1. Background
Severe acute respiratory syndrome Coronavirus 2 (SARS-COV-2) is a virus with positive single-stranded RNA as a genome and is responsible for the worldwide COVID-19 pandemic that started in Wuhan, China. SARS-CoV-2 is a beta-coronavirus of the Coronaviridae family (1). The genome structure and sequence of SARS-CoV-2 indicate high similarity with those of SARS coronavirus 1 and have several structural and nonstructural proteins that have a pivotal role in virus replication, pathogenicity, and transmission (2, 3). RNA synthesis is a necessary process that involves several steps, including genome replication and sub-genomic RNA transcription. Viral RNA synthesis is not a perfect process, and some errors such as point mutation and recombination can occur during the process, which can affect the detection process by PCR detection kits (4).
Some processes, such as sequencing the SARS-CoV-2 genome in patient's samples, provide beneficial information about viral genome characteristics, reducing false negatives, and increasing the sensitivity and specificity of real-time PCR kits (5). Orf 1ab, N, and RdRp are three regions mainly used for virus detection in different PCR diagnostic kits (6). In some studies, the S region was also used for detection, but the false-negative rate increased compared to that in the other regions (7, 8). On the other hand, the sensitivity of the PCR test is another factor that can affect the rate of false negative. Several studies have revealed that real-time PCR is not the best technique to diagnose infectious agents such as SARS-CoV-2. Nested PCR can indicate sensitivity ten times better than real-time polymerase chain reaction (RT-PCR) (9). A decrease in the false-negative rate can help prevent virus transmission through the community and disrupt the replication cycle of the virus.
2. Objectives
This study aimed to evaluate samples of patients with symptoms of COVID-19 but negative real-time PCR results. Of interest was also to test the samples again to detect the presence of viral RNA using the nested PCR method.
3. Methods
One hundred eighty-four nasopharyngeal swab samples of patients with negative SARS-COV-2 PCR results were collected and stored at -80°C in the laboratory. All the cases had negative real-time PCR test results for SARS-CoV-2 but had symptoms like fever, sneezing, myalgia, and headache. All the specimens were tested using a real-time PCR kit (Pishtaz Teb, Iran) that targeted the RdRp and N genes of the detected viral genome.
3.1. Primer Design
The outer set of PCR primers was used for the previous study (10). Inner primers were designed based on first-round products using NCBI primer blast software. All the primers were then checked using in silico PCR and primer blast software. The primers are indicated in Table 1.
| Gene and Amplicon Size | Primer Sequence |
|---|---|
| Orf1ab | |
| 200bp | Outer F: GTTACCTTCTCTTGCCACTGT |
| Outer R: TCCTAGCACCATCATCATACA | |
| 120bp | Inner F: TGACATGGTTGGATATGGTTGA |
| Inner R: CATCATCATACACAGTTCTTGC | |
| N | |
| 151bp | Outer F: CCTCTTCTCGTTCCTCATCAC |
| Outer R: CTCTCAAGCTGGTTCAATCTGT | |
| 80bp | Inner F: CAACTCCAGGCAGCAGTAG |
| Inner R: CAGCAGCAAAGCAAGAGC | |
| RdRp | |
| 190bp | Outer F: ATCTCACTTGCTGGTTCCTAT |
| Outer R: TAGTCCTCACTTCTCTCAAAGA | |
| 160bp | Inner F: GGTCCTATTCTGGACAATCTAC |
| Inner R: GTCCTCACTTCTCTCAAAGAAA |
3.2. RNA Extraction
The extraction process was done using the viral RNA extraction kit (Rojeh, IRAN) according to the manufacturer's instructions. Briefly, 560 µL lysis buffer supplemented with carrier RNA and 140 µL of viral transport media were mixed and incubated at room temperature for 10 min. After incubation, 560 µL ethanol was added to the mixture, which was then transferred to the filter tube after the vortex. The filter columns were washed with wash buffers 1 and 2, and finally, the nucleic acid was eluted into a 60 µL elution buffer.
