1. Introduction
Helicobacter pylori are spiral-shaped, microaerophilic, Gram-negative bacteria, and the main cause of gastroduodenal diseases (1). The prevalence of H. pylori infection in some developing countries is more than 80% and below 20% in some developed countries (2). Man is a major reservoir of the bacteria and colonization remains for a lifetime unless treated. The mode of transmission of H. pylori is from person to person with 2 route of transmission being proposed: fecal-oral, oral-oral, and/or stomach-oral routes. Studies suggest that the infection transmits through saliva and dental plaque, and normal and diarrheal stools (3). However, several studies show that low standard of living and crowded households are the major risk factors for contracting H. pylori infection (4, 5). Researches suggest environmental reservoirs such as food, water and domestic animal for H. pylori. To support this hypothesis, numerous studies are carried out to report the presence of H. pylori in such environments (6-10).
The presence of Helicobacter spp. is investigated in natural environmental waters, including ground water, fresh water streams, and off-shore marine waters using molecular technique, but it is rarely isolated from these kinds of samples (11-16). Since H. pylori have the propensity for exhibiting 2 forms: spiral and coccoid. The spiral-shaped form is metabolically active while the coccoid shape is formed under stressful conditions (17, 18). Some authors consider the coccoid form as dormant and viable (17, 19-21). Morphological change from spiral to coccoid makes the organism non-culturable (22). The coccoid form can change into the spiral form under certain conditions and this supposition increases the risk of H. pylori transmission through water. Studies show that the coccoid form of H. pylori, with a smaller amount of DNA and mRNA relative to the spiral form, has oxidative metabolism and respiration (18, 23). Therefore, water can play a major role in H. pylori transmission and the bacteria can survive as long as 20 days in distilled water, while their pathogenic ability remains (24).
The survival of H. pylori in water may also depend on specific factors such as the presence of free living amoeba and zooplanktons (25). Some studies show that H. pylori can be incorporated into the biofilm, which is an important factor for successful survival in the aquatic environment (26, 27). For successful long-term colonization in the human stomach, H. pylori harbors a set of bacterial virulence determinants; Cytotoxin-associated antigen (cagA) is 1 of widely disseminated genes with 90% prevalence among Asian population and is believed to increase the risk of gastric cancer (28). Therefore, the presence of this gene in water could be a risk factor for cancer development.
In the current study, the following hypothesis was devised based on the author’s previous studies: the presence of H. pylori was detected using PCR method targeting ureC gene. Therefore, the current study was used to determine if the analytical method commonly used for other bacteria can be used to evaluate and estimate the presence of H. pylori in water. Also, in the current study, cagA and 16s rRNA genes were targeted by PCR, and ureC for the Loop-mediated isothermal amplification (LAMP) method. For this purpose, a highly specific PCR method was developed and then the obtained results were compared with those of LAMP to evaluate the sensitivity and specificity of the LAMP method to detect H. pylori DNA in water or human samples; where a very high rate of H. pylori infection is recently shown (29, 30). In addition, samples were collected and analyzed with rigorous controls for false positive or negative results.
2. Methods
2.1. Sample Collection and Preparation
As mentioned in the authors’ previous study, samples were collected from urban tap and well water sources in Kermanshah from June to November 2012 (12). Kermanshah is a mountainous city with mild climate, and is the capital city of the province located in the Western part of Iran (31). The drinking water supplies Kermanshah come from 133 wells and Gavshan dam, which is connected to 21 reservoir tanks and distributed around the city (32).
2.2. DNA Preparation
The drinking water samples were obtained from taps of all water supplies. For each sample, 1 liter of water was obtained and filtered through 0.45 μm nitrocellulose filters (12). Finally, the DNA of the filtered bacteria was extracted using QIAAmp Mini DNA kit (Qiagen, Germany), according to the manufacturer’s instructions. In the authors’ previous study, the presence of H. pylori was analyzed using only PCR. Here, to evaluate the presence of 16s rRNA, cagA, and ureC genes, PCR was amplified using specific primers; LAMP targeted species specific ureC gene on the DNAs extracted from the samples.
