1. Background
Hepatitis C virus (HCV) infection is the most important risk factor for liver function abnormalities and failure. Hepatitis C virus infection is involved in the development of hepatic diseases such as chronic hepatitis, cirrhosis, and hepatocellular carcinoma (1, 2). According to the world health organization (WHO) estimation of HCV prevalence, there are 170 - 200 million HCV-infected patients in the world (up to 3% of the world’s population), and nearly three to four million people are become infected with this virus each year (3). Although HCV infection has been found to be asymptomatic (similar to its carrier), approximately 80% of HCV patients develop chronic hepatitis and 20% suffer from chronic liver disease (4). The seroprevalence of HCV in the general population of Iran was reported to be 0.5% - 1% (5), which increases to 6.7% in hemodialyzed patients (6, 7) and 20% in multiple-transfused patients with thalassemia (8, 9).
In 1989, HCV, after its characterization, was introduced as the major cause of post-transfusion hepatitis. Hepatitis C virus is a single-stranded, positive-sense RNA virus, which belongs to the Hepacivirus genus of the family Flaviviridae with a genome of approximately 9600 base pairs. This virus is classified into six major genotypes and multiple subtypes (10, 11). These genotypes can differ by up to 50% in terms of their nucleotide sequences; also, the virus has a high propensity to mutate. These characteristics might help explain some of the difficulties in the development of effective vaccines and treatments (12). The HCV genotypes are distributed around the world (13, 14). According to the literature, HCV genotype 1 is predominant in Italy, Austria, and Mexico (15-17). On the other hand, genotype 3a is predominant in India (18), while genotypes 4, 5, and 6 are commonly detected in Egypt, South Africa, and Southeast Asia, respectively (19). Based on various studies, HCV infection is a highly common problem in Iran.
In a previous study performed in Southwest of Iran indicated that the dominant genotypes in this region are 1a and 3a (20). According to a study in central Iran, the distribution of HCV 1a/b and 3a genotypes were 50% and 18.2% among patients, respectively (21). In addition, 3a genotype was the most common HCV genotype in the Northwest of Iran (22). In a population-based study in Iran, 1a and 3a genotypes were found to be predominant (23). On the other hand, the most common HCV genotype was 1a/b among Iranian hemodialysis and thalassemia patients (24, 25), while in non-uremic patients 3a was the most prevalent genotype (24). There is a relationship between HCV genotype and response to antiviral therapy; accordingly, genotyping is prior to treatment. Based on previous findings, HCV genotype 1 does not respond to therapy, similar to genotypes 2 and 3 (26).
In addition, genotype 1 may be associated with severe liver disease and a high risk of hepatocellular carcinoma (14, 27). In former study, the serological findings were different from the results of the molecular assay (28, 29). Also, in a previous study performed in Iran, 93 (74%) out of 126 anti-HCV-positive patients were positive on the molecular assay (30). The gold standard for the detection of HCV infection seems to be antibody screening, combined with the PCR test (31).
2. Objectives
In this study, we aimed to identify HCV RNA and perform genotyping in anti-HCV-positive patients through polymerase chain reaction (PCR). We compared the results of PCR test with those of ELISA test among patients referred to the healthcare centers of Mazandaran province, situated in North of Iran, during 2015 - 2016.
3. Methods
3.1. Samples
This cross-sectional study was performed during 2015 - 2016; blood samples were collected from 157 anti-HCV-positive patients who were referred to the healthcare centers of Mazandaran province, situated in North of Iran. The blood samples were transferred to the department of virology at Mazandaran University of Medical Sciences; subsequently, serum was separated and frozen at -70°C for 1 to 6 days. All the serum samples were examined by indirect enzyme-linked immunosorbent assay (ELISA) using a ELISA kit (Pishtaz Teb, Iran), according to the manufacturer’s instructions for reconfirmation. Afterwards, anti-HCV-positive samples were examined for HCV genotypes using RNA extraction kit with the standard protocol by using polymerase chain reaction assay (PCR).
Characteristics such as age, gender, and suspected sources of infection (high-risk sexual intercourse, intravenous drug abuse, and blood transfusion) were collected by using a questionnaire. For statistical analysis, Fisher’s exact test was performed using SPSS version 17. P value less than 0.05 was considered statistically significant. This study was reviewed and approved by the ethics committee (nr. 1767) of Mazandaran University of Medical Sciences, Sari, Iran, and written informed consent was obtained from all the patients participating in the study
3.2. RNA Extraction
RNA virus was extracted from the serum samples using a PureLinkTM Viral RNA Kit (Invitrogen) according to the manufacturer’s instructions. RNA quantity and purity were determined by Nanodrop (Biochrom WPA, Cambridge, UK). Thereafter, DNase treatment of RNA was performed using RNase free DNase I (TaKaRa Bio, Otsu, Japan) and stored at -70°C prior to use.
