1. Background
Pseudomonas aeruginosa is a saprophytic gram-negative bacterial species that can be isolated from soil, plants, and hospital environments (1). Pseudomonas aeruginosa-related infections cause death in patients who suffer from bacteremia (2), intensive care unit (ICU)-related septicemia (3), ventilator-related pneumonia, and even urinary tract infections (4, 5). Multiple drug-resistant (MDR) and even extensively drug-resistant (XDR) phenotypes among P. aeruginosa have been reported frequently and at elevated prevalence (4 - 60%) (6, 7). Reduced cytoplasmic membrane permeability, expression of efflux pump-related genes, the release of antibiotic-destroying enzymes such as beta-lactamases, and alginate production are the most common antibiotic resistance mechanisms among P. aeruginosa strains (8-10).
Efflux pumps are a group of proteins transferring energy packages, antibiotics, and other small molecules out of the bacterial cytoplasm or periplasmic space. Continuous transfer of antibiotics out of bacteria will increase the minimum inhibitory concentrations (MIC) for such antibiotics and antiseptic agents. It helps the bacteria continue growing while the antibiotic or antiseptic target is still present in the bacterial cytoplasm (8-10). Also, blaOXA-23, blaOXA-24, and blaOXA-40 are genes responsible for resistance against different carbapenem antibiotics (11).
Utilizing disinfectants with quaternary ammonium compounds (QAC) and biguanide compounds included is one of the most applied prevention strategies against nosocomial infection (12). Reduced susceptibility against the active agents of the mentioned antiseptics has been reported before (13, 14). Antiseptic resistance may result from the expression of chromosomal genes and plasmid-located genes, including biocide resistance genes (BRGs) and qac (15). The qacE gene has been reported as one of the genes encoding resistance against QACs and chlorhexidine digluconate (CHG) among Enterobacterales and different Pseudomonas spp. (16).
2. Objectives
The current study investigated the prevalence of antiseptic resistance genes and the resulting increase in MIC against QACs and CHG among P. aeruginosa isolates from different infections among patients in the Imam Hassan Hospital (IHH). The IHH is the biggest referral teaching and care hospital in North Khorasan province of Iran.
3. Methods
3.1. Study Samples
All P. aeruginosa isolates causing infections at different anatomical sites were collected by tracheal aspirate culture, blood culture, urine culture, and wound culture. Clinical specimens originated from hospitalized patients in various hospital wards (ICU, Cardiology, Emergency Department, Infectious Diseases Department, and Neurology) in the IHH during 2020. Bacteria were identified at the species level in the hospital laboratory, and this was confirmed in the microbiology laboratory at the Faculty of Medicine by Gram staining, oxidase testing, motility testing, and defining the ability of growth at 42°C according to Clinical and Laboratory Standard Institute (CLSI) guidelines (17). All isolates were stored at -30°C in trypticase soy broth (TSB) supplemented with 20% glycerol.
3.2. Antiseptic Susceptibility Testing
Bacterial susceptibilities to benzethonium chloride (BTC), benzalkonium chloride (BKC), and biguanide compounds such as CHG (Sigma-Aldrich, Steinheim, Germany) were determined using the Mueller-Hinton broth microdilution method (BMD) (18).
3.3. Antibiotic Susceptibility Testing
Antimicrobial susceptibility testing (AST) was performed on Mueller-Hinton agar (Merck, Germany) using the disk diffusion (Kirby-Bauer) technique, with zone size interpretation based on CLSI (2021) guidelines (17). The 11 antimicrobials agents used in characterizing the isolates of P. aeruginosa were: Carbapenems (doripenem, meropenem, imipenem), a tetracycline (tigecycline), aminoglycosides (amikacin, tobramycin), beta-lactamase/beta-lactamase inhibitor combinations (ampicillin + sulbactam, piperacillin + tazobactam), cephalosporins (cefepime, ceftazidime), and a fluoroquinolone (ciprofloxacin). Isolates were categorized as XDR when they revealed resistance to one or more antimicrobial agents in at least six categories or showed resistance to all except one or two antibiotics. Resistance to one or more agents in three or more categories was used for grouping the bacteria as MDR (19).
3.4. Detection of Genes
Chromosomal DNA was extracted using a commercial DNA extraction kit (Poyagene Azma, Iran) according to the manufacturer's instructions and kept at -30°C for further molecular investigations. All isolates were confirmed at the gene level as P. aeruginosa by detecting the gyrB gene using polymerase chain reaction (PCR) (20). All isolates were screened for antiseptic and antibiotic resistance genes such as qacE, qacEΔ1, and blaOXA-23 genes as described before (21, 22). The Supplementary File shows the primer sequences and related PCR protocols.