2.3. Primer Design
The primers used in PCR in the current study were retrieved from previously published articles. Oligonucleotide primers targeting ureC (GenBank accession no. AE000511) for LAMP were designed using LAMP primer design software (http://primerexplorer.jp/e/index.html). The 2 outer primers were designated as forward outer primer (UreCF3) and backward outer primer (UreCB3). The inner primers were designated as Forward Inner Primer (UreC FIP) and Backward Inner Primer (UreC BIP). The loop primers were designated as forward and backward Loop Primers, (UreC LF) and (UreC LB), respectively. Complete sets of primers and their sequences are shown in Table 1.
| Primer | Sequence (5’ – 3’) | Position | Product Size | Reference |
|---|---|---|---|---|
| HP-UreCBIP | CTCGCCTCCAAAATTGGCTTGCGATTGGGGATAAGTTTG | The current study | ||
| HP-UreCFIP | GCATATCATTTTTAGCGATTACGCTCACTAACGCGCTCACTTG | |||
| HP-UreCB3 | TCCCAAGATTTGGAATTGAAG | |||
| HP-UreCF3 | GCTTACCTGCTTGCTTTC | |||
| HP-UreCLB | TCAATTGCATGCATTCGCTCA | |||
| HP-UreCLF | CAGGCGATGGTTTGGTGTG | |||
| ureC | ureCF: CAT CGC CAT CAA AAG CAA AG | 605 - 625 | 214 | (12) |
| ureCR: AGT TTA AGG ATC GTG TTA G | 798 - 819 | |||
| 16s rRNA | Rg; GCTATG ACG GGT ATC C | 276 - 291 | 400 | (33) |
| Fg: GAT TTTACC CCT ACA CCA | 681 - 698 | |||
| cagA | F1:GATAACAGGCAAGCTTTTGAGG | 349 | (34) | |
| B1:CTGCAAAAGATTGTTTGGCAGA |
2.4. PCR Assay
Optimal PCR reaction was carried out according to the procedure laid down in a previous article of the authors. In summary, the process was carried out using a final volume of 15 μL containing 20 mM Tris-HCl (Cinacolon, Iran) , 50 mM KCl, 200 μM dNTP mix (Cinacolon, Iran), 1.5 mM MgCl2, 0.5 μM of each forward and reverse primers (Takapouzist, Iran), 1 unit Taq DNA polymerase (Cinacolon, Iran), and 5 μL of template DNA. Initial denaturation of the target DNA was at 95°C for 5 minutes as well as 214, 422, and 349 bp target sequences were amplified in the reaction mix through 35 cycles as follows: 92°C for 30 seconds, 55°C for 30 seconds (ureC), 58°C for 30 seconds (cagA), 45°C for 30 seconds (16s rRNA gene) and 72°C for 30 seconds, followed by 72°C for 5 minutes. Electrophoresis through 1% agarose gel and staining with ethidium bromide were done on PCR product.
2.5. Loop-Mediated Isothermal Amplification Assay
Loop-mediated isothermal amplification was carried out in a total volume of 30 μL reaction volume. The LAMP system contained 1.6 μM of each FIP and BIP, 0.2 μM of each F3 and B3, 0.8 μM of each LF and BF, 8 U Bst polymerase, 3 μL of genomic DNA, 2 mM each dNTP (TransGen Biotech), 0.8 M betaine (Sigma, St. Louis, MO, USA), 20 mM Tris-HCl, 10 mM KCL, 10 mM (NH4)2SO4, 12 mM MgSO4 that was mixed and mixture transferred to microtubes. The reaction mixture was heated at 95°C for 3 minutes in thermocycler and then chilled on ice, 8 U Bst polymerase (New England Biolabs, USA) were added followed by incubation at 65°C for 60 minutes, and was heated at 96°C for 2 minutes and cooled at 4°C for 5 minutes to terminate the reaction (35).
