, DDX3Y, EIF1AY, PRS4Y2, KDM5D, and DAZ have been observed in NOA patients. All genes belonged to the azoospermia factor (AZF) proximal (a), middle (b), or distal (c) Yq11 subregions and regulate cell cycle and germ cell survival. To our knowledge, microdeletions of the AZFc region is the most prevalent microdeletion of Yq, which have been detected in more than 80% of NOA patients (16).
The gene ubiquitin-specific peptidase 9, Y-linked (USP9Y) on the AZFa region, is one of the critical spermatogenesis modulators removing the ubiquitin residue from ubiquitin-linked precursors. The gene contains 47 exons and spanning 170 kb of DNA on the Y chromosome. It has been documented that microdeletions in this gene have been associated with phenotypes from oligozoospermia to azoospermia. However, routine screening for AZF gene-specific deletions is not recommended since AZF gene-specific deletions are extremely rare, and USPY9 deletions also have heterogeneous phenotypes (17). In a recently published article by Kalhor et al., the five polymorphisms of this gene have been analyzed in 100 men suffering from NOA. They found no significant association between these nucleotide changes and increased risk of NOA. This may arise from the limited sample size in this study, and a larger sample size may rescue this result (18).
Diagnosis of NOA: The diagnosis of male infertility usually starts with taking a complete medical history, as well as a clinical examination. Semen analysis, hormonal assays, radiological checkups, and genetic studies, particularly in cases of severe defects, are the main tests (2, 19). The patient’s medications, such as taking anticancer chemotherapy or special drugs, should be considered. Some drugs, like steroids and 5a-reductase inhibitors, can damage spermatogenesis. Additionally, undescended testis or the presence of the secondary sexual characteristics should be evaluated according to the Tanner stages (20). Genitalia or poor pubic hair is probably linked to hypogonadism. Notably, the size of the testes reflects the level of spermatogenesis. The testicular measurement volume is important during NOA judgment (19, 20). The volumes less than 15 cc with a flat epididymis are probably attributed to NOA. Ultrasonography is the gold standard in this regard (20).
Available Genetic Testing for Investigation of NOA: Three types of commonly used genetic tests are 1) cytogenetic analysis, like karyotyping to evaluate the presence of chromosomal aneuploidy and structural aberration, 2) blood-based multiplex polymerase chain reaction (PCR) to detect YCMs, and 3) mutational analysis of a specific gene through DNA sequencing (21). In recent years, the application of next-generation sequencing (NGS) technology, such as whole-exome-sequencing (WES) has facilitated the discovery of causative genes in infertile men. In spite of previously used mono-targeted tests in medical genetic laboratories, which were done based on the clinical presentation of patients on one gene or the small number of genes, during the WES, all of the exonic regions in the genome are sequenced and screened for the presence of probable mutations. The technique is cost- and time-effective to study the rare variant mapping in complex disorders, discovery of causative genes in Mendelian disorders, and also case studies (22, 23). Using RNA sequencing, around 430 genes have been linked to spermatogenesis and NOA (24). There has been limited prospective, point-of-care, next-generation sequencing assessment of males with idiopathic NOA to determine the fraction with genetic causes (13). The challenge in identifying genetic variants in idiopathic males is that the high prevalence of NOA, despite the detrimental effect on reproductive fitness, suggests that high genetic heterogeneity underlies this condition.
Management of NOA: Although there is no treatment to reestablish spermatogenesis in NOA, these patients can have their own children by means of in vitro fertilization (IVF) and the intracytoplasmic sperm injection (ICSI) in patients with a low count of sperm (25). However, genetic diagnosis is necessary to avoid vertical transmission of genetic anomalies in the newborn (23, 25).
Discussion and Conclusions: NOA is the most severe form of male infertility, which is differentiated from obstructive form based on semen analysis as the cornerstone of azoospermia diagnosis, hormonal assays, radiological examination, and genetic studies, like karyotyping to check up the chromosome aberration or PCR to screen the known mutations in male infertility, like YCMs or CFTR gene defect (23, 26). During the last decades, using the advanced approaches of genomics and high throughput sequencing, a number of genes with direct effects on the regulation of spermatogenesis have been discovered. These findings on male fertility-related genes make the causative molecular mechanisms of infertility more clear (23). Previously published studies have focused on the family-based approach to discover candidate genes with high potency in male infertility (27). However, more experimental jobs and deeper knowledge are needed before recommending these genes for screening male infertility in clinics. If this goal is achieved, prognosis, diagnosis, and even treatment of infertility will be improved, especially for idiopathic patients to help these patients to produce healthy and fertile offspring.