3.3. RT-Nested PCR
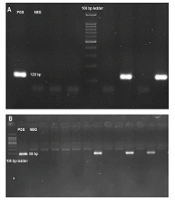
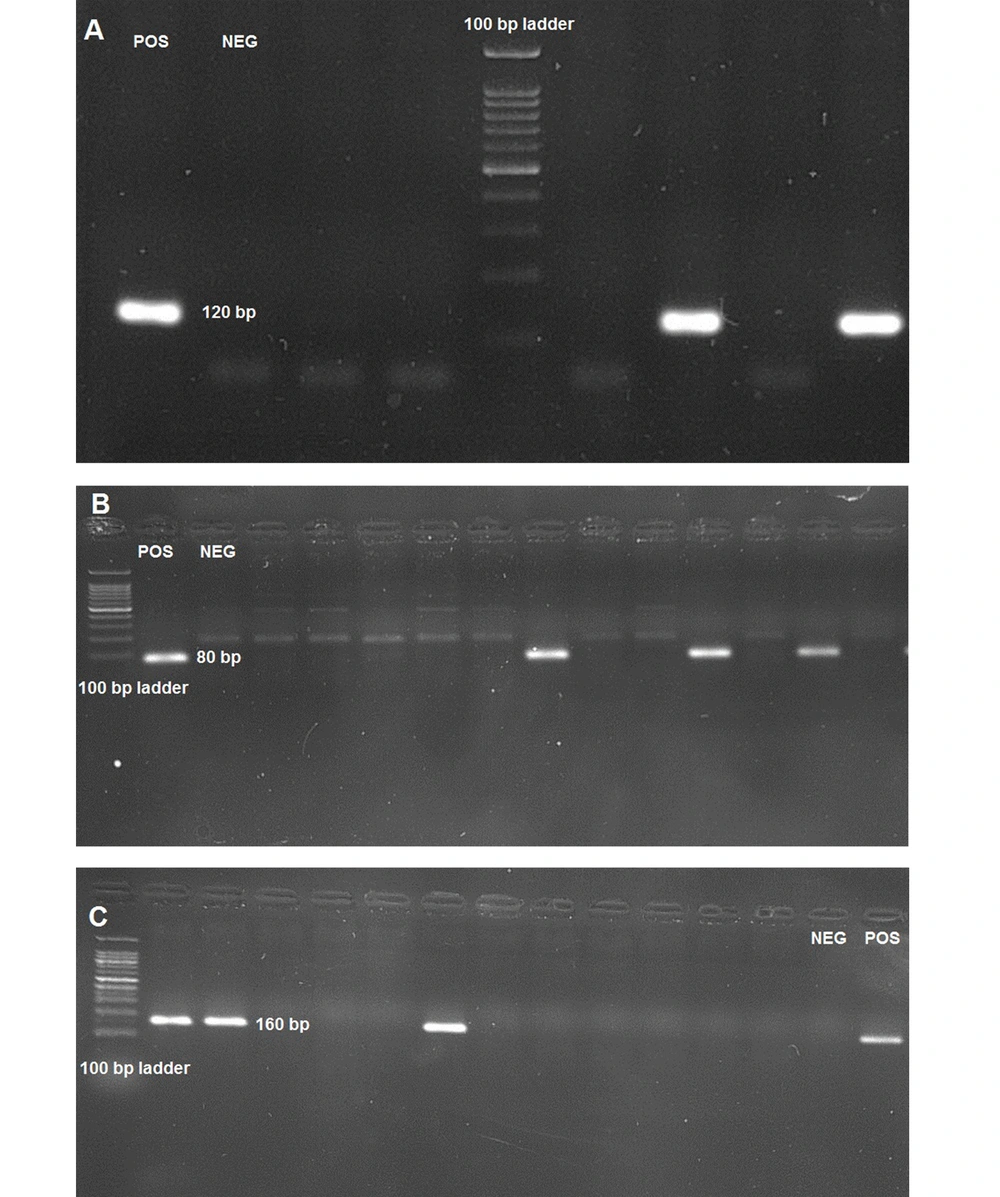
All the extracted negative samples were subjected to cDNA synthesis using First Strand cDNA Synthesis Kit (SinaClon, Iran) according to the manufacturer's instructions. The synthesized cDNA was subjected to the first round of nested PCR with external sets of Orf1ab, N, and RdRp primers to evaluate potential positive samples among the negatives. The PCR condition was as follows: (1) 10 pmol of each primer; (2) 12.5 µL of hot start 2xPCR master mix (amplicon, Denmark); (3) 2.5 µL synthesized cDNA as template; and (4) water were mixed in a PCR tube to complete 25 µL PCR reaction and subjected to 35 cycle PCR with the following program: (1) initial denaturation at 95°C for 2 min; (2) denaturation at 94°C for 30 sec; (3) annealing at 55°C for 30 sec; and (4) extension at 72°C for 30 sec. Second-round PCR was done the same as the first-round except for primers and template measurement. The inner set of primers was used, and 1µL of the first-round PCR product was added to the second-round PCR mix as a template. The second-round PCR products were subjected to electrophoresis on 2% agarose gel for gene-specific amplicon detection. Amplicons with sizes of 120 bp, 80 bp, and 160 bp were detected for Orf1ab, N, and RdRp genes, respectively. A positive real-time PCR result sample was used as a positive control, and distilled water was used as a negative control.