2.6. Analysis of LAMP Products
The LAMP products were centrifuged for 3 minutes at 3000 rpm, positive samples can be detected through observable magnesium sulfide (MgSO4) with sediment at the bottom of microtubes. Negative LAMP products with no observable sediment were subjected to electrophoresis through 1.5% agarose gels then visualized under UV light after staining with ethidium bromide (Cinacolon, Iran) to prove negativity.
2.7. Determination of Primers Specificity in LAMP Reaction
To determine the specificity of LAMP primers, the primers were set up in a LAMP reaction with the DNA of other bacterial species such as Citrobacter, Campylobacter, Klebsiella, Yersinia, Shigella, and Pseudomonas, in addition to Enterococcus faecalis and Salmonella typhi.
2.8. Determination of Analytical Sensitivity
Furthermore, analytical sensitivity was carried out to detect the sensitivity of LAMP. Here, a suspension of 3 day old culture of H. pylori was prepared in a PBS buffer and the number of H. pylori was counted using Petroff-Hasser counting chamber. The average number of counted bacteria was used as the basis for multiple suspensions, which was approximately 1 to 100,000 bacteria per liter of water. The sensitivity of LAMP was evaluated with a detection limit and after DNA extraction of H. pylori, the concentration of DNA was measured 3 times using the Nonodrop spectrophotometer device and the average result was considered as a base to prepare serial dilution with concentrations of 10 ng/reaction to 0.01 fg/reaction.
2.9. Detection Limit
The sensitivity, specificity, and negative and positive predicting values of LAMP method, in line with PCR, were measured.
2.10. Statistical Analysis
Data were analyzed by SPSS software version 16. To calculate the degree of agreement between the 2 methods, Kappa coefficient was used. The gold standard for H. pylori detection was PCR method. Loop-mediated isothermal amplification sensitivity and specificity, negative and positive predicting values and its agreement with PCR was calculated using Table 2.
| Gold Standard (PCR) | |||
|---|---|---|---|
| Positive | Negative | ||
| LAMP | Positive | a (True positives) | b (False - positives) |
| Negative | c (False - negatives) | d (True negatives) | |
3. Results
3.1. PCR and LAMP Results
In a total of 60 drinking water samples, 16 (26.67%) were positive for ureC by PCR, 20 (33.33%) were positive for ureC by LAMP, and 37 (61.67%) were positive for 16s rRNA by PCR. Overall, detection rate based on ureC gene by PCR and LAMP was 26%.
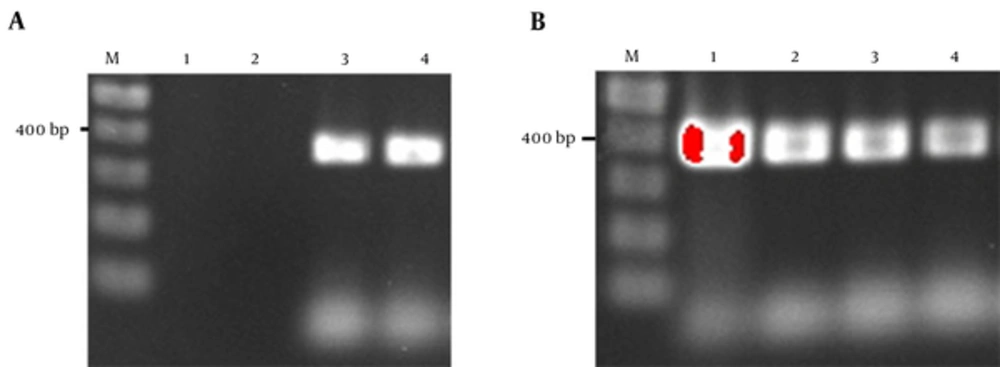
The prevalence of cagA gene among ureC positive samples was 3/16 (18.75%). But among 37 samples positive for 16s rRNA, 5 samples (13.51%) were also positive for cagA gene. The overall detection rate of cagA was 10% (6/60). Figure 1A and 1B show the PCR electrophoresis products of 16s rRNA and cagA genes.
