1. Background
Staphylococcus aureus is one of the Gram-positive, facultatively anaerobic, and ubiquitous cocci that often colonizes the skin and mucous membranes (1, 2). Staphylococcus aureus is an important nosocomial opportunistic pathogen with colony stabilization and transient rates of approximately 20% and 30%, respectively (3). It is also a major cause of infection in burn patients. Among the pathogens causing infection in burn patients, S. aureus, which is a methicillin-resistant S. aureus (MRSA), has been proposed as a prominent cause of mortality (4-6). In recent years, the emergence of MRSA has been widely studied worldwide. It has been found that MRSA infections are the cause of 20,000 annual deaths in the United States. The frequency of MRSA infections in Iran was reported as 43% (7, 8). Methicillin-resistant S. aureus was formerly known as a hospital-acquired pathogen, but recently there has been an increasing trend in the reports of community-related MRSA (CA-MRSA). Some studies have reported that HA-MRSA strains are being replaced by CA-MRSA strains (9, 10).
Staphylococcus aureus has many virulence factors, including capsules, peptidoglycan, protein A, proteases, hemolysins, and leukocidin, which cause a wide range of problems, including wound infections, folliculitis, abscesses, and sepsis. This pathogen causes toxicity, endocarditis, bacteremia, osteomyelitis, and food poisoning (11, 12). Leukocidins are two-part virulence factors and pyrogenic super-antigenic toxins that can destroy host cell membranes and control immune reactions via the activation of immune cells (13). These bacteria produce seven different leukocidins, including Pantone-valentine Leukocidin (PVL), hemolysin gamma, Luk M, and Luk E/D. Moreover, PVL has been introduced as a virulence factor of S. aureus, a type of gamma toxin with two protein components called F (32 kDa) and S (38 kDa) that are controlled by the Lukf/PV and Luks/PV genes (14).
These toxins can spread among other Staphylococci due to transmission via bacteriophages. Their mechanism of action is against the outer membrane of polymorphonuclear cells, monocytes, and macrophages. Depending on the concentration of these toxins, they open calcium channels and cause apoptosis and necrosis of human leukocytes (15). The high prevalence of PVL has also been reported as a result of increasing MRSA strains (16). In addition, researchers have identified the PVL genes in CA-MRSA isolates and LukS/lukF-PV genes in methicillin-susceptible S. aureus (MSSA) (17). Staphylococcus aureus strains, which carry leukocidins, have a high level of virulence and are more common in infections such as furuncles, skin abscesses, and severe necrotic infections. In addition, S. aureus isolates containing these genes have a high potential for infection in burn patients due to their high antibiotic resistance, which, in turn, disrupts the healing process (18, 19).
2. Objectives
No comprehensive study has been recently conducted on the rate of leukocidin genes in S. aureus isolates in burn patients in Kermanshah, Iran. Thus, we aimed at determining the incidence of PVL and LucED genes in methicillin-susceptible and resistant isolates of S. aureus in burn patients who visited the Burn Unit of Imam Khomeini Hospital in Kermanshah.
3. Methods
3.1. Isolate Collection and Recognition
This descriptive cross-sectional study was performed for 11 months from December 2018 to November 2019. Sampling was performed from burn wounds of 73 patients in the Burn Unit of Imam Khomeini Hospital in Kermanshah who had not consumed any antibiotics according to patient reports and files for one week before hospitalization. The isolates were transferred to the laboratory in brain heart infusion (BHI) broth (Merck, Germany). After culture in blood agar and nutrient agar (Himdia, India), they were incubated at 37°C for 24 h. The isolates were identified by the approved biochemical tests such as Gram staining, oxidase disc, catalase, DNAase, coagulase, mannitol sugar fermentation (MSA), and novobiocin susceptibility test. For the definitive identification of S. aureus isolates, a PCR was carried out for the gene encoding deoxyribonuclease (nuc) (Table 1). After the ultimate recognition of S. aureus isolates, we kept them in a trypticase soy broth (TSB) containing 20% glycerol at a temperature of -70°C.
