1. Background
As of November 27, 2022, coronavirus disease (COVID-19), caused by severe acute respiratory syndrome coronavirus 2 (SARS-CoV-2), has resulted in over 645 million confirmed cases and more than 6.6 million deaths (1). Severe acute respiratory syndrome coronavirus 2 infection can lead to a range of clinical outcomes, from asymptomatic carriers to patients with severe acute respiratory disease (ARD) (2-4). Typically, PCR tests turn negative within three weeks for patients recovering from COVID-19; however, there are reports of prolonged positive PCR tests in recovered individuals, persisting for months post-recovery despite non-infectivity (5). While re-infection cases have been documented, cohort studies indicate that some instances of PCR re-positivity months after recovery are not due to re-infection, as these individuals remained in quarantine (6). The phenomenon of prolonged or recurrent viral RNA shedding remains poorly understood due to the novel nature of the disease and warrants further study (6). Studies have detected SARS-CoV-2 RNA in peripheral blood mononuclear cell (PBMC) samples by analyzing sequence data from various studies (7).
Research shows that SARS-CoV-2 is a positive-sense single-stranded RNA virus (8, 9). Recent studies, however, suggest that SARS-CoV-2 DNA may be produced via reverse transcription from the viral RNA genome. This DNA could potentially integrate into the host genome, leading to the production of viral RNA through host-dependent transcription pathways (10). Two mechanisms are proposed: Some viruses, like human immunodeficiency virus (HIV), can encode enzymes such as reverse transcriptase and integrase to synthesize complementary ssDNA and integrate it into the host genome. Another mechanism involves the viral genome recombining with the host genome through components of endogenous transposons like intracisternal A-particle (IAP) and long interspersed element-1 (LINE-1) (11)
Integrating the viral genome into host chromosomal DNA can lead to various consequences, including gene disruption, premature cell death, and oncogene activation, and may contribute to species evolution through inherited genomic inclusions. While integration is a necessary stage for some viruses, such as retroviruses, it may occur incidentally in others (12). Establishing the capability of the SARS-CoV-2 genome to integrate into the host genome opens new avenues for future studies to understand the pathogenic mechanisms of SARS-CoV-2. Such studies could explore the viral integration sites within the host genome, the stage of the viral life cycle at which integration occurs, and the cellular and viral factors involved in this process. They could also aim to develop biomarkers for the persistent presence of the viral genome in recovered COVID-19 cases and determine whether the virus can integrate in all individuals infected with SARS-CoV-2.
2. Objectives
This study aims to investigate the presence of the DNA form of the SARS-CoV-2 genome in oropharyngeal, nasopharyngeal, and PBMC samples from individuals who have recovered from COVID-19 as well as from a healthy control group.
3. Methods
3.1. Study Population
From January 2022 to October 2022, this cross-sectional study enrolled eighty individuals diagnosed with SARS-CoV-2, referred to clinics or hospitals affiliated with Iran University of Medical Sciences (IUMS) in Tehran, Iran. The study consisted of forty outpatient respondents with no specific complications (group 1) and another forty patients who were hospitalized due to significant clinical manifestations and remained molecularly positive for COVID-19 45 days post-onset of the illness (group 2). Additionally, forty healthy individuals served as controls (group 3). Notably, none of the COVID-19 patients or healthy control participants had co-infections with human cytomegalovirus (HCMV), Mycobacterium tuberculosis, hepatitis B virus (HBV), hepatitis C virus (HCV), or HIV.
3.2. Sample Collection and Processing
To assess the presence of the integrated SARS-CoV-2 genome in the participants’ specimens, oropharyngeal and nasopharyngeal samples were collected and stored in viral transport media (VTM). Additionally, 5 mL of peripheral blood was drawn from each participant and placed into sterile vacutainer tubes containing Ethylenediaminetetraacetic acid (EDTA). PBMCs were then isolated from the blood samples using Ficoll-Hypaque (Lympholyte H, Cedarlane, Hornby, Canada) density gradient centrifugation. The resulting PBMC pellet was resuspended in 350 µL of RNALater solution (Ambion, Inc., Austin, TX) and stored at -80°C for subsequent analysis.
3.3. Genomic DNA Isolation
Total DNA was extracted from the oropharyngeal, nasopharyngeal samples, and pellets of 4 - 5 × 106 PBMCs using the QIAamp® DNA Mini kit (Qiagen GmbH, Hilden, Germany), following the manufacturer's protocols. The quality and quantity of the isolated DNA were assessed using a NanoDrop spectrophotometer (Thermo Scientific, Wilmington, MA).