4. Results and Discussion
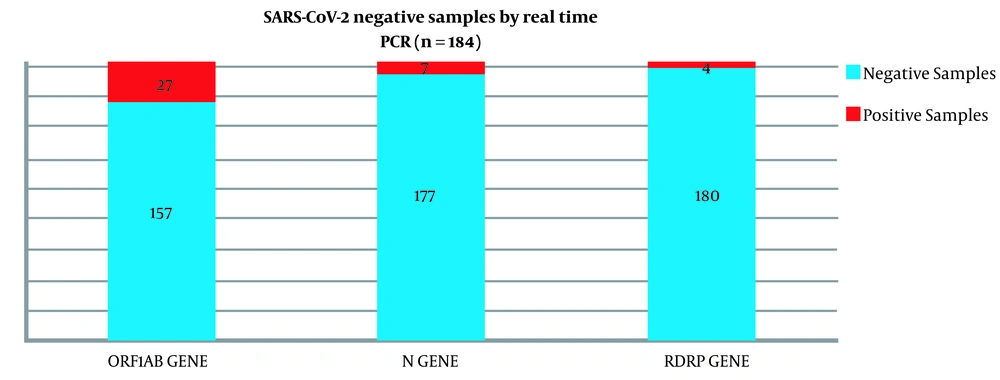
In previous reports, the Orf1ab region showed better results in SARS-CoV-2 detection (6, 10). Thus, Orf1ab was firstly used to evaluate negative samples. Among 184 negative swab samples from SARS-CoV-2, 27 (14.6%) cases were positive for the Orf1ab gene. These samples were tested by N and RdRp primer sets. Seven (3.8%) cases were positive for the N gene, and four (2.1%) cases were positive for the RdRp gene (Figure 1). Best results in SARS-CoV-2 detection using nested PCR were obtained with an Orf1ab primer. Electrophoresis results of all the genes are observable in Figure 2. SARS-CoV-2 pandemics put high and unpredictable pressure on different aspects of human life, such as health, economy, and social relationships. The best way to control the pandemic based on the WHO recommendations is to consider social distance and vaccination (11). However, fast and precise virus detection is still essential to quarantine infected people and prevent fast transmission in the community (12).
The most reliable and fast technique to detect the SARS-CoV-2 viral genome is real-time PCR, which is based on detecting different viral genes such as RdRp, N, S, Orf1ab, and E. Among them, Orf1ab, N, and RdRp are very frequent in different real-time PCR kits (13). One of the recurring problems in the SARS-CoV-2 detection is the false-negative rate. This rate highly causes sample kind (swab sample), sampling site, sampling technique, and sensitivity of PCR kits (13-16). In this study, 184 samples with negative real-time PCR test results were collected and tested again using nested PCR as a much more sensitive technique than real-time PCR to evaluate possible positive samples among negative real-time PCR samples.
Our results showed that 14.6% (27 samples) of the 184 cases with negative real-time PCR test results were positive with nested PCR using the Orf1ab primer set. Among the 27 samples, seven (3.8%) cases were positive with the N primer set, and four (2.1%) cases were positive using the RdRp primer set. These results indicated a 14.6% false negative in real-time PCR results of the Pishtaz Teb SARS-CoV-2 detection kit. Our results are in line with the results from Davda et al.'s study. They reported 13% false-negative among samples with negative real-time PCR results (17). Also, Wang et al. reported 14 positive real-time PCR results among 156 samples (8.97%) with negative real-time PCR results (9). The difference between the results can be because of differences in real-time PCR kit sensitivity, PCR instrument detection sensitivity, and region of viral genome targeting to primer design.
Wang et al. (9), in their study, used a Sansure qPCR kit for virus detection. In this kit, Orf1ab and N regions were targeted for primer design, but in the Pishtaz Teb real-time PCR kit used in this study, RdRp and N genes were targeted for primer design and detection. Some studies revealed that Orf1ab was a better choice than RdRp for primer design and indicated a better positivity rate, which is in line with our results (18). These results reveal that nested PCR can be a good alternative to real-time PCR because it can be up to 10 times more sensitive than PCR (9). The most significant negative aspect of the nested PCR technique is the risk of contamination. Other studies resolved this problem using a single tube real-time nested PCR (19-21). These approaches can decrease the false-negative rate of SARS-CoV-2 detection, thereby decreasing the transmission rate. Sample size, use of only three genes for virus detection, and sequencing could be mentioned as limitations of the study. Sequencing could be considered to detect possible point mutations, insertion, and deletions affecting PCR kit detection.
5. Conclusions
RT-nested PCR can be more sensitive than real-time PCR and reduce the false-negative rate.