| Primer | Sequence (5’ - 3’) | 35 Cycles | Product Size, bp | ||
|---|---|---|---|---|---|
| Denaturation 92°C | Annealing 30 s | Extension 72°C | |||
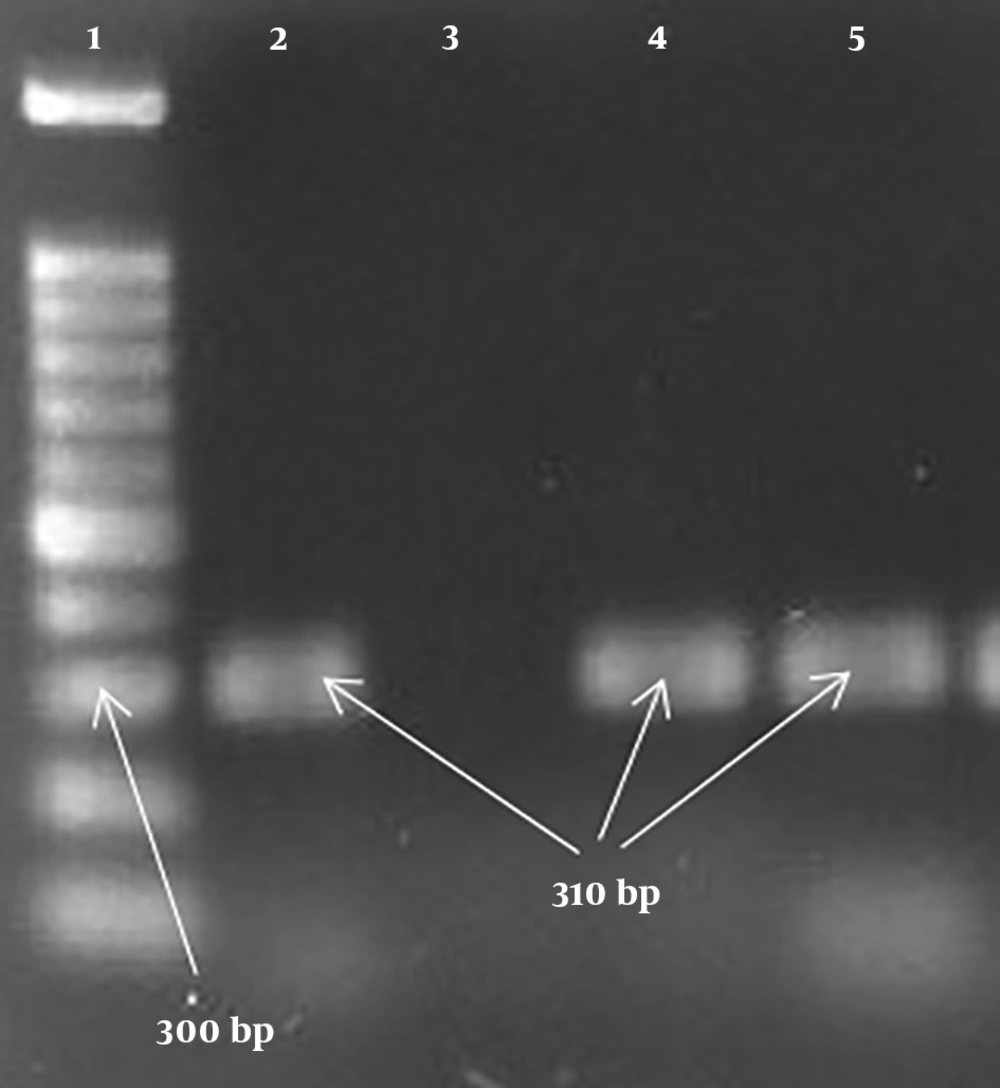
| mecA | GTAGAAATGACTGAACGTCCGATAA | 60 s | 50°C | 60 s | 310 |
| CCAATTCCACATTGTTTCGGTCTAA | |||||
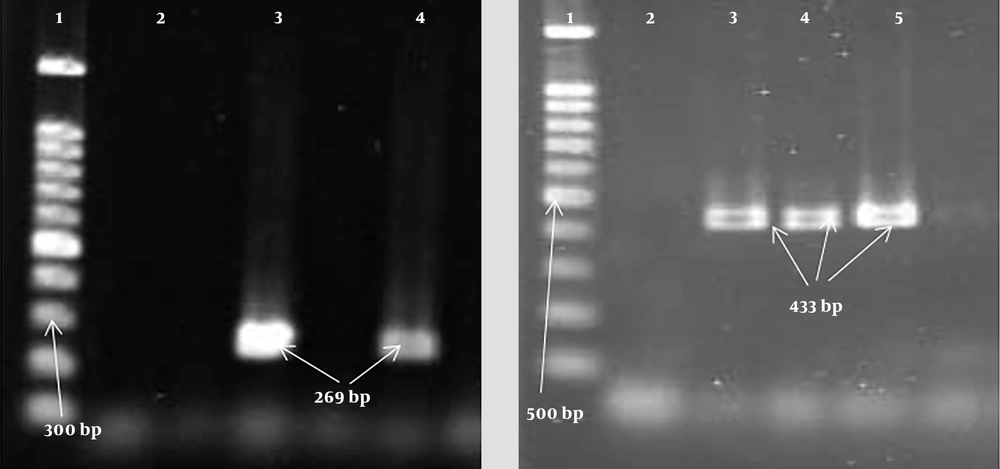
| PVL | ATCATTAGGTAAAATGTCTGCACATGATCCA | 30 s | 55°C | 45 s | 433 |
| GCATCAASTGTATTGGATAGCCAAAAGC | |||||
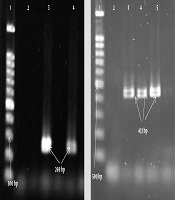
| luk-E/D | ATTCCATAGCATAAGCACTGC | 30 s | 55°C | 45 s | 269 |
| TGAAAAACCTTCAAAGTTGATACCAG | |||||
3.2. Antibiotic Susceptibility Testing
According to the guidelines put forth by the Clinical and Laboratory Standards Institute (CLSI), we used the Kirby-Bauer disc diffusion technique to perform anti-bacterial susceptibility testing. According to these studies (1, 20), the antibiotics included gentamicin (10 µg), amikacin (30), tobramycin (10 µg), rifampicin (5 µg), penicillin (5 µg), linezolid (30 µg), chloramphenicol (30 µg), clindamycin (2 µg), ciprofloxacin (5 µg), levofloxacin (5 µg), and co-trimoxazole (25 µg). We used the disk diffusion method and antibiotic discs of oxacillin (1 µg) and cefoxitin (30 µg) according to the CLSI protocol for the identification of MRSA isolates. This was accomplished by adding a direct colony of S. aureus to Müller-Hinton broth and comparing it with the 0.5 McFarland standard. The suspension was