3.5. Typing of Bacterial Isolates
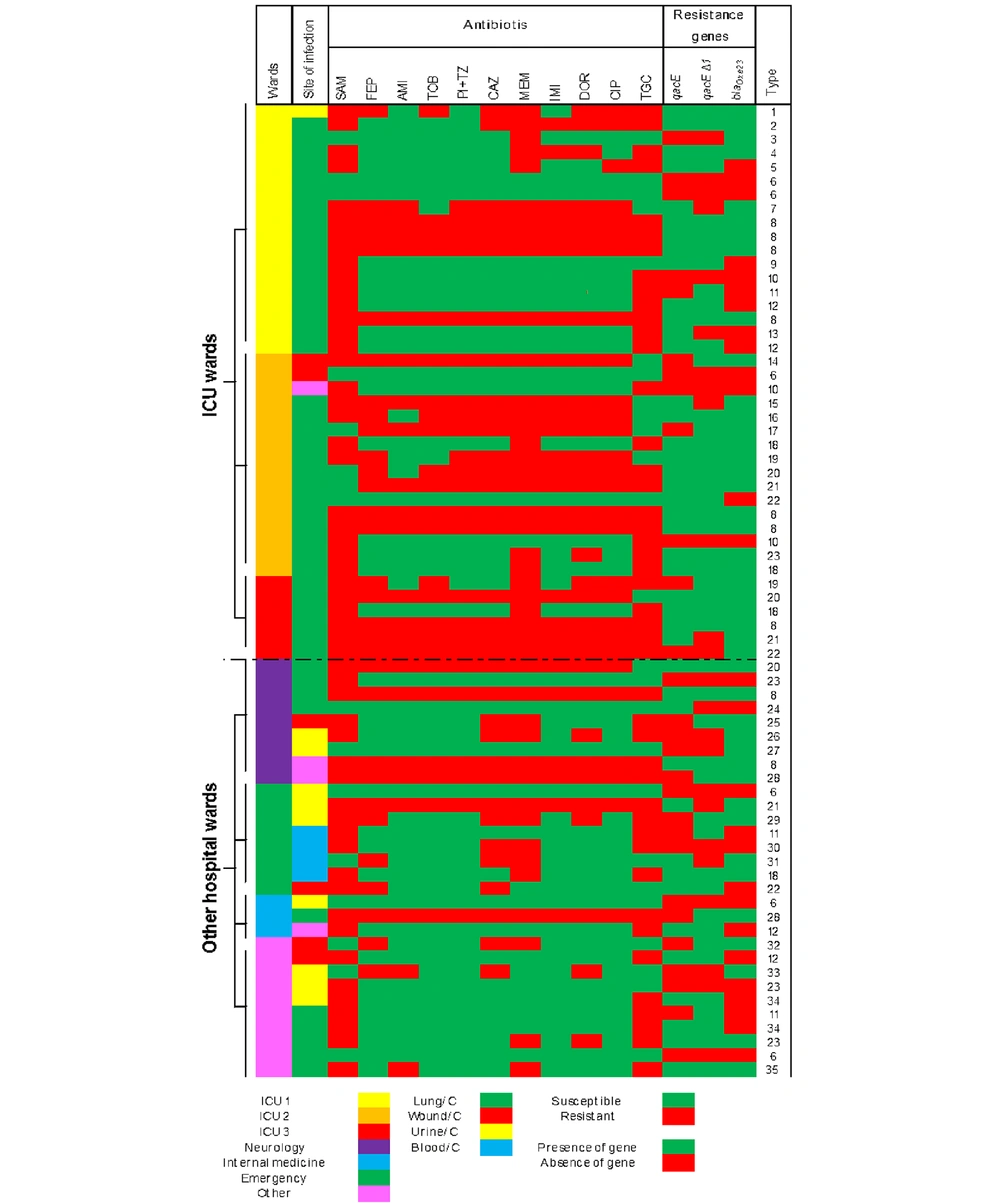
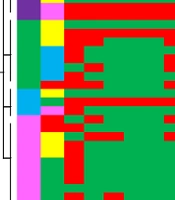
In order to verify that isolates were sufficiently heterogeneous and that past or ongoing outbreaks did not bias our study, we accumulated all data gathered above in a single figure. We defined the uniqueness of the strains by comparing phenotypic and PCR data (Figure 1). Typing the strains was performed using the combined similarities of isolates in their antibiotic resistance patterns and their resistance gene content. Similar P. aeruginosa isolates were placed under the same type and numbered accordingly.
