1. Background
Helicobacter pylori is one of the most prevalent human pathogens, contributing to some gastrointestinal related diseases such as gastritis, peptic ulcer, and gastric cancer (1). Over 50% of people are silent carriers of H. pylori, hence it has been defined as type I carcinogen in terms of the WHO classification (2). Gastric cancer is ranked as the fourth common cancer, and is also the second mortality cause worldwide (3). The bacteria secrete urease to neutralize the acidic condition of gastric mucosa, which then adheres to the cellular surface via blood-antigen binding protein A (BabA) and sialic acid-binding adhesion A (SabA) contributing to tissue damage by releasing several toxins such as cytotoxin-associated gene A (CagA) and vacuolating cytotoxin A (VacA) (4). Accordingly, VacA is presented in all strains contributing to the apoptosis and suppression of the T and B-lymphocytes proliferation, which finally result in chronic infection. Also, it induces the production of radical oxygen species (ROS) leading to mitochondrial DNA mutation in gastric mucosa (5).
Notably, CagA is presented in approximately 60% of H. pylori strains, which promotes severe gastritis, ulcer, and gastric cancer. Moreover, it disturbs the apical-junctional function through the interaction with tight-junction scaffolding protein ZO-1 (6). In addition, it enhances NF-κB pathways by interacting with TNF-receptor associated factor 6 (TRAF6) and TFG-B activating kinase, which consequently result in a higher level of interleukin-8 (IL-8) production. CagA also has some properties such as methylation of MGMT DNA repair gene as the epi-genic regulator, interfering with tumor suppressor RUNx3, and TP53 degradation with proteasome (7, 8). However, only some colonized bacteria may ever develop neoplasia and cancer, suggesting that the progression to cancerous type depends on several factors such as strain-specific bacteria, virulence factors, host response, and environmental factors (7).
MicroRNAs (miRNAs) are 18 - 28 base pair noncoding RNAs that regulate target gene expression through degradation or translation inhibition of mRNA (9). In bioinformatics studies, several miRNAs have been identified in the human genome. Correspondingly, it is estimated that about 30 - 92% of human genes are regulated by miRNAs (10, 11). They play important roles in apoptosis, cell proliferation, and differentiation. Since miRNAs play a major role in the expression of tumor suppressor genes and oncogenes involving in the proliferation and apoptosis, it also contributes to the initiation and progression of malignancy (12). In the case of any gastric cancer, some miRNAs are down-regulated, which results in tumor suppression while others like mir-21 are overexpressed in gastric mucosa indicating to oncogenes (13). Although some studies demonstrated the relationship between miRNAs and gastric cancer, these data could not be generalized for all the regions of the world, as the pattern of disease differs according to ethnic and spatial localization of people worldwide. Moreover, the effect of mirNAs on the progression of gastric cancer has been evaluated in Iran, but there is no data indicating that these markers are presented since the beginning of H. pylori infection. Accordingly, this later contributes to neoplasia or some other factors may affect the expression of miRNA at neoplasia of gastric related to H. pylori.
2. Objectives
In the current study, the expression of mir-21 in biopsy samples of gastritis has been evaluated compared to healthy adjacent part of endoscopic candidates referring to the Taleghani Hospital.
3. Methods
3.1. Patients
Seventy patients who were referred to the Taleghani Hospital diagnosed with dyspeptic symptoms by a specialist between February and December 2017, were enrolled in this study. Demographic characteristics of these patients and their lifestyles such as smoking, diets, and history of disease were separately recorded (Table 1).
