1. Background
Pseudomonas aeruginosa is a ubiquitous, opportunistic pathogen known for causing a wide range of infections, particularly in immunocompromised individuals (1, 2). Its ability to thrive in various environments, coupled with an inherent resistance to many antibiotics, makes it a formidable challenge in clinical settings (3, 4). The rise of antibiotic resistance in P. aeruginosa has become a significant public health concern, as this pathogen is increasingly implicated in difficult-to-treat infections, particularly in hospital settings (5). According to the INFORM database, P. aeruginosa strains that are multidrug-resistant (MDR) and extensively drug-resistant (XDR) are common and formidable pathogens in healthcare settings, with prevalence rates consistently between 11.5% to 24.7% for MDR strains and 9.0% to 11.2% for XDR strains (6, 7).
The emergence of MDR and XDR strains of P. aeruginosa has further complicated treatment options (8). These strains exhibit resistance to multiple classes of antibiotics, including carbapenems, which are often reserved as last-resort treatments for severe infections (9). The ability of P. aeruginosa to form biofilms, which protect bacterial communities from antimicrobial agents and the host immune response, further exacerbates the challenge of eradicating infections caused by this pathogen (10). Pseudomonas aeruginosa forms biofilms, helping it to survive and dominate in different environments, including the lung environment associated with cystic fibrosis and its association with chronic infections in humans. It can adhere to various surfaces, including medical equipment and contact lenses, and prefers to stick to surfaces in the food industry, like mixing tanks and tubes (4, 11).
Biofilms are structured communities of microorganisms enveloped in a self-generated extracellular matrix, which shields them from environmental threats, including antimicrobial agents. This feature plays a vital role in microbial physiology and has significant public health implications (12-14). The later study underscores the pivotal roles of the biofilm formation genes algD, pelF, and pslD in P. aeruginosa. These genes are responsible for the biosynthesis of three distinct exopolysaccharides: Alginate, Psl, and Pel, which are critical components of the biofilm matrix (15, 16).
2. Objectives
The present study aims to evaluate the prevalence of antimicrobial R, MDR, and XDR in clinical isolates of P. aeruginosa. Additionally, it investigates the relationship between resistance phenotypes and various factors, such as demographic characteristics, specimen types, and disease conditions. The study also examines the correlation between the presence of the algD, pelF, and pslD genes and the biofilm formation capacity of clinical isolates of P. aeruginosa. This study contributes to supporting the development of effective strategies for managing and treating infections caused by highly R strains of P. aeruginosa.
3. Methods
3.1. Pseudomonas aeruginosa Clinical Isolates
Twenty-five laboratory-confirmed P. aeruginosa clinical strains were obtained from the Microbiology Department of King Fahad Medical City (KFMC), Riyadh, KSA. The strains were isolated from patients’ clinical samples, and phenotypic identification of P. aeruginosa involved morphological, cultural, and biochemical tests. The strains were designated as PA1 - PA25. All identified strains were preserved in tryptic soy broth with 20% glycerol and stored at -20°C. For the experiment, the strains were cultured in tryptic soy broth (TSB; Oxoid) at 37°C to ensure optimal growth conditions.
3.2. Antibiotic Susceptibility
The antimicrobial susceptibility test was performed using the Kirby-Bauer disk diffusion method on 25 P. aeruginosa clinical strains. Strains were cultured in Mueller-Hinton broth (MHB) for 4 - 6 hours and adjusted to a 0.5 McFarland standard turbidity. The suspension was spread on Mueller-Hinton agar (MHA) plates to form a bacterial lawn, followed by gentle air drying. Antimicrobial disks used included piperacillin-tazobactam (PIT), cefepime (FEP), ceftazidime (CAZ), gentamicin (GEN), ciprofloxacin (CIP), amikacin (AMK), imipenem (IPM), meropenem (MEM), colistin (COL), and levofloxacin (LEV). Clear zone diameters were measured and interpreted as sensitive or resistant as per CLSI guidelines, using ATCC strains as references (17, 18). Isolates were classified based on susceptibility: Extensively drug-resistant if non-susceptible to ≥ 1 agent in all but ≤ 2 antimicrobial categories, MDR if non-susceptible to ≥ 1 agent in ≥ 3 categories, and resistant if non-susceptible to one or two antibiotics (19).