3.6. Statistical Analysis
A one-way ANOVA test was performed to evaluate possibly significant differences using SPSS statistics 21. P-values < 0.05 were considered significant.
4. Results
From 78 documented P. aeruginosa infections detected in 2020, 70 P. aeruginosa isolates were still available and subjected to the current study. These P. aeruginosa isolates were obtained from different clinical samples such as tracheal aspirates (n = 46; 65.7%), wounds (n = 6; 8.5%), blood (n = 4; 5.7 %), and urine (n = 10; 14.2 %) (Appendix 2 in the Supplementary File). Among the infected patients, 58.5% were male (n = 41/70). The mean age of the infected patients was 63.3 (2 - 92) years, with 61.8 (2 - 92) years for males and 64.9 (42 - 87) years for females (Table 1).
The vast majority of P. aeruginosa infections occurred in the ICU (n = 43, 61.4%), comprising ICU I (n = 19, 27.1%), ICU II (n = 18, 25.7%), and ICU III (n = 6, 8.5%). An extensive diversity among ICU isolates (23 out of 43 strains, 53.4%) was spotted when an accumulation of all the strain-specific variables determined was used (Figure 1). Of note, when all isolates were considered, we documented 35 different types among all 70 isolates included. The main isolation site of P. aeruginosa isolates was lung infection (65.7%), followed by urine infection (14.2%), wound infections (8.5%), blood infection (5.7%), and other (5.7%) (Supplementary File).
4.1. Antibiotic Susceptibility Test
Among the 11 tested antibiotics, the highest resistance rate was detected against ampicillin + sulbactam (77.1%), followed by tigecycline (64.3%). The lowest resistance rate was for amikacin (32.9%) (Table 1). Carbapenem-resistant P. aeruginosa (CRPA) was detected in 48.5% of all cases.
| Resistance Phenotype | Pattern | Values | Resistant Antibiotic | No. | Sensitive | No. | Having at Least A Gene | qacE | qacEΔ1 | qacE + qacEΔ1 | blaOXA-23 |
|---|---|---|---|---|---|---|---|---|---|---|---|
| XDR | A | 14 (20) | SAM, FEP, AMI, TOB, PI + TZ, CAZ, MEM, IMI, DOR, CIP, TGC | 11 | - | - | 9 (64.2) | 2 (14.28) | 2 (14.28) | 9 (64.2) | 14 (100) |
| MDR | B | 3 (4.2) | SAM, FEP, AMI, TOB, PI + TZ, CAZ, MEM, IMI, DOR, CIP | 10 | TGC | 1 | 1 (33.3) | 0 | 1 (33.3) | 2 (66.6) | 3 (100) |
| C | 2 (2.8) | SAM, CAZ, MEM, TGC | 4 | FEP, AMI, TOB, PI + TZ, IMI, DOR, CIP | 7 | 1 (50) | 0 | 1 (50) | 0 | 1 (50) | |
| D | 2 (2.8) | SAM, MEM, TGC | 3 | FEP, AMI, TOB, PI + TZ, IMI, DOR, CIP, CAZ | 8 | 2 (100) | 0 | 0 | 2 (100) | 2 (100) | |
| E | 2 (2.8) | SAM, DOR, MEM, TGC | 4 | FEP, AMI, TOB, PI + TZ, IMI, CAZ, CIP | 7 | 2 (100) | 0 | 0 | 2 (100) | 2 (100) | |
| Non-MDR | F | 13(18.5) | SAM, TGC | 2 | FEP, AMI, TOB, PI + TZ, CAZ, MEM, IMI, DOR, CIP | 9 | 4 (30.7) | 1 (7.6) | 3 (23) | 6 (46.1) | 0 |
| G | 3 (4.2) | SAM | 1 | FEP, AMI, TOB, PI + TZ, CAZ, MEM, IMI, DOR, CIP, TGC | 10 | 1 (33.3) | 0 | 0 | 1 (33.3) | 0 | |
| H | 9 (12.8) | 0 | 0 | SAM, FEP, AMI, TOB, PI + TZ, CAZ, MEM, IMI, DOR, CIP, TGC | 11 | 2 (22.2) | 1 (11.1) | 0 | 1 (11.1) | 1 |
Eight antibiotic resistance patterns were detected (A-H). Five contained XDR and MDR phenotypes (n = 23 isolates, 32.8%) (Table 1). Non-MDR patterns (F-H) were the dominant resistance phenotypes detected among 25 (35.7%) isolates. Also, P. aeruginosa showing pattern A (XDR) expressed resistance against 11 antibiotics (Table 1). The MDR isolates were spotted among pattern B isolates (10 antibiotics) (N = 3, 4.3%), pattern C (four antibiotics) (N = 2, 2.8%), pattern D (three antibiotics) (N = 2, 2.8%), and pattern E (four antibiotics) (N = 2, 2.8%) (Table 1). Twenty-two isolates showed a unique resistance pattern comprising 15 ICU-collected isolates and seven isolates of other wards. The majority of P. aeruginosa infections were observed among ICU patients (n = 46/70, 65.7%), comprising 29 (63%) male and 17 (36.9%) female patients. Moreover, MDR and XDR phenotypes were detected in 29/46 (63%).