| Characteristic | Values |
|---|---|
| Mean age | 48.50% (18 - 83) |
| Sex | |
| Male | 32 (45.7) |
| Female | 38 (54. 3) |
| Smokers | 21 (30) |
| Canned food consumption | |
| non-consumer | 19 (27.1) |
| Once in a month | 18 (27.7) |
| Twice in a month | 9 (12.9) |
| Once in a week | 11 (15.7) |
| Twice in a week | 13 (18.6) |
| Underlying disease | |
| Heart diseases | 8 (11.4) |
| Gastric diseases | 33 (47.1) |
| Diabetes | 5 (7.1) |
| Respiratory diseases | 7 (10) |
Demographic and Clinical Characteristics of Patients with Pyelonephritisa
3.2. Endoscopy and Biopsy Collection
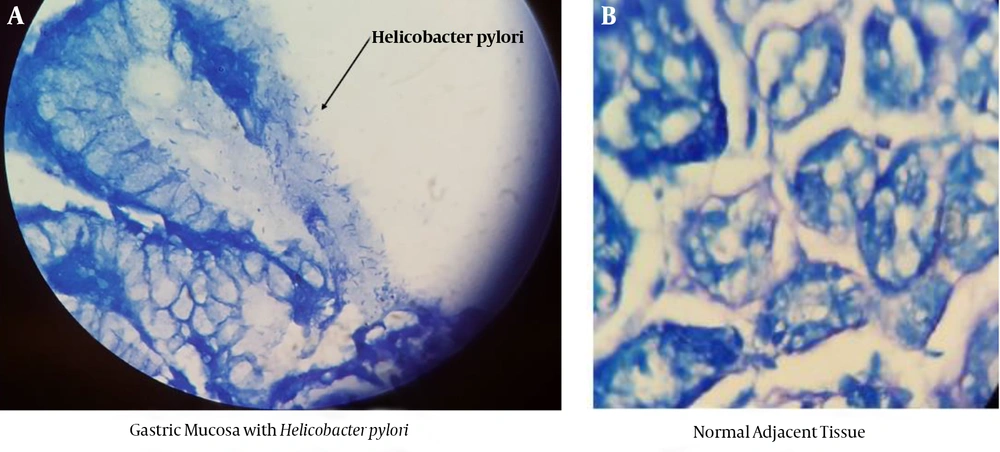
Endoscopy was performed by gastroenterologists and five biopsy samples (4 from gastritis parts of antrum and 1 from adjacent healthy part) were subsequently taken. One tissue sample was embedded in paraffin for performing pathological evaluation and the presence of H. pylori was confirmed by Giemsa staining of the paraffin-embedded blocks. The endoscopic findings were described as normal, gastritis, and gastric ulcer. Three other biopsy samples were immediately transferred to the laboratory in a transfer media, to culture and analyze the molecular characteristic. The fifth biopsy, as the healthy adjacent part, was administered as a control sample. Simultaneously, 5 mL of each blood sample from the participants was collected on EDTA.
3.3. Helicobacter pylori Culture and Identification
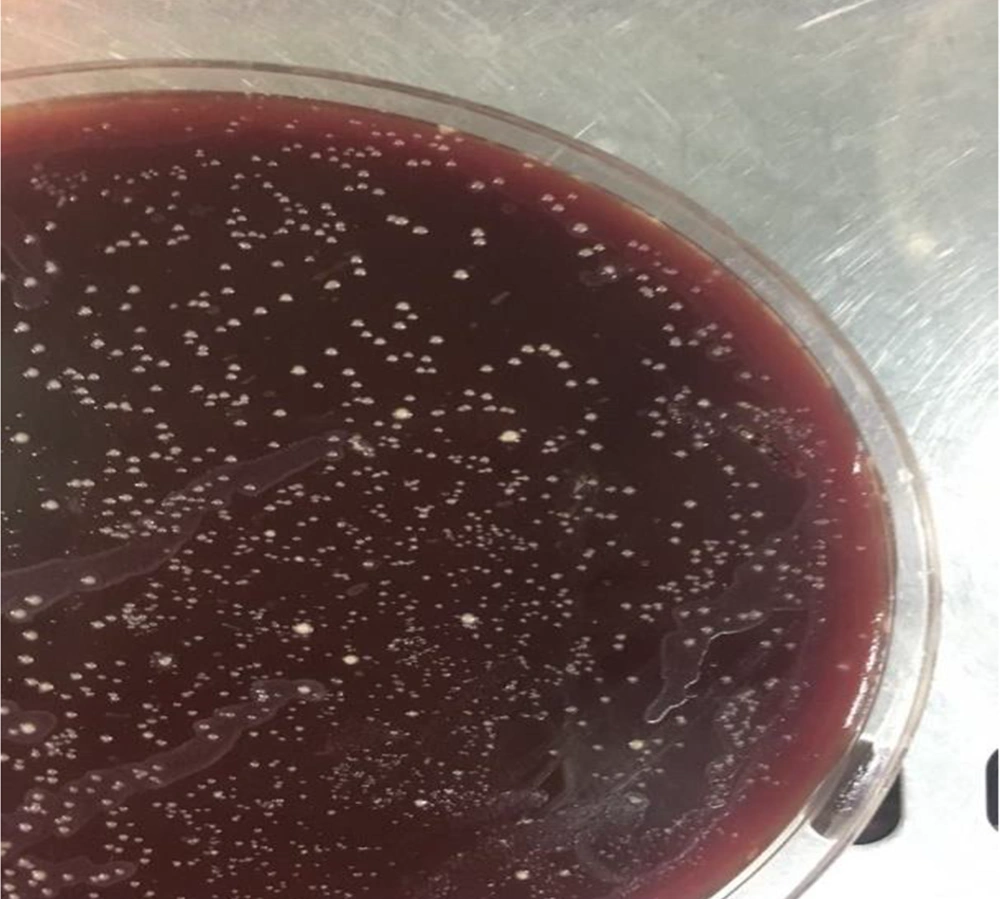
One of the fresh gastric biopsy sample was homogenized and then cultured on Brucella agar (Merck, Germany) media containing vancomycin (6 mg/L), TMP (5 mg/L), and fungizone (2 mg/L) (all antibiotics purchased from Sigma Chemical Co., St.Louis, Mo.). Afterward, the inoculated plates were incubated for 3 - 5 days at 37°C under a microaerophilic condition using CO2 gas pack type C (Anaerocult C, Merck, Germany). Subsequently, colonies were transferred to Brucella blood agar with no antibiotics after the first passage and incubated for another 48 - 72 hours at 37°C under a microaerophilic condition (Anerobic jar and gas pack type C). Based on the reference protocol, to confirm the H. pylori identification, the oxidase, catalase tests, and Gram staining were performed (14).