3.3. Quantification Biofilm Assay
The biofilm quantification was performed using a previously described microtiter biofilm formation assay with minor modifications. Pseudomonas aeruginosa isolates were cultured in TSB broth for 24 hours at 37°C. Pseudomonas aeruginosa (PA01) served as a positive control. For microtitration, 200 μL of bacterial suspensions were added in triplicate to 96-well polystyrene plates and incubated at 37°C for 24 hours. After incubation, suspensions were removed, and wells were washed three times with PBS (pH 4.0). Wells were stained with 200 μL of 0.1% crystal violet (CV) solution for 30 minutes, washed with PBS, and treated with 200 μL of 30% glacial acetic acid for 15 minutes at room temperature. Then, 100 μL of the solubilized CV was transferred to a flat-bottom microtiter dish, and absorbance was read at 595 nm using an ELISA reader. Strains were classified based on OD values (OD < 0.071, non-biofilm; OD 0.071 - 0.142, weak; OD 0.142 - 0.284, moderate; OD > 0.284, high).
3.4. Bacterial DNA Extraction
The DNA extraction from P. aeruginosa clinical strains was performed according to the manufacturer’s instructions of the genomic DNA purification kit (K0512, Thermo Scientific). The DNA isolated from P. aeruginosa clinical strains was subjected to polymerase chain reaction (PCR) to detect the biofilm-associated genes using specific primers listed in Table 1.
3.5. Polymerase Chain Reaction
The PCR assay was performed according to the instructions provided in the kit (PCR Master Mix 2X- K0171, Thermo Scientific). Amplification was carried out in a PCR thermo cycler (AB Applied Biosystems, GeneAmp 9700) with the optimized PCR program that consisted of an initial denaturation at 95°C for 3 minutes, followed by 40 cycles (21). Each cycle comprised denaturation at 95°C for 30 seconds, annealing temperature (56/58°C) (22) for 30 seconds, and extension at 72°C for 1 minute. The final extension included one cycle at 72°C for 15 minutes. The known DNA sample containing the target genes (algD, pelF, and pslD) was used as a positive control, and the reaction without the DNA sample was used as a negative control. The amplicons were resolved by electrophoresis in 1.2% agarose gel, stained with SYBR Safe DNA gel stain, and visualized in a gel documentation system (Bio-Rad, USA).
3.6. Statistical Analysis
The demographic and resistant phenotypes were analyzed by contingency table analyses and Fisher’s exact test to compare resistant phenotypes with demographic data, biofilm formation capacity, and presence of biofilm genes. The results were considered significant if the P-value was ≤ 0.05. Simple frequencies (n) and percentages (%) were calculated for each variable. All the statistical analyses were performed using GraphPad Prism 10.2.3 software.
4. Results
This study evaluated 25 clinical isolates of P. aeruginosa to determine their resistance, MDR, and XDR phenotypes. The isolates were analyzed in relation to patient age, gender, specimen type, disease conditions, the presence of biofilm-forming genes, and their biofilm formation capacity.
4.1. Antibiotic Resistance
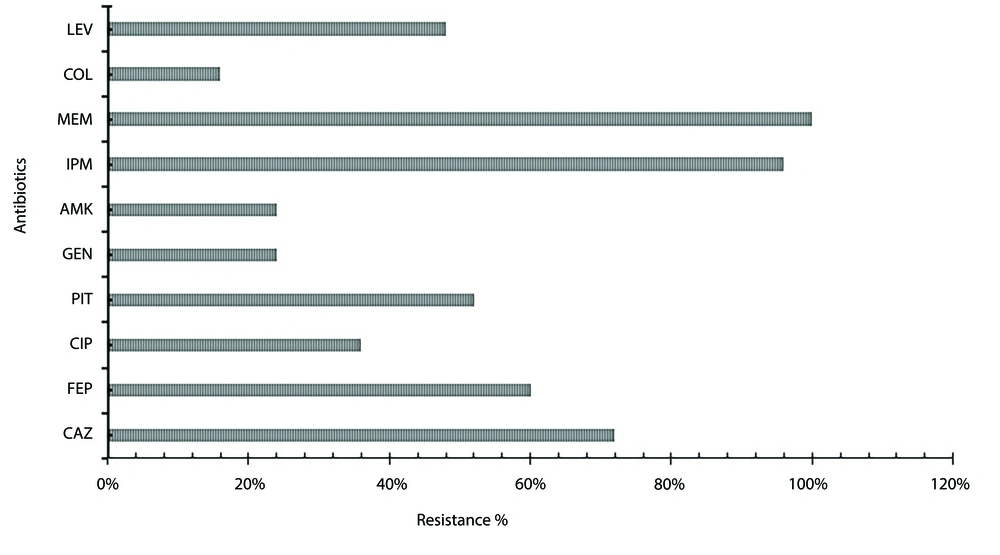
The highest resistance was observed against MEM (100%), IPM (96%), and CAZ (72%), whereas the least resistance was observed in COL (16%) (Figure 1).