3.3. cDNA Synthesis
An equal amount of RNA treated with DNase was subjected to cDNA synthesis by using cDNA synthesis kit (Aria Tous Kit, Iran) based on the instructions of the manufacturer. Briefly, 10 µL of RNA was added to 1 µL Random Hexamer and 3 µL of DEPC water; then, the solution was incubated at 65°C for 5 minutes, and it was put on ice for 5 minutes. Thereafter, 10 µL of RT premix (2x) was added and the mixture was incubated at 25°C for 10 minutes. The reaction was stopped by heating at 70°C for 10 minutes.
3.4. PCR Assay
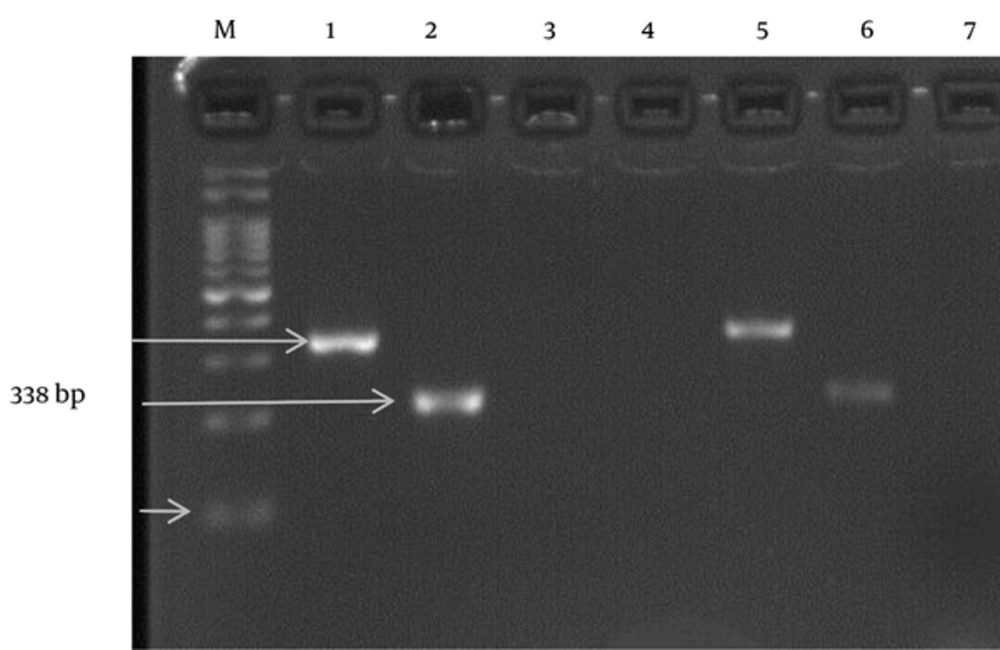
Total RNA was extracted from the serum samples and PCR assay was performed using the RNA PCR kit (Amplisense-Russia genotype kit; Invitrogen, USA). Briefly, 5 µL of cDNA sample was added to 10 µL of PCR-Mix-2Red into the surface of the wax layer of each tube. In the next stage, 1 drop of mineral oil was added for PCR; the total reaction volume was 25 µL. The PCR program for amplification consisted of initial denaturation at 95°C for 5 minutes, followed by 35 cycles at 94°C for 1 minute, at 58.5°C for 1 minute, at 72°C for 1 minute, and finally, at 72°C for 10 minutes. The PCR products were subjected to agarose electrophoresis on a 1.5% agarose gel followed by staining with SYBR Green.
4. Results
In this cross-sectional study, 157 anti-HCV-positive specimens were evaluated. The mean age of the patients was 39 ± 14.3 years (range: 20 - 68 years). In total, 110 (70.06%) anti-HCV-positive patients were male, while 47 (29.94%) cases were female. Based on the evaluations, 123 (78.34%) and 34 (21.65%) anti-HCV-positive samples were married and single, respectively. As the findings revealed, 62 (39.49%) anti-HCV-positive patients were intravenous drug users (IVDUs). In addition, 33 (21.02%) anti-HCV-positive patients had received hemodialysis for a long time. Blood transfusion was reported in 16 (10.19%) anti-HCV-positive patients.