3.4. DNA Extraction
DNA extraction was performed using the boiling method on the confirmed bacteria colonies (15). One of the remained gastric biopsy sample was homogenized for DNA extraction of bacteria using the Stratec molecular kit (Berlin, Germany) in terms of the manufacture’s structure. Also, DNA extraction was done on the blood samples using a similar method. The OD of the extracted DNA samples was detected by a Nanodrop instrument (DeNovix, USA) at 260 nm and then stored at -20°C until analysis.
3.5. Helicobacter pylori Identification by PCR
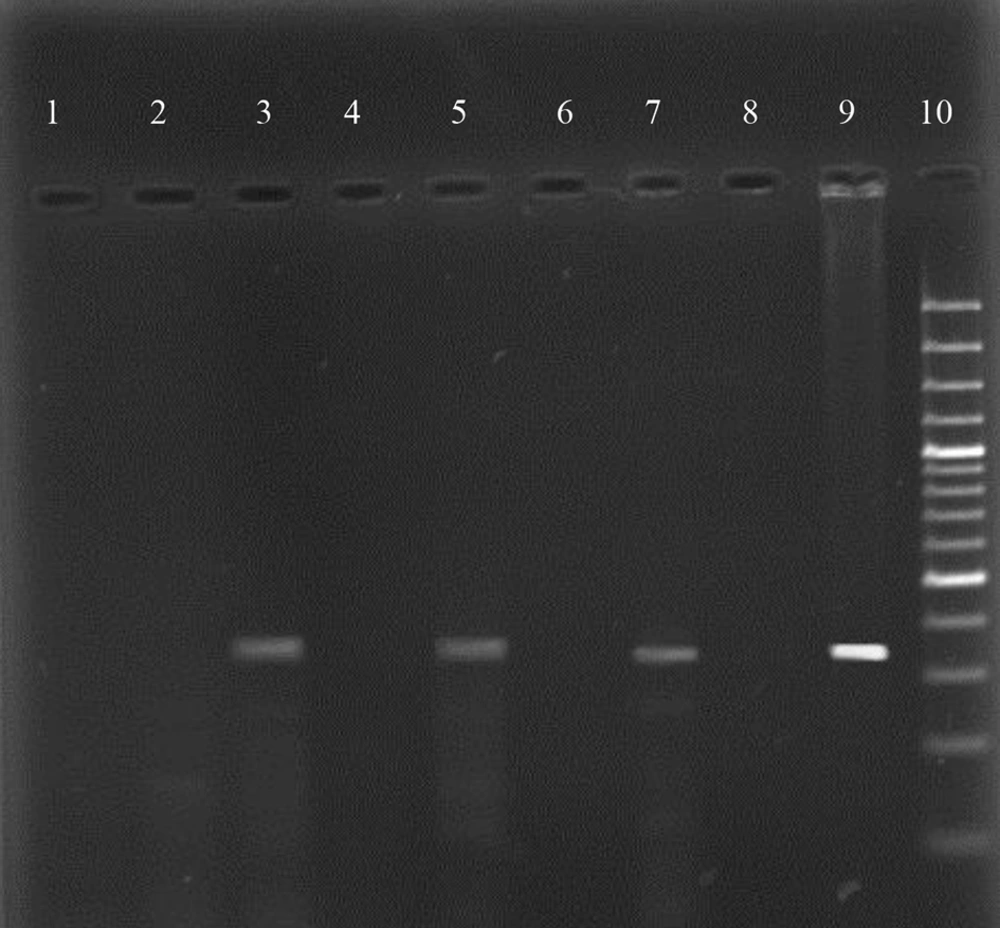
The presence of H. pylori according to molecular analysis was determined using the H. pylori PCR detection kit (Cinnagen co, Tehran. Iran) in terms of the following procedure; 1× PCR mix, Tag DNA polymerase, and DNase free deionized sterile water. Also, DNA samples were then amplified using the following program: Initial denaturation for 45 sec at 94°C, for 20 sec at 50°C, and for 20 sec at 72°C with 5 cycle repeat following 30 cycle for 20 sec at 94°C, annealing for 20 sec at 50°C, and the final extension for 30 sec at 72°C. The presence of 492 bp in 1% agarose gel indicated the positive samples for H. pylori.
Helicobacter pylori-positive samples were amplified with specific primers of the CagA gene (Metabion, Germany) to determine the invasive form of H. pylori (Table 2). In addition, PCR was carried out in a final volume of 25 µL containing 12.5 µL of Taq PCR Master Mix (Qiagen, Hilden, Germany), 2 µL (0.5 µg/µL) of template DNA, 1.0 µL (1 µM) of each primer, and 8.5 µL of PCR-grade distilled water (provided with the Taq PCR Master Mix). Amplification was performed as follows: an initial denaturation step for 4 min at 94°C, followed by 35 cycles, which included denaturation for 1 min at 94°C, primer annealing for 1 min at 59°C, and extension for 1 min at 72°C. Finally, an extension step was