Antibiotic resistance percentage of Pseudomonas aeruginosa clinical isolates. Figure 1 represents the clinical isolates of P. aeruginosa, which showed the highest resistance with antibiotic meropenem (MEM) and imipenem (IPM) (96 - 100%), moderate resistance with antibiotics ceftazidime (CAZ), cefepime (FEP), piperacillin-tazobactam (PIT), and levofloxacin (LEV) (72 - 48%) and the least resistance were observed with antibiotics gentamicin (GEN), amikacin (AMK) and colistin (COL) (24 - 16%).
4.2. Demographics
The highest percentage of R strains was detected in the 51 - 60 and 71 - 80 age groups (12%), while in the 81 - 90 age group, MDR (16%) strains were isolated. In males, the highest percentage of R strains (28%) was detected, and in females, MDR (24%) strains were detected. The difference between genders was not statistically significant (Table 2).
| Variables | Strains (n = 25) | |||
|---|---|---|---|---|
| R | MDR | XDR | P-Value | |
| Age | 0.125 | |||
| 0 - 10 | 1 (4) | 0 (0) | 1 (4) | |
| 11 - 20 | 0 (0) | 0 (0) | 0 (0) | |
| 21 - 30 | 0 (0) | 2 (8) | 1 (4) | |
| 31 - 40 | 0 (0) | 0 (0) | 0 (0) | |
| 41 - 50 | 0 (0) | 0 (0) | 1 (4) | |
| 51 - 60 | 3 (12) | 0 (0) | 1 (4) | |
| 61 - 70 | 2 (8) | 3 (12) | 0 (0) | |
| 71 - 80 | 3 (12) | 1 (4) | 1 (4) | |
| 81 - 90 | 1 (4) | 4 (16) | 0 (0) | |
| Gender | 0.403 | |||
| Male | 7 (28) | 4 (16) | 2 (8) | |
| Female | 3 (12) | 6 (24) | 3 (12) | |
| Type of specimen | 0.679 | |||
| Sputum | 6 (24) | 6 (24) | 3 (12) | |
| Wound | 3 (12) | 4 (16) | 1 (4) | |
| Abdominal wall | 1 (4) | 0 (0) | 0 (0) | |
| Ear swab | 0 (0) | 0 (0) | 1 (4) | |
| Disease condition | 0.592 | |||
| Diabetes mellitus | 5 (20) | 2 (8) | 1 (4) | |
| Cardiopulmonary arrest | 1 (4) | 0 (0) | 0 (0) | |
| Encephalitis | 0 (0) | 1 (4) | 0 (0) | |
| Immunodeficiency | 0 (0) | 0 (0) | 1 (4) | |
| Ovarian mass | 0 (0) | 1 (4) | 0 (0) | |
| Pneumothorax | 0 (0) | 1 (4) | 0 (0) | |
| Sepsis | 4 (16) | 4 (16) | 2 (8) | |
| Traumatic brain injury | 0 (0) | 1 (4) | 1 (4) | |
| Biofilm formation genes positive | 0.721 | |||
| algD | 9 (36) | 8 (32) | 5 (20) | |
| pelF | 4 (16) | 10 (40) | 3 (12) | |
| pslD | 9 (36) | 10 (40) | 5 (20) | |
| Biofilm formation strains | 0.502 | |||
| Strong biofilm formation | 3 (12) | 5 (20) | 3 (12) | |
| Moderate biofilm formation | 7 (28) | 4 (16) | 2 (8) | |
| Weak biofilm formation | 0 (0) | 1 (4) | 0 (0) | |
Abbreviations: R, resistant; MRD, multidrug resistance; XDR, extensively drug resistance.