4.2. Antiseptic Resistance Gene Distribution
Among 70 isolates, 53 isolates harbored at least one antiseptic resistance gene (75.7%), comprising 11 (20.7%) isolates with the qacE Δ1 gene alone and seven (13.2%) isolates with the qacE gene alone. Simultaneous occurrence of qacE and qacEΔ1 genes were spotted in 35 (66%) isolates. As stated above, grouping the isolates according to their antibiotic resistance phenotypes and resistance genes pattern similarity defined 35 overall combination types among the 70 (50%) strains we studied (Figure 1). The highest prevalence of antiseptic resistance genes (qacE and qacEΔ1) was detected in antibiotic resistance patterns B, D, and E (100%), followed by pattern A (64.2%). The most frequent single occurrence of qacE and qacEΔ1 (50%) genes was in MDR pattern C, followed by XDR pattern A (14.28%). The highest co-occurrence of resistance genes was observed in pattern D (50%) (Table 1).
4.3. Antibiotic Resistance Gene Distribution
The blaOXA-23 gene was spotted in 59 (84.3%) isolates. Most blaOXA-23 gene-positive isolates had at least one antiseptic resistance gene (n = 44/59, 74.5%) (Table 2). The highest occurrence of the blaOXA-23 gene was among patterns A, B, D, and E (100%), following pattern C (50%) (Table 1).
| PCR Result for Antiseptic Resistance Genes | Values | BTC | P b | BKC | P b | CHG | P b | Resistance Gene, Distribution Pattern | BTC | BKC | CHG | blaOXA-23 c |
|---|---|---|---|---|---|---|---|---|---|---|---|---|
| Positive | 53 (75.7) | 24.0 (1.9 - 62.5) | 0.001 d | 46.1 (1.9 - 125) | 0.001 d | 107.7 (31.2 - 250) | 0.001 d | qacE, 7 (13.2) | 15.90 (1.9 - 31.2) | 28.1 (1.9 - 62.5) | 53.3 (31.2 - 62.5) | 4 (6.7) |
| qacEΔ1, 11 (20.7) | 21.4 (1.9 - 62.5) | 27.8 (1.9 - 62.5) | 56.6 (31.2 - 62.5) | 9 (15.4) | ||||||||
| qacEΔ1 + qacE, 35 (66.0) | 26.5 (7.8 - 62.5) | 55.4 (7.8 - 125) | 134.7 (31.2-250) | 31 (52.5) | ||||||||
| Negative | 17 (24.3) | 10.56 (7.8-15.6) | 17.22 (3.9 - 62.5) | 29.4 (15.6 - 31.2) | 10.56 (7.8 - 15.6) | 17.22 (3.9 - 62.5) | 29.4 (15.6 - 31.2) | 15 (25.4) |
Distribution of Antiseptic Resistance Genes and Minimum Inhibitory Concentrations Among A. baumannii Isolates a
4.4. Minimum Inhibitory Concentrations Against Antiseptics
The MICs ranged from 1.9 to 62.5 µg/mL for BTC, 1.9 to 125 µg/mL for BKC, and 31.2 to 250 µg/mL for CHG. The mean MICs for BTC (24.0 versus 10.56 µg/mL, P = 0.001), BKC (46.1 versus 17.22 µg/mL, P = 0.001), and CHG (107.7 versus 29.4 µg/mL, P = 0.001) among antiseptic resistance gene-harboring isolates were statistically significantly higher than those for strains that did not possess resistance genes (Table 2). There was a statistically significant difference in the mean MICs between isolates harboring qacE, qacEΔ1, and qacE + qacEΔ1 for BTC, BKC, and CHG and those having no gene (P = 0.001) (Table 2).