performed for 10 min at 72°C. A negative control (without template DNA) was included in each run. PCR products (397 bp) were separated by electrophoresis using 2% agarose gel run in Tris acetate-EDTA (TAE) buffer and stained with ethidium bromide. The gel was visualized under ultraviolet transillumination. Also, a molecular size marker 100 bp DNA ladder (Cleaver Scientific, UK) was used to determine the size of the bands.
| Forward | Reveres | |
|---|---|---|
| U6 gene | CTCGCTTCGGCAGCACATATACT | ACGCTTCACGAATTTGCGTGTC |
| CagA gene | AATACACCAACGCCTCCAAG-3’ | TTGTTGCCGCTTGCTCTC |
| Mir-21 gene | GCTTATCAGACTGATGTTGACTG | CAGCCCATCGACTGGTG |
List of Primers
3.6. RNA Extraction and Detection of mir-21
Micro RNA was extracted from the gastric biopsy samples using Bio basic kit (Canada inc, kit) in terms of the manufacturer’s instruction. Moreover, DNase (deoxyribonuclease) was administered to remove any any contamination of DNA from the extracted RNA, which was carried out in total volume 10 microliter containing 1λ MgCl2, 0.5λDDNase, extracted RNA, and DEPC water. Afterward, they were incubated for 30 min at 37°C following by an incubation for 10 min at 65°C after the addition of 1λ EDTA. As the RNA is unstable, cDNA was immediately synthesized using Maxime RT PreMix (oligo DT primer.iNTron, Korea). Accordingly, the protocol was carried out according to the manufacturer’s structure. Detection of mir-21 was performed using real Q plus 2× Master Mix Green, without using ROX Kit (AmpliQon, Denmark) according to the standard protocol, which was carried out in a total volume 23λ with 12.5λ master mix cyber green NO ROX, 2λ forward and reverse primers, 8.5λDEPC water. The real-time program was performed as follows: initial denaturation for 5 min at 95°C, following 25 cycles with denaturation for 20 sec at 95°C, annealing for 30 s at 60°C, and extension for 20 s at 72°C. Quantitative real-time data were analyzed using 2-ΔΔCT method. Also, the mir-21 was amplified with U6 gene as an internal control. Primers sets for each gene are listed in Table 2.