4.3. Specimen and Disease Condition
From the sputum samples, the highest resistance (24%), MDR (24%), and XDR (12%) strains were isolated, whereas in sepsis cases, the highest resistance (16%), MDR (16%), and XDR (8%) strains were detected. Differences by specimen type and disease condition were not statistically significant (Table 2).
4.4. Biofilm Formation Genes
The algD gene was found to be most prevalent in R (36%) strains, while pelF and pslD were highest in MDR strains (40%). No significant differences were observed between the presence of genes and resistance phenotypes (Tables 2 and 3 and Figure 2) (Appendices 1 - 3 in Supplementary File).
| Pseudomonas aeruginosa Strains | algD | pelF | pslD | Resistant Phenotype |
|---|---|---|---|---|
| PA1 | + | + | + | MDR |
| PA2 | + | + | + | R |
| PA3 | - | + | + | MDR |
| PA4 | + | + | + | XDR |
| PA5 | + | - | + | R |
| PA6 | + | - | + | XDR |
| PA7 | + | - | + | XDR |
| PA8 | + | - | + | R |
| PA9 | - | - | + | R |
| PA10 | + | + | + | R |
| PA11 | + | + | + | MDR |
| PA12 | + | + | + | MDR |
| PA13 | + | + | + | MDR |
| PA14 | + | + | + | MDR |
| PA15 | + | + | + | MDR |
| PA16 | + | + | + | R |
| PA17 | + | - | + | R |
| PA18 | + | + | + | XDR |
| PA19 | + | + | + | XDR |
| PA20 | + | + | + | MDR |
| PA21 | + | - | + | R |
| PA22 | - | + | + | MDR |
| PA23 | + | - | + | R |
| PA24 | + | + | + | MDR |
| PA25 | + | + | - | R |
| PAO1 | + | - | + | R |
Abbreviations: R, resistant; MRD, multidrug resistance; XDR, extensively drug resistance.
a Correlation analysis of algD in comparison with resistance phenotype r = 0.065, P ≤ 0.755; correlation analysis of pelF in comparison with resistance phenotype r = 0.275, P ≤ 0.183; correlation analysis in comparison with resistance phenotype r = 0.218; P ≤ 0.295.
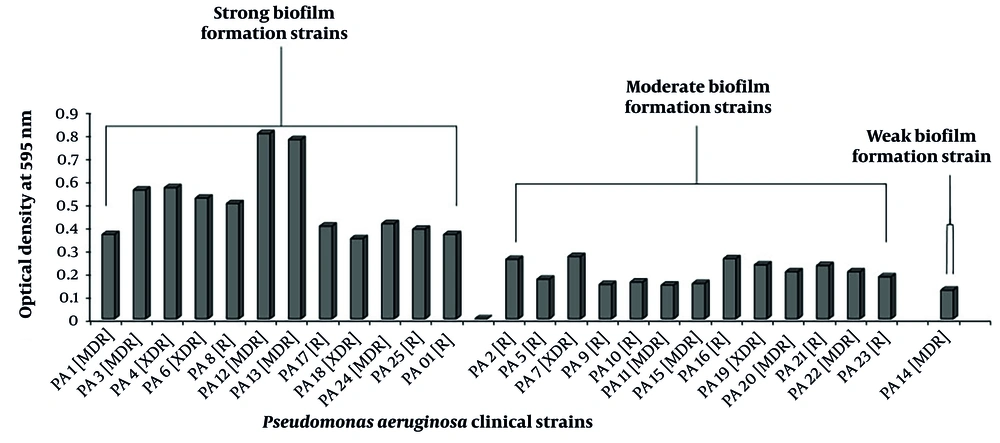
Evaluation of biofilm formation capacity in Pseudomonas aeruginosa clinical isolates. R, resistant; MDR, multidrug resistance; XDR, extensively drug resistance. Correlation analysis in comparison with resistance phenotype r = 0.188, P ≤ 0.366 (OD ≤ 0.071 non-biofilm; OD ≤ 0.071 - 0.142 wk; OD ≤ 0.142 - 0.284 moderate; OD ≤ 0.284 high).
4.5. Biofilm Formation Strength
Strong biofilm formation was most common in MDR (20%) and XDR (12%) strains. Differences in biofilm formation capacity between resistance phenotypes were not statistically significant (Table 2 and Figure 2).