cultured on a Müller-Hinton agar medium (Himdia, India), and then antibiotic discs were placed on the medium. After 24 h of incubation at 37°C, we measured the diameters of antibiotic growth inhibition zones and compared the obtained values with the standard CLSI tables. Those isolates resistant to at least three classes of antibiotics were regarded as Multidrug-resistant (MDR) isolates. Staphylococcus aureus ATCC 25923 was used for quality control in the antibiogram test.
3.3. Polymerase Chain Reaction
To examine the existence of the methicillin resistance gene (mecA), we carried out PCR on the studied isolates (Table 1) to investigate the frequency of PVL and LucED genes with the specific primers (21). Afterward, S. aureus NCTC13300 and ACTC49775 standard strains were utilized as the positive controls to detect the PVL and LukE/D genes. First, a total genomic DNA of the bacterium was extracted by the boiling method. For this purpose, several bacterial colonies were dissolved in 0.5 mL of sterile distilled water (DW) and centrifuged at 7,000 g for one minute, followed by five minutes of boiling and cooling. The supernatant solution was transferred to new Eppendorf tubes for PCR. The concentration of DNA samples was quantified by measuring optical density (OD) at 260 nm using a NanoDrop Synergy HTX (Bio Tek Instrument, Inc. Highland Park, USA) equal to 34 pmol/µL. Also, the DNA purity at 260/280 nm was 1.84.