3.7. Statistical Analysis
All statistical analyses were performed using SPSS 18.0 software (SPSS Inc, Chicago, and IL., USA). The chi-square (χ2), one-way ANOVA test, and odds ratio were determined for the H. pylori presence and histopathological finding. The correlation of gene expression, type of gastric diseases, and the subjects’ lifestyle, were analyzed using unpaired t-test. A P value less than 0.05 was considered as statistically significant.
4. Results
The demographic characteristics of the patients as well as the source of their infection were documented. The results show that 54.3% and 45.7% of the participants were female and male, respectively, with the mean age of 50.53 ± 16.91 years old. Moreover, the lifestyle habits such as smoking, diets, and history of disease for each patient are shown in Table 2. About 10 out of 50 samples were successfully isolated from Brucella blood agar (Figure 1). The frequency of positive H. pylori culture was 14% (10/70). Also, pathology examinations showed that 32 (45.7%) participants were infected by H. pylori (Figure 2A and B). Also, 74.3% and 57.1% of the participants were suffering from gastritis and reflux, respectively.
4.1. Helicobacter pylori Identification and CagA Gene by PCR
The results of PCR product of H. pylori identification kit are shown in Figure 3. Also, the CagA gene of PCR product in 1.5% agarose gel is shown in Figure 4.
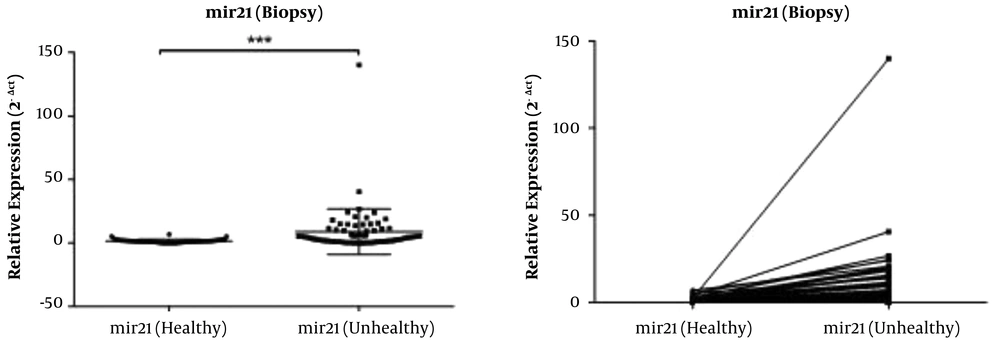
Comparative RT-qPCR experiments of miR-21 level in infected samples with Helicobacter pylori and healthy samples. The level of GAPDH and U6 were used as internal controls for normalization of miRNA expression. Data are reported as mean value ± SE (n = 3). Student’s t-test was used for data analysis. ***, Represents P value < 0.001.
4.2. Expression of mir-21 in Unhealthy and Healthy Tissue Samples
The miRNA RT-qPCR experiments were carried out on antrum biopsy samples and the adjacent healthy samples obtained from the participants, which showed that the expression level of mir-21 was significantly higher in the infected samples compared to the healthy samples (P < 0.001) (Figure 4). Thus, our data confirmed that, mir-21 expression was significantly up-regulated in the infected samples with H. pylori compared to the healthy samples.