5. Discussion
The current study illustrated the high prevalence of antiseptic resistance genes (qacE and qacEΔ1) and a significant relationship between their presence and increased phenotypic resistance against BTC, BKC, and CHG among infectious P. aeruginosa. The reported rate of MDR and XDR isolates (68.5%) was slightly higher than the average rate for Iran (58%) (23). The accumulation of all the strain-specific variables determined illustrated an extensive diversity among all isolates (35 out of 70 strains, 50%), highlighting that data interpretation had no biases by clonality among isolates; even the isolates from ICU departments showed significant diversity (23 out of 43 strains, 53.4%). Multi-drug resistant and XDR phenotype frequencies were different in China (MDR 18.5%, XDR 3.5%) (24), Pakistan (MDR 36.3%, XDR 18.1%) (25), Iraq (MDR 50%, XDR 45%) (26), Thailand (46.4%) (27), and Nigeria (MDR 61%, XDR 5%) (28). Comprehensive monitoring of antibiotic resistance among P. aeruginosa isolates in 30 European Union countries revealed an MDR rate of zero to 49.4% when among a group of 27 European countries, the reported average rate was under 25% (29).
The prevalence of CRPA isolates was reported higher in Tehran (55.8%) (30) and the Southwest of Iran (52.2%) (31), while the rate of resistance was lower in other parts of Iran, including Yazd (37%) (32) and Golestan provinces (28.1%) (33) than in the present study (48.5%). The CRPA phenotype was detected differently among Asian P. aeruginosa isolates (10.2% to 72.7%) (34-36). The reported rate of CRPA in Egypt was from 42.5% to 100%, depending on hospital localization (37). The rate of CRPA among isolates was significantly lower in European countries (17.2%) and the U.S. (12%) than in the present study (38). Resistance mechanisms in P. aeruginosa are either intrinsic or acquired. Mutations in efflux pumps have been observed in carbapenem-resistant isolates, causing strains to display an MDR phenotype (32). In combination with gene mutations and the acquisition of genetic elements such as qacEΔ1 and qacE genes, efflux pumps play a critical role in this process (39). The rate of qacE∆1-positive P. aeruginosa isolates in Iran (73.7% to 92.5%) (Table 3) was relatively higher than in our current study (20.7%). The situation differed for the qacE gene, which was reported differently from prior Iranian studies (1.1% to 26.3%) (Table 3).
| Place of Study | Genes (Rate in %) | References | |
|---|---|---|---|
| qacE | qacE∆1 | ||
| Iran | |||
| North Khorasan | 13.2 | 20.7 | Current study |
| Ardabil | 26.3 | 73.7 | (40) |
| Hamadan | 1.1 | 36.9 | (41) |
| Qazvin | 17.5 | 92.5 | (42) |
| Tehran and Esfahan | 59 | 91.5 | (43) |
| Other countries | |||
| Australia and India | 100 | 46.2 % | (44) |
| Brazil | - | 48 | (45) |
| Egypt | 13.4 | 47.2 | (46) |
| Egypt | 33 | 78 | (44) |
| Germany | 2.7 | 10 | (22) |
| Saudi Arabia | 18.2 | - | (39) |
| Iraq | - | 97.1 | (47) |
| Bangladesh | 34.72 | 93.05 | (39) |
. Prevalence of Genes Among Pseudomonas aeruginosa Isolates in Iran and Other Countries
The documented rate of the qacEΔ1 gene was lower in Germany (10%) (22) when its reported rate was higher in other countries (Table 3) (39, 44-49). The qacE gene frequency was lower among P. aeruginosa isolates from Germany (2.7%) (22), but several other studies reported a higher rate of qacE genes (33% - 100%) (39, 44, 46, 48, 49) (Table 3). The coexistence of antiseptic resistance genes (qacEΔ1 and qacE) and carbapenem resistance gene (blaOXA-23) among P. aeruginosa isolates here can be explained by the location of genes on the same plasmid (48).
The significant increases in MIC against BTC, BKC, and CHG among P. aeruginosa isolates that harbor qacE and qacEΔ1 genes were already reported in Iran (50) and Egypt (48). However, in studies conducted in Saudi Arabia (39) and Brazil (45), no relation between the presence of qacE and qacEΔ1 genes and increased MIC against biocides was recognized. Bacterial distribution and spread reduce susceptibility among them, affecting biological, socio-economic, and physical aspects (39). The recommended working concentrations for BKC, BTC, and CHG in commercial disinfectants (2000, 1000, and 5000 µg/mL, respectively) (40) are higher than the highest measured MIC in the current study. Still, increased MICs for antiseptic agents active against P. aeruginosa indicate the importance of targeted screening for such P. aeruginosa isolates in hospitals.
5.1. Conclusions
The results of our study highlighted the importance of close monitoring of P. aeruginosa isolates causing infection in hospitals for antiseptic resistance development. This will ultimately help prevent the spread of such organisms.