The PCR of the genes was performed separately with a thermal cycler (Biorad, USA). The final volume for a typical 25 µL reaction was as follows: 12.5 µ of Master Mix (SinaClon Company, Iran), 3 µL of bacterial DNA, 1 µL of each primer, and 25 µL of DW. Moreover, the general formula started with an initial denaturation step at 97°C for 6 min, followed by 35 cycles according to Table 1, and finally, 10 minutes of final extensions at 72°C. The PCR products were analyzed on a 1.5% agarose gel at 85 V for 45 min. They were stained with an ethidium bromide solution and visualized under UV light in a gel documentation system (Biorad, USA).
3.4. Statistical Analysis
Fisher’s exact test and chi-square test were run to analyze the data in SPSS, version 20. A P-value of less than 0.05 was considered statistically significant.
4. Results
A total of 73 S. aureus samples were collected from burn patients [54 (74%) male patients and 19 (26%) female patients] with a mean age of 38.14 ± 16.9 years. All the isolates were from burn wound samples. It was found that the highest antibiotic resistance of the isolates was to penicillin (100%) and gentamicin (63%), whereas the lowest resistance was to linezolid (1.4%) and chloramphenicol (12.3%). The frequencies of MRSA and MSSA isolates were 58.9% (43 isolates) and 41.1% (30 isolates), respectively. In MRSA isolates, the highest antibiotic resistance was to penicillin (100%) and gentamicin (81.4%), whereas the lowest resistance was to linezolid (2.3%) and chloramphenicol (16.3%) (Table 2). The frequency of MDR isolates was determined to be 95.9% (70 isolates). Moreover, the PCR results showed that the highest frequency among the genes was related to the LucED gene with a frequency of 76.7% (56 isolates). In addition, the frequency of the PVL gene equaled 27.4% (20 isolates). Based on the results, there was a significant relationship (P < 0.05) between the existence of the LucED gene and resistance to some antibiotics, including amikacin, tobramycin, and ciprofloxacin (Table 3). The highest frequencies of the LucED and PVL genes were observed in MRSA (81.4%) and MSSA (40%) isolates, respectively (Table 4). Figures 1 and 2 depict the PCR results for the mecA, LucED, and PVL genes.
| Antibiotics | MRSA (43 Isolates) | MSSA (30 Isolates) | P-Value | ||||
|---|---|---|---|---|---|---|---|
| R | S | I | R | S | I | ||
| Gentamicin | 35 (81.4) | 5 (11.6) | 3 (7) | 11 (36.7) | 19 (63.3) | 0 | 0.0001b |
| Amikacin | 23 (53.5) | 20 (46.5) | 0 | 9 (30) | 21 (70) | 0 | 0.039b |
| Tobramycin | 21 (48.8) | 22 (51.2) | 0 | 5 (16.7) | 23 (76.6) | 2 (6.7) | 0.149 |
| Rifampin | 21 (48.8) | 22 (51.2) | 0 | 6 (20) | 26 (80) | 0 | 0.011b |
| Penicillin | 43 (100) | 0 | 0 | 30 (100) | 0 | 0 | - |
| Linezolid | 1 (2.3) | 42 (97/7) | 0 | 0 | 30 (100) | 0 | 0.589 |
| Oxacillin | 19 (44.2) | 24 (55.8) | 0 | 14 (46.7) | 16 (53.3) | 0 | 0.511 |
| Ciprofloxacin | 20 (46.5) | 21 (48.8) | 2 (4.7) | 20 (66.7) | 10 (33.3) | 0 | 0.404 |
| Levofloxacin | 21 (48.8) | 20 (46.5) | 2 (4.7) | 10 (33.3) | 19 (63.3) | 1 (3.4) | 0.189 |
| Clindamycin | 21 (48.8) | 18 (41.9) | 4 (9.3) | 10 (33.3) | 19 (63.3) | 1 (3.4) | 0.64 |
| Chloramphenicol | 17 (16.3) | 36 (83.7) | 0 | 2 (6.7) | 28 (93.3) | 0 | 0.195 |
| Cotrimoxazole | 13 (3.2) | 30 (69.8) | 0 | 5 (16.7) | 25 (83.3) | 0 | 0.147 |
aValues are expressed as No. (%).
bSignificant.
aSignificant.
| Genes | ||||
|---|---|---|---|---|
| LucED (N = 56) | PVL (N = .20) | |||
| Isolates (No.) | Positive | Negative | Positive | Negative |
| MRSA (43) | 35 (81.4) | 8 (18.6) | 8 (18.6) | 35 (81.4) |
| MSSA (30) | 21 (70) | 9 (30) | 12 (40) | 18 (60) |
aValues are expressed as No. (%).