3.4. Amplification of SARS-CoV-2 Genomic DNA Via Real-Time PCR
To determine the presence of the SARS-CoV-2 genome in the samples, cDNA synthesis was omitted, and DNA sequences of the virus were directly tested. Real-time PCR was conducted using specific TaqMan probes and primers targeting the genomic DNA of SARS-CoV-2 in the isolated DNA. The PCR reactions were performed on a Rotor-Gene Q system (QIAGEN, Germany), targeting conserved regions of the N (Nucleocapsid) (13), E (Envelope), and RdRp (RNA-dependent RNA polymerase) (14) genes, with RNase P serving as an internal control (13, 15) (Table 1). The reaction mixture for RT-PCR included 10 pmol of each primer and 5 pmol of each TaqMan probe for the N, E, RdRp, and RNase P genes, 12.5 μL Premix Ex TaqTM (Probe qPCR, TaKaRa Bio Inc., Shiga, Japan), and 5 μL of total DNA as the template. The thermal profile included an initial step at 50°C for 2 minutes, 95°C for 10 minutes, followed by 40 cycles of 95°C for 15 seconds and 60°C for 60 seconds.
| Assay Use and Polarity | Name | Sequences |
|---|---|---|
| RNase P | ||
| Forward primer | RP2-F | AGA TTT GGA CCT GCG AGC G |
| Reverse primer | RP2-R | GAG CGG CTG TCT CCA CAA GT |
| Probe | RP2-probe | ROX- TTC TGA CCT GAA GGC TCT GCG CG -BBQ |
| E | ||
| Forward primer | E-Sarbeco | ACA GGT ACG TTA ATA GTT AAT AGC GT |
| Reverse primer | E-Sarbeco | ATA TTG CAG CAG TAC GCA CAC A |
| Probe | E-Sarbeco | FAM- ACA CTA GCC ATC CTT ACT GCG CTT CG -BBQ |
| N | ||
| Forward primer | N-Pearson | CCA GAA TGG AGA ACG CAG T |
| Reverse primer | N-Pearson | TGA GAG CGG TGA ACC AAG A |
| Probe | N-Pearson | Cy5- GCG ATC AAA ACA ACG TCG GCC CC -BBQ |
| RdRp | ||
| Forward primer | RdRP-SARSr | GTG ARA TGG TCA TGT GTG GCG G |
| Reverse primer | RdRP-SARSr | CAR ATG TTA AAS ACA CTA TTA GCA TA |
| Probe | RdRP-SARSr | VIC- CAG GTG GAA CCT CAT CAG GAG ATG C -BBQ |
Abbreviations: RNase P, ribonuclease P; E, envelope; N, nucleocapsid; RdRp, RNA-dependent RNA polymerase.
3.5. Statistical Analysis
Data were analyzed using SPSS version 20 (SPSS Inc., Chicago, IL, USA). The Kolmogorov-Smirnov test was used to assess data normality. Categorical variables were compared using Fisher's exact test or the chi-square test, as appropriate. A P-value of less than 0.05 was considered statistically significant.
4. Results
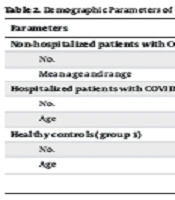
As previously stated, 80 patients with COVID-19 were referred to hospitals associated with IUMS in Tehran, Iran. In addition, 40 healthy individuals (group 3) were recruited for this cross-sectional study. Among the COVID-19 cases, forty were treated on an outpatient basis (group 1) and forty were hospitalized due to the severity of their symptoms (group 2). The mean age of the non-hospitalized COVID-19 patients (group 1) was 36.1 ± 11.0 years (range 22 - 63 years), 61.6 ± 18.4 years (range 13 - 92 years) for hospitalized COVID-19 patients (group 2), and 39.0 ± 8.7 years (range 25 - 51 years) for the healthy participants (group 3). In groups 1 and 2 of patients with COVID-19, and in the healthy controls, 17 (42.5%), 27 (67.5%), and 17 (42.5%) were males, respectively, as detailed in Table 2.
| Parameters | Male | Female | Total | P-Value a |
|---|---|---|---|---|
| Non-hospitalized patients with COVID-19 (group 1) | ||||
| No. (%) | 17 (42.5) | 23 (57.5) | 40 (100.0) | - |
| Age; mean (range) | 37.4 ± 8.3 (29 - 53) | 35.2 ± 12.8 (22 - 63) | 36.1 ± 11.0 (22 - 63) | 0.055 |
| Hospitalized patients with COVID-19 (group 2) | ||||
| No. (%) | 27 (67.5) | 13 (32.5) | 40 (100.0) | - |
| Age; mean (range) | 60.3 ± 19.2 (13 - 85) | 64.5 ± 17.3 (28 - 92) | 61.6 ± 18.4 (13 - 92) | 0.942 |
| Healthy controls (group 3) | ||||
| No. (%) | 17 (42.5) | 23 (57.5) | 40 (100.0) | - |
| Age; mean (range) | 34.4 ± 5.7 (25 - 40) | 42.4 ± 9.1 (25 - 51) | 39.0 ± 8.7 (25 - 51) | < 0.001 b |
a Mann-Whitney U test.
b Statistically significant.