Expression of mir-21 in the blood of the patients infected with H. pylori and healthy subjects real-time PCR analysis showed that the mir-21 level was numerically up-regulated compared to the H. pylori-positive and H. pylori-negative tissues. However, it was not statistically significant (P value > 0.05).
5. Discussion
Although deregulation of many miRNAs has been recently documented in various diseases extensively, the correlation between chronic bacterial infections and alteration in the expression level of miRNAs still is a challenging issue for researchers. Recently, some authors reported that H. pylori can affect the expression of various miRNAs, which may mediate the epigenetic misregulation of oncogenes and tumor suppressor genes as well as illustrating the bridge between H. pylori infection and gastric cancer (16, 17). Accordingly, a study conducted by Matsushima et al. (14) aimed to characterize the expression patterns of diverse miRNAs in gastric mucosa infected by H. pylori. Using high throughput profiling analysis, 31 miRNAs, including some members of the let-7 family, was recognized to be differentially expressed between the H. pylori-infected and H. pylori-uninfected gastric mucosa. More recently, by the use of some in vitro experiments, it was indicated that the cytotoxin-associated gene (CagA) of H. pylori can induce mir-584, mir-1290, and mir-146 (18, 19). Several studies have also revealed the specific alterations in the expression levels of some miRNAs in human gastric cancer compared to non-cancerous adjacent tissue (13, 20). The present study was designed to investigate mir-21 expression among H. pylori-infected compared to the healthy biopsy samples as well as the blood samples of endoscopy candidates.
In the current study, it was confirmed that mir-21 was significantly overexpressed in H. pylori infected samples compared to normal samples. According to these findings, mir-21 was the primary miRNA found in gastric cancer and H. pylori infected tissues (21, 22). Inflammatory cytokines, as well as NF-κB, mediated the inactivation of AP-1 transcription, which is able to induce mir-21 that could explain mir-21 overexpression during H. pylori infection (7). In a study conducted by Liu et al. (22), the expression of miRNA-21 was remarkably up-regulated in sporadic gastric tumor tissues compared to the non-cancerous tissues. Shiotani et al. (23) reported a higher expression of mir-21 in metaplastic gut mucosa compared with non-metaplastic mucosa. Correspondingly, it was shown that the eradication of H. pylori promotes miRNA deregulation in the gastric mucosa, rather than metaplastic sites. They suggested that the long-term colonization of H. pylori might induce epigenetic modifications, which cannot return to its normal status with H. pylori eradication alone (23). The overexpression of mir-21 in gastric cancer is probably related to H. pylori infection, as reported by Zhang et al. (13). Therefore, it can be concluded that H. pylori infection can up-regulate mir-21 expression followed by the development of gastric cancer.
Similar to other studies, some advantages and limitations exist in the current study. One of the advantages of this study is the use of a healthy adjacent part of each patient, as a control, to compare the miRNA-21 expression levels. Also, the expression level of miRNA-21 was detected in the blood samples of each patient. However, the main limitation of the current study was its design that was descriptive, so a prospective cohort study should be performed to support a causal relationship among H. pylori infection, miRNA expression, and the subsequent risk of gastric cancer development. Another limitation was that our experiments were not performed based on microarray miRNA expression measurements and there will be a significant indication in the expression of various miRNAs related to H. pylori infection and development of gastric cancer. Therefore, further in vitro and in vivo evaluations are required to achieve a comprehensive association among H. pylori pathogenesis, deregulation of miRNAs, and the associated abnormalities in cellular signaling in gastric cancer. Also, identification of the target genes in future studies is recommended for achieving a better understanding on the H. pylori infection and its impact on miRNA expression. Although there are some studies in this subject, most of them were done based on bioinformatics, which needs to be confirmed in vitro conditions.
5.1. Conclusions
In conclusion, it should be noted that H. pylori can alter the expression of various miRNAs in gastric epithelial cells and gastric mucosal tissues. Therefore, the altered miRNAs, especially overexpression of Oncomirs, may identify a potential link between miRNAs and pathogenesis of H. pylori-related complications. However, the way that the dysregulation of these miRNAs is involved in the pathological development of gastric cancer is not quite clear yet. So, high throughput sequencing should be performed to obtain complete descriptions of tissue miRNA dysregulation in H. pylori-related diseases.