A, Agarose gel electrophoresis of PCR products amplified from drinking water. Lane M, 100 bp DNA ladder; lane 1, negative control; lane 2, drinking water sample negative for H. pylori; lane 3 drinking water sample positive for H. pylori; lane 4, positive control; B, agarose gel electrophoresis of PCR products amplified from drinking water targeted cagA; Lane M, 100 bp DNA ladder; lane 1, positive control; lane 2, drinking water sample positive for H. pylori; lane 3 drinking water sample negative for H. pylori; lane 4, negative control.
3.2. Loop-Mediated Isothermal Amplification Specificity
The evaluation of primers specificity in LAMP reaction for ureC gene of H. pylori was done to decline any complementary relevancy between these primers and other bacterial genes in water. The results showed that the primers designed for ureC to be used in LAMP reaction were 100% specific for H. pylori.
3.3. Loop-Mediated Isothermal Amplification Sensitivity
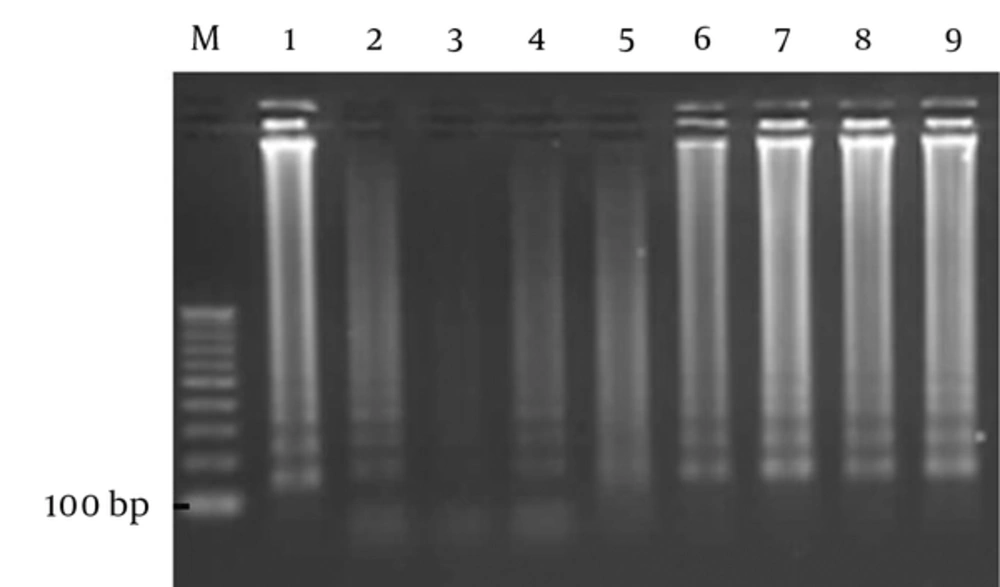
The analytical sensitivity of the LAMP method was 50 bacteria per liter with observable sediment, but after electrophoresis, the sensitivity improved to 1 bacterium per liter. The ability of the LAMP reaction to detect H. pylori, using pure DNA of the bacterium, was 10 fg/reaction (Figure 2). In the current study, the sensitivity, specificity, negative and positive predicting values of the LAMP to detect H. pylori in water samples were 100.00%, 90.91%, 100.00%, and 80.00%, respectively and the efficacy of the LAMP was 100% in comparison with that of PCR, which is the gold standard. Kappa coefficient and agreement between LAMP and PCR were 0.84 and 93%, respectively. The agreement between the 2 tests was excellent and can be used parallel to PCR.
From left to right: lane M. 100 bp DNA Vivantis ladder; lane 1, positive control (151 bp); lane 2, Fifty bacteria per liter; lane 3, negative control; lane 4, one bacterium per liter; lane 5, ten bacteria per liter, lane 6, one hundred bacteria per liter; lane 7, one thousand bacteria per liter; lane 8, ten thousands bacteria per liter; lane 9, one-hundred thousand bacteria per liter.
4. Discussion
Adequate knowledge regarding the reservoirs and modes of transmitting H. pylori could help to explain the high prevalence of the bacteria. The incidence of H. pylori is high in the developing countries (90%), whereas in the industrialized countries, the figure is lower (50%) and tends to decrease. Childhood is the critical period for infection, and transmission usually occurs from person to person (2, 10, 36, 37). In an endemic area, a common source of infection is suspected (38).