The laboratory and clinical characteristics of the participants, as well as those of the healthy controls, are presented in Tables 3 and 4. The laboratory results for the three groups are summarized in Table 5. After conducting the real-time TaqMan® RT-PCR assay with an internal amplification control to detect the presence of the integrated SARS-CoV-2 genome in the host cell genome, no integration of the virus genome into the host genome was detected. It is important to note that three conserved regions of the virus genome were tested—genes N, E, and RdRp—and all tests returned negative results. Consequently, it can be concluded that the virus genome does not convert into DNA within the host cells, nor does it integrate into the host genome.
| Parameters | Group 11 | Group 22 | Group 33 | P-Value b, c |
|---|---|---|---|---|
| Male/female ratio | 17/23 | 27/13 | 17/23 | 0.036 |
| Fever | 33 (82.5) | 34 (85.0) | 0 (0.0) | < 0.001 |
| Chills | 24 (60.0) | 25 (62.5) | 0 (0.0) | < 0.001 |
| Headache | 24 (60.0) | 27 (67.5) | 0 (0.0) | < 0.001 |
| Weakness | 7 (17.5) | 16 (40.0) | 0 (0.0) | 0.008 |
| Skeletal pain | 26 (65.0) | 29 (72.5) | 0 (0.0) | < 0.001 |
| Chest pain | 15 (37.5) | 19 (47.5) | 0 (0.0) | < 0.001 |
| Shortness of breath | 11 (27.5) | 17 (42.5) | 0 (0.0) | 0.002 |
| Dry cough | 24 (60.0) | 25 (62.5) | 0 (0.0) | < 0.001 |
| Sputum cough | 3 (7.5) | 6 (15.0) | 0 (0.0) | 0.001 |
| Deceased smell | 6 (15.0) | 5 (12.5) | 0 (0.0) | < 0.001 |
| Deceased taste | 9 (22.5) | 10 (25.0) | 0 (0.0) | < 0.001 |
| Runny nose | 24 (60.0) | 6 (15.0) | 0 (0.0) | < 0.001 |
| Cape of nose | 25 (62.5) | 12 (30.0) | 0 (0.0) | < 0.001 |
| Diabetes | 0 (0.0) | 13 (32.5) | 0 (0.0) | < 0.001 |
| Bleeding stomach | 4 (10.0) | 2 (5.0) | 0 (0.0) | 0.025 |
| Gastrointestinal symptom | 22 (55.0) | 8 (20.0) | 0 (0.0) | < 0.001 |
a Values are expressed as No. (%).
b Chi-square test.
c P-values are significant at 0.05.
| Parameters | Group 1 b | Group 2 c | P-Value d, e |
|---|---|---|---|
| Male/female ratio | 17/23 | 27/13 | 0.021 |
| Positive result of PCR for SARS-CoV-2, day | 0.659 | ||
| ≤ 15 | 35 (87.5) | 32 (80.0) | |
| 16 - 30 | 3 (7.5) | 5 (12.5) | |
| 31 - 45 | 2 (5.0) | 3 (7.5) | |
| Fever | 33 (82.5) | 34 (85.0) | < 0.001 |
| Chills | 24 (60.0) | 25 (62.5) | 0.090 |
| Headache | 24 (60.0) | 27 (67.5) | 0.012 |
| Weakness | 7 (17.5) | 16 (40.0) | 0.390 |
| Skeletal pain | 26 (65.0) | 29 (72.5) | 0.022 |
| Chest pain | 15 (37.5) | 19 (47.5) | 0.039 |
| Shortness of breath | 11 (27.5) | 17 (42.5) | 0.500 |
| Dry cough | 24 (60.0) | 25 (62.5) | 0.036 |
| Sputum cough | 3 (7.5) | 6 (15.0) | 0.033 |
| Deceased smell | 6 (15.0) | 5 (12.5) | 0.057 |
| Deceased taste | 9 (22.5) | 10 (25.0) | 0.006 |
| Runny nose | 24 (60.0) | 6 (15.0) | < 0.001 |
| Nasal congestion | 25 (62.5) | 12 (30.0) | < 0.001 |
| Diabetes | 0 (0.0) | 13 (32.5) | < 0.001 |
| Bleeding stomach | 4 (10.0) | 2 (5.0) | 0.259 |
| Gastrointestinal symptoms | 22 (55.0) | 8 (20.0) | < 0.001 |
a Values are expressed as No. (%).