5. Discussion
Methicillin-resistant S. aureus can be isolated from nosocomial infections. Moreover, it has long been known to cause serious infections in burn patients and escalate their mortality rates. According to the analyses conducted in this study, the frequency rates of MRSA and MSSA isolates were 58.9% and 41.1%, respectively. Concerning studies performed in Iran, the prevalence of MRSA ranged from 29% to 40% in non-burn patients and 60% to 90% in burn patients, which indicates a higher prevalence of MRSA strains in burn patients than in non-burn ones. Indeed, these results are in line with our findings (22-26).
Other studies conducted abroad have shown different results regarding the prevalence of MRSA. A study conducted by Jiang et al. on burn samples for five years revealed that out of 259 S. aureus isolates, 239 (92.28%) were MRSA, which is higher than our finding. Also, in Bangladesh, Australia, and China, the rates of MRSA were much lower than our obtained results (27-29). In MRSA isolates in this research, the highest antibiotic resistance was to penicillin (100%) and gentamicin (81.4%), and the highest sensitivity was observed to linezolid (97.7%) and chloramphenicol (83.7%). Other studies in Iran and abroad have reported high levels of resistance in MRSA isolates to different antibiotics, including penicillin, gentamicin, and ciprofloxacin (30-32). In a study carried out by Moosavian et al. (25), all MRSA isolates were sensitive to linezolid and vancomycin (25).
In this study, among 73 S. aureus isolates obtained from burn samples, the frequency rates of LucED and PVL genes were determined to be 76.7% and 27.4%, respectively. In two studies conducted in Iran that were similar to our study, the prevalence of LucED was observed to be higher than that of PVL. Khosravi et al. (22) reported that the prevalence rates of these two genes in S. aureus isolates isolated from burn samples were 66.26% and 7.2%, respectively. In addition, Havaei et al. (21) reported the frequencies of 73% and 24.8% for LucED and PVL, respectively, which are consistent with our results. Methicillin-resistant S. aureus strains carrying the PVL gene cause health problems; thus, they should be detected quickly. The first MRSA strain to carry PVL was observed in the late 1990s, and it has spread worldwide in recent years. Furthermore, PVL is a virulence factor that is transmitted to other Staphylococci through bacteriophage and contributes to the increased virulence and pathogenicity of S. aureus (33-37). In general, studies have shown that the incidence of the PVL gene in HA-MRSA is lower than the incidence of this gene in CA-MRSA. These results are completely in line with our findings, as our samples were obtained from a hospital setting (38).
In this study, the highest frequencies of the LucED and PVL genes were observed in MRSA (81.4%) and MSSA (40%) isolates, respectively. A study by Enwuru et al. (32) showed that 100% of MRSA samples carried the LucED gene, and 98% of MSSA strains had the PVL gene. In addition, a study by Khosravi et al. (22) reported the prevalence of the LucED gene to be 66.26% among MRSA strains, and the prevalence of the PVL gene in MSSA isolates was equal to 33.3%. One of the criteria for identifying nosocomial MRSA isolates is the lower prevalence of the PVL gene. In our study, the prevalence of this gene was lower than that of LucED, indicating the incidence of nosocomial infections among burn patients.
5.1. Conclusions
Despite the higher drug resistance of MRSA isolates than MSSA isolates, the frequency of the PVL gene was higher in MSSA isolates than in MRSA isolates, which indicates that burn patients in this study had acquired these infections in the hospital. The high frequency of leukocidin genes was also explained by the inherent nature of this toxin. These results can be justified by the fact that in sites such as burn wounds, bacterial death is prevented due to the active role of white blood cells, including neutrophils in wound healing, the cascading process of inflammation, and lysis of white blood cells. Considering the potential for life-threatening infections caused by the mentioned isolates and the possibility of the early detection of MSSA isolates based on the PVL gene, a wide range of nosocomial infections could be prevented by a timely diagnosis and appropriate treatment.