Data of the present study showed that H. pylori can be detected in Kermanshah municipal tap water and the consumption of such water could be associated with gastric colonization of the organism. These findings were in line with those of the previous studies of the authors, but further investigation is required to determine whether the organism is viable or not (39). These findings also confirmed the previous observations in Isfahan, Iran (13), Colombia (40), Peru (41), Mexico (42), England (43), Sweden (38, 39, 41, 42, 44), Japan (45), and the United States (46). The high prevalence of H. pylori detected in drinking water samples strengthens the evidence of H. pylori transmission through drinking water. Considering that the cagA is associated with increased virulence and risk of peptic ulcer and cancer, the present study was the first to report on the presence of cagA in drinking water samples. A prevalence of 13.51% of this gene in drinking water is an alarming situation. In a similar study carried out in Pakistan, the prevalence of 16s rRNA and cagA were 40% and 0, respectively (47).
The result obtained for 16s rRNA was considerably high, 61.67% and 25% (15/60) of 16s rRNA positive samples were negative for the LAMP of ureC gene. This indicates the likely presence of other Helicobacter spp. other than H. pylori in the water samples. Poor sanitation of water and allowing domestic animals, which could be a carrier of non-pylori Helicobacter species to roam near water supplies, lead to water contamination (48-52). Another probability is the presence of H. pylori that has lost its pathogenicity genes (53).
The current study was also the first to report on the possible existence of Helicobacter ssp. in water samples. Non-pylori Helicobacter species are associated with some human diseases and could exacerbate some situations such as inflammatory bowel disease (IBD), and hepatocellular carcinoma (HCC) (54-56). Among the ureC positive samples, 18.75% were positive for the cagA gene and the overall detection of cagA gene was 13.51%. No study is carried out to detect cagA gene in water, but in a previous study on gastric biopsies, in terms of prevalence, cagA was 84.4% (30).
The cagA is a 40 kbp gene located in the cag pathogenicity island (PaI) of the H. pylori chromosome (57). It is shown that the presence of cagA gene is associated with peptic ulcer disease (58), atrophic gastritis (59), and gastric adenocarcinoma (60). The cagA positive strains are more virulent than other strains (61). The presence of cagA gene in water sample could be a potential risk for cancer development in Kermanshah, Iran. All the mentioned previous studies were based on the PCR of ureC gene or 16s rRNA gene, but in the current study 2 methods of detection were considered; PCR and LAMP. In addition, various genes were employed as a target of amplification, which caused an increase in detection accuracy.
It is noteworthy that it was the first study to report on the use of LAMP reaction to detect H. pylori in water samples.
Loop-mediated isothermal amplification is easy to perform if the appropriate primers are prepared, which requires only 6 pairs of primers, DNA polymerase, a bain-marie bath, and a thermocycler for reaction. Loop-mediated isothermal amplification is 10-100-fold more analytically sensitive than PCR. Compared to the other amplification methods, the DNA amplification reaction in LAMP method is carried out under isothermal condition and the efficiency of the amplification is higher and, the detection limit is lower (62, 63). The analytical specificity of the LAMP is attributed to 6 sets of primers that recognize 8 distinct regions on the target DNA. The amplified products can be also confirmed using sequencing or digestion with restriction enzyme (64-66). Since the LAMP method is much more analytically sensitive than PCR, therefore, more positive results are obtained in the reactions.
Considering the fact that cagA is associated with increased virulence, risk of peptic ulcer and cancer, the high prevalence of H. pylori and the presence of cagA gene in drinking water is fast becoming an alarming situation. In the current study, 25% of samples were positive for non-pylori Helicobacter species. However, non-pylori Helicobacter species are linked with chronic infection of the intestinal and hepatobiliary tract. They also disturb immune responses of the intestinal epithelial cells by modulating its inflammatory response, which increases the risk of bacterial infection in the intestine. The contamination of water by these bacteria could be a potential risk to develop some gastrointestinal diseases.