b Non-hospitalized patients with COVID-19.
c Hospitalized patients with COVID-19.
d Fisher’s exact test except for positive result of PCR for SARS-CoV-2 (chi-square test).
e P-values are significant at 0.05.
| Parameters | Group 1 b | Group 2 c | Group 3 d | P-Value e, f |
|---|---|---|---|---|
| Male/female ratio | 17/23 | 27/13 | 17/23 | 0.036 |
| WBC | 7.9 ± 1.0 (5.7 - 9.6) | 7.4 ± 5.3 (2.0 - 32.4) | 7.6 ± 1.4 (4.1 - 9.7) | 0.051 |
| RBC | 4.4 ± 0.4 (3.4 - 5.1) | 4.3 ± 1.1 (1.0 - 7.0) | 4.4 ± 0.4 (3.4 - 5.4) | 0.665 |
| Hb | 13.6 ± 1.2 (11.8 - 15.4) | 12.7 ± 3.6 (1.6 - 20.6) | 13.7 ± 1.4 (10.6 - 16.5) | 0.054 |
| Hct | 41.4 ± 3.8 (35 - 48) | 37.5 ± 9.4 (7.6 - 59.2) | 41.9 ± 4.4 (32 - 49) | 0.001 |
| Platelet | 240 ± 108 (105 - 437) | 184.1 ± 110.1 (20 - 571) | 242.8 ± 113.2 (115 - 465) | 0.004 |
| INR | 1.0 ± 0.1 (0.9 - 1.3) | 1.3 ± 0.7 (1.0 - 5.3) | 1.0 ± 0.1 (0.8 - 1.2) | 0.001 |
| PTT | 30.0 ± 3.2 (26 - 38) | 36.7 ± 14.4 (24 - 84) | 30.1 ± 4.0 (24 - 38) | 0.021 |
| FBS | 86.3 ± 9.3 (77 - 110) | 175.4 ± 121.6 (77 - 512) | 81.8 ± 8.3 (69 - 102) | < 0.001 |
| Urea | 20.5 ± 4.2 (14 - 31) | 25.2 ± 14.3 (7 - 77) | 19.8 ± 4.0 (14 - 29) | 0.186 |
| Cr | 0.93 ± 0.2 (0.5 - 1.2) | 2.3 ± 4.2 (0.6 - 20) | 0.96 ± 0.2 (0.5 - 1.2) | 0.009 |
| AST | 17.6 ± 9.4 (9 - 33) | 56.0 ± 32.0 (19 - 154) | 14.7 ± 5.0 (9 - 24) | < 0.001 |
| ALT | 19.5 ± 10.1 (10 - 39) | 47.0 ± 30.6 (10 - 145) | 16.4 ± 5.2 (10 - 27) | < 0.001 |
| LDH | 262 ± 90.0 (120 - 439) | 658.0 ± 332.0 (121 - 1479) | 222.7 ± 89.4 (109 - 430) | < 0.001 |
| CPK | 61.5 ± 36.3 (24 - 143) | 201.9 ± 503.4 (19 - 3200) | 58.2 ± 31.8 (22 - 140) | 1.000 |
| ALP | 92.4 ± 43.2 (41 - 178) | 301.2 ± 529.5 (45 - 3431) | 91.1 ± 34.3 (40 - 135) | < 0.001 |
| Na | 140 ± 2.8 (136 - 145) | 136 ± 4.6 (115 - 146) | 140 ± 2.7 (134 - 145) | < 0.001 |
| K | 4.0 ± 0.5 (3.3 - 5.0) | 4.1 ± 0.8 (2.0 - 5.7) | 4.0 ± 0.4 (3.3 - 5.4) | 0.126 |
| Ca | 10.0 ± 0.6 (9.0 - 11.2) | 8.7 ± 0.7 (6.5 - 10.2) | 9.9 ± 0.7 (8.9 - 11.0) | < 0.001 |
| Ph | 4.0 ± 0.6 (2.8 - 4.9) | 2.8 ± 0.9 (1.6 - 5.8) | 4.0 ± 0.4 (3.0 - 4.8) | < 0.001 |
| CRP | 7.2 ± 3.2 (2.0 - 12.0) | 26.0 ± 13.0 (4 - 48) | 1.7 ± 0.7 (1.0 - 3.0) | < 0.001 |
| Vitamin D | 23.4 ± 10.5 (11 - 44) | 25.5 ± 12.7 (6 - 47) | 33.3 ± 16.0 (11 - 65) | 0.006 |
Abbreviations: WBC, white blood cell; RBC, red blood cell; Hb, hemoglobin; Hct, hematocrit; FBS, fast blood sugar; PTT, partial thromboplastin time; AST, aspartate aminotransferase; ALT, alanine aminotransferase; CPK, creatine phosphokinase; ALP, alkaline phosphatase; Cr, creatinine; LDH, lactate dehydrogenase; Na, sodium; K, potassium; Ca, calcium; CRP, C-reactive protein; Ph, phosphorus.