In total, 12 (7.64%) cases suffered from thalassemia and 9 (5.73%) cases had undergone major surgery. Moreover, 25 (15.92%) anti-HCV-positive patients had an unknown source of infection. Distribution of HCV infection showed a significant difference between IVDUs and unknown group. (P = 0.182). In this study, HCV-RNA was found in 84 (53.50%) out of 157 HCV-antibody-positive patients. Also, 39 (62.90%) and 18 (54.54%) patients positive for HCV-RNA were IVDUs and had a prior history of hemodialysis, respectively. In addition, 8 (50%) HCV-RNA-positive patients had received blood transfusion, 7 (58.33%) HCV-RNA-positive patients had thalassemia, and 5 (55.56%) cases had undergone major surgery. Seven (28%) HCV–RNA-positive patients had an unknown source of infection (Table 1).
| Test | HCV Positive by ELISA | HCV Positive by PCR | P Value |
|---|---|---|---|
| Intravenous drug abusers (IVDUs) | 62 (39.49) | 39 (62.90) | - |
| Hemodialysis | 33 (21.02) | 18 (54.54) | 0.511 |
| Blood transfusion | 16 (10.19) | 8 (50) | 0.398 |
| Thalassemia | 12 (7.64) | 7 (58.33) | 0.757 |
| Surgery | 9 (5.73) | 5 (55.56) | 0.723 |
| Unknown | 25 (15.92) | 7 (28) | 0.004 |
| Total | 157 (100) | 84 (53.50) | - |
Abbreviations: ELISA: enzyme-linked immunosorbent assay; HCV, hepatitis C virus.
aValues are expressed as No. (%).
A total of 72 (85.72%) and 12 (14.28%) HCV-RNA-positive specimens were detected in men and women, respectively. The prevalence rate of HCV infection between men and women was 6 to 1. There was a significant difference in the distribution of HCV-infection between women and men (P < 0.001), although no significant difference was found between different types of genotypes among men and women. In this study, genotype 3a was detected in 53 (63.10%) samples, including 45 (84.9%) male and 8 (15.1%) female samples.
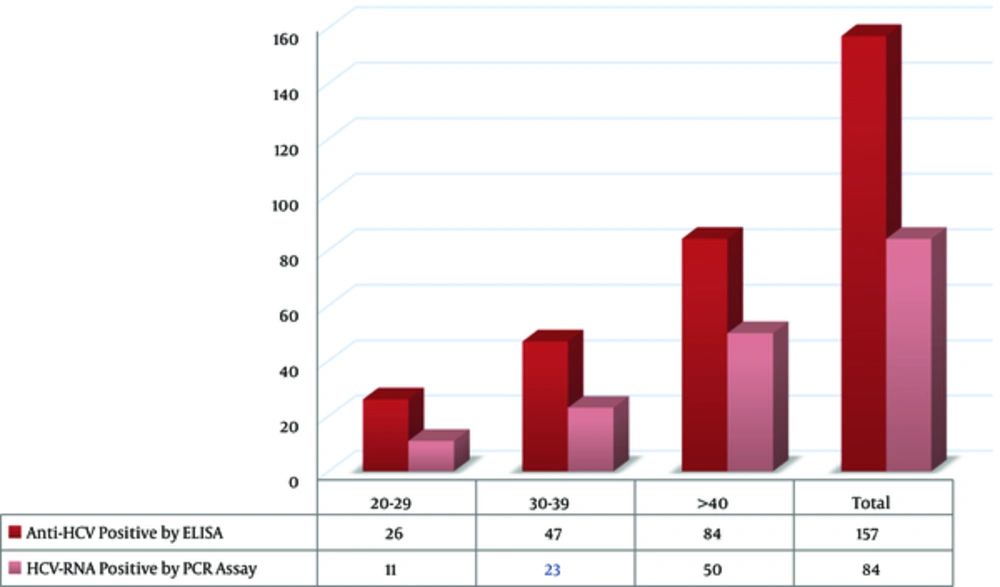
Genotype 1a/b was found in 31 (36.9%) samples, including 27 (87.1%) male and 4 (12.9%) female cases (Table 2). Also, genotype 3a had the highest prevalence in IVDUs, whereas in other patients, genotype 1a/b was predominant. The distribution of HCV genotypes did not show any significant difference between IVDUs and non-IVDU patients. Overall, 26 (16.56%) samples were within the age range of 20 - 29 years, 47 (29.93%) samples were in the age range of 30 - 39 years, 84 (53.5%) cases were above 40 years old. Based on our findings, 11 (13.09%) HCV-RNA-positive samples were within the age range of 20 - 29 years, 23 (27.38) cases were in the age range of 30 - 39 years, and 50 (59.52%) HCV-RNA-positive samples were above the age of 40 years. There was no significant relationship between the type of HCV genotype and HCV infection with age of the patients (P = 0.231; Figures 1 and 2).
| Sex | HCV 3a Genotype | HCV 1a/b Genotype |
|---|---|---|
| Male | 45 (84.9) | 27 (87.1) |
| Female | 8 (15.1) | 4 (12.9) |
| Total | 53 (100) | 31 (100) |
aValues are expressed as No. (%).