a Values are expressed as mean ± SD.
b Non-hospitalized patients with COVID-19.
c Hospitalized patients with COVID-19.
d Healthy controls individuals.
e Mann-Whitney U except for male/female ratio (chi-square test).
f P-values are significant at 0.05.
5. Discussion
When COVID-19 escalated into a pandemic, several reports documented instances where individuals, including recovered and asymptomatic patients, continued to test positive for the virus weeks later, stirring considerable debate (16, 17). These reports raised the possibility of SARS-CoV-2 genome integration into human cellular DNA, akin to what is observed with retroviruses (18). Our study sought to detect the DNA form of SARS-CoV-2 as an indicator of viral integration in the PBMC, oropharyngeal, and nasopharyngeal samples of COVID-19 patients using a TaqMan® RT-PCR assay. Our results did not yield any positive detections in these samples. However, the absence of detected integration does not conclusively disprove our hypothesis about the potential integration of the SARS-CoV-2 genome.
Unlike retroviruses, where genome integration is a crucial part of the viral lifecycle (19), such integration is rare in other viruses. Yet, there are exceptions, such as lymphocytic choriomeningitis virus (LCMV) and bornavirus, which may integrate into the host genome under certain conditions facilitated by host factors like IAP and LINEs, respectively (20-22). Long interspersed element-1, a non-long terminal repeat retrotransposon that constitutes about a fifth of the human genome, includes two open reading frames that encode an endonuclease/reverse transcriptase and a nucleic acid-binding protein (23).
It is hypothesized that SARS-CoV-2 RNA could be reverse transcribed and integrated into the host genome by the endogenous reverse transcriptase protein encoded by LINE-1. Due to this protein's high affinity for RNA, it may bind to viral RNAs and facilitate their retro-integration. Zhang et al. (6) proposed two mechanisms for the integration of SARS-CoV-2, noting that LINE-1 expression is significantly increased in SARS-CoV-2-infected or cytokine-exposed cells. Consequently, the retro-integration of viral RNAs may activate the host's immune system, potentially triggering severe immune responses and cytokine storms (6). In this study, we also investigated the integration of the SARS-CoV-2 genome in long-term hospitalized patients with symptoms and inflammatory conditions, and as mentioned, no positive samples were detected in this group.
Several publications have discussed the integration of the SARS-CoV-2 genome into infected cells under both in vitro conditions and in clinical samples. In these studies, the integrated viral genome was not detected. The negative results might be attributed to increased virus-induced cell death in culture mediums before sample collection and the relatively low and rare occurrence of this phenomenon, which should be investigated with larger sample sizes. In one study, the small sample size could have contributed to the lack of findings (10, 24). Furthermore, given the random nature of the integration process, the likelihood of integration occurring at the same genomic locus across different cases and/or tissues is low (11).
Smits et al. were unable to find any evidence of SARS-CoV-2 integration when investigating with long-read DNA sequencing, aligning with the results of this study (24). It should be noted that the negative results in both the current and previous studies might be influenced by the therapeutic drugs used to treat COVID-19 (6, 25). However, these negative results do not conclusively dismiss the hypothesis of viral genome integration, suggesting that further and more detailed studies are necessary to understand the mechanisms and effects of virus integration into the genomes of infected host cells.
In some viruses, such as human papillomavirus (HPV), only 14 of over 200 HPV types, known as high-risk HPV types, are capable of integrating into the host genome (26, 27). No studies have yet investigated the potential for integration among various SARS-CoV-2 variants. We collected all samples while the delta variant was dominant, yet no positive samples were detected in this study. The small sample size and focus on cases infected with the delta variant are limitations of this study. These factors may impact our findings, so our interpretations should be approached with caution.
5.1. Conclusions
The potential integration of SARS-CoV-2 RNA into host cells remains uncertain, as this study found no evidence of virus-related DNA sequences. Nonetheless, further research is required to explain the phenomenon of long-term PCR positivity in recovered COVID-19 patients.