5. Discussion
Hepatitis C virus infection is a major public health problem, leading to chronic liver disease, cirrhosis, and hepatocellular carcinoma (32). Distribution of genotypes varies in different areas and patients (13). In this study, out of 157 patients with anti-HCV-positive infection, 84 (53.50%) cases were found to have HCV-RNA based on the PCR assay. The present result revealed that genotype 3a (63.1%) is the most predominant genotype in this region, followed by genotype 1a/b (36.9%).
In line with the present findings, several studies in Pakistan, India, and Malaysia showed that HCV genotype 3a is predominant in these countries (29, 33, 34). Furthermore, in a study performed in Pakistan, out of 582 anti-HCV-positive patients, 329 (56.5%) cases were HCV-RNA-positive. Genotype 3a was detected as the predominant genotype in Pakistan and was found in 68.1% of patients, followed by genotype 1a as reported in 11.3% of the cases (33); these results were consistent with the present findings. On the other hand, among 187 anti-HCV-positive samples, only 135 (72.19%) cases were HCV-RNA-positive, which is in contrast with the current study (35). Moreover, in another study performed in Isfahan province (Iran), during 2007 - 2009, among 146 anti-HCV- positive patients, 97 were determined as HCV-RNA-positive and also the most predominant genotype was genotype 3a (61.2%), followed by genotypes 1a (29.5%), 1b (5.1%) and 2 (2%), respectively (36). In contrast, in a study carried out in Southwest of Iran in 2008 - 2009, the overall prevalence rate of anti-HCV was 54% in 87 patients.
Forty two of anti-HCV patients (89.3%) were HCV-RNA-positive. In this study, the most frequent genotypes were related to genotypes 1b and 3a detected in 26.1% (11/42) and 11.9% (5/42) of the patients respectively. The results of distribution of genotypes are not compatible with our study (37). Recently Ghaderi-Zefrehi et al. (2016) performed a systematic review and meta-analysis and announced that in Turkey (82%), Israel (68%), Cyprus (68%), and Iran (55%), genotype 1 was the most prevalent HCV genotype; moreover, HCV genotype 4 was the most prevalent genotype in the Middle East (74.7%), followed by genotype 1 (15.1%) (38). According to the literature, HCV genotype 4 is the most prevalent among chronic liver patients in Egypt and Saudi Arabia (2, 38), while in a previous study in Iran, genotype 1a was the predominant genotype, followed by 3a and 1b (39); these findings were inconsistent with the current results. Furthermore, in a previous study, 1a genotype was the most frequent genotype (44.9%), followed by genotypes 3a (39.6%) and 1b (11.3%) (33, 40); these results were contradictory with our findings.
In comparison with other countries, the highest number of IVDUs has been reported in Iran; also, the prevalence of HCV infection is the highest in this population (38% - 46.6% among IVDUs) (41). Similarly, our findings revealed the highest prevalence of anti-HCV-positive cases in the IVDU group; also, genotype 3a was found to be predominant. In a previous study, in a total of 888 HCV-RNA-positive patients, the highest frequency of HCV infection was reported in the IVDU group (42), which is in line with the present findings. The genotype 1 was found as a predominant genotype among IVDUs in that study (42), this finding was inconsistent with the current results. In line with the current study, 7 (14%) out of 50 samples were determined as HCV-RNA-positive by PCR assay in Kermanshah, Iran, and genotype 3a was found to be predominant (43). In the Netherlands, HCV-RNA was detected in 57 (81.43%) out of 70 anti-HCV-positive patients based on the PCR assay (31). This finding was compatible with our results comparing serological and molecular methods. Inconsistent with the present findings, in a study in Northeast of Croatia on anti-HCV-positive patients, HCV-RNA was found in 124 (61.1%) out of 203 HCV-positive patients.
Hepatitis C virus-RNA increased in older patients compared with their younger counterparts, whereas there was no significant difference between men and women; RNA percentage was found to be 6.1% in both groups (28), which is incompatible with our findings. In addition, according to a previous study in Iran, the prevalence of HCV infection in men was higher than women (44). In this study, the highest prevalence of HCV infection was reported among men (85.5%). In conclusion, in the present study, the prevalence of active HCV infection was high among IVDUs in Mazandaran province, Iran. A high rate of HCV infection was reported in patients within the age range of over 40 years, and genotype 3a was found to be the most common genotype. Further information is required on the epidemiology of HCV genotypes, as response to treatment is reported to vary in different types of HCV.