5. Discussion
The findings of this study underscore the significant prevalence of antibiotic resistance among P. aeruginosa clinical isolates, with a particular emphasis on carbapenem resistance. Alarmingly, 100% of isolates were resistant to MEM, and 96% were resistant to IPM, highlighting the critical threat posed by carbapenem-resistant Pseudomonas aeruginosa (CRPA). These results reflect a persistent global health concern, as CRPA infections are associated with limited treatment options and poor clinical outcomes. The high rates of resistance observed in this study emphasize the urgent need for robust antimicrobial stewardship programs and the development of novel therapeutic approaches to combat these MDR pathogens.
The demographic analysis revealed higher rates of R strains in older age groups, particularly in the 51 - 60 and 71 - 80 age ranges. This trend may be attributed to the increased susceptibility of older patients to infections due to weakened immune systems and prolonged healthcare exposure to antibiotic usage (20). The highest percentage of MDR strains was observed in the 81 - 90 age group, a population already at heightened risk for severe infection-related complications. Gender-based analysis showed a higher prevalence of R strains in males and MDR strains in females, though the differences were not statistically significant. Interestingly, prior studies have also reported a higher prevalence of MDR strains in females, potentially linked to the higher incidence of urinary tract infections (UTIs) in the female population (23). Such trends may reflect differences in infection types, healthcare exposure, or antibiotic usage patterns, warranting further investigation.
In sputum samples, the highest percentage of resistance (24%), MDR (24%), and XDR (12%) strains were isolated, which aligns with the known propensity of P. aeruginosa to cause respiratory infections, particularly common in patients with underlying lung conditions, such as cystic fibrosis or chronic obstructive pulmonary disease (COPD) (24, 25). Similarly, in sepsis disease conditions, the highest percentage of resistance (16%), MDR (16%), and XDR (8%) strains were isolated, underscoring the severe clinical implications of P. aeruginosa infections in critically ill patients. These findings align with previous reports that MDR P. aeruginosa increases the risk of adverse outcomes in sepsis, including treatment failure and mortality. Although differences in resistance patterns across specimen types and disease conditions were not statistically significant, the observed trends are consistent with the clinical behavior of this P. aeruginosa opportunistic pathogen (26-28).
Biofilm formation is a well-known factor contributing to antibiotic resistance, as it can protect bacterial communities from antimicrobial agents and the host immune system (29). The presence of biofilm-formation genes, particularly algD, pelF, and pslD, highlights their potential role in the resistance mechanisms of P. aeruginosa. In this study, the algD gene was most commonly detected in R strains (36%), suggesting its involvement in strong biofilm formation, which contributes to the ability of P. aeruginosa to evade antimicrobial agents and host defenses. In contrast, pelF and pslD genes were most prevalent in MDR strains (40%), reflecting their role in the development of a robust biofilm matrix, which enhances bacterial persistence and resistance (30, 31). Although no statistically significant differences in the presence of these genes were observed among resistance phenotypes, the trends align with the established understanding of biofilm formation as a critical factor in antimicrobial resistance. The lack of statistical significance may be attributed to the small sample size or the complexity of resistance mechanisms, which likely involve a combination of genetic and environmental factors. Further research with larger sample sizes and a broader scope of genetic analysis is needed to clarify the contribution of these genes to resistance and to explore additional factors that may influence biofilm-formation resistance in P. aeruginosa.
The high prevalence of carbapenem resistance, coupled with the significant biofilm formation capacity of MDR and XDR strains, represents a formidable challenge for infection control and treatment. These findings underscore the critical need for integrated strategies, including antimicrobial stewardship, biofilm-targeting therapies, and ongoing surveillance of resistance trends. Further studies exploring additional molecular mechanisms and resistance determinants may provide deeper insights into the complex interplay of factors involved in the resistance of P. aeruginosa.
5.1. Conclusions
To conclude, these findings emphasize the role of biofilm-formation genes (algD, pelF, and pslD) in resistance mechanisms, suggesting their involvement in enhancing bacterial persistence and antimicrobial evasion. Although the lack of statistically significant differences in gene distribution across resistance categories may reflect the study’s limited sample size, the trends observed align with the well-established role of biofilm formation in promoting bacterial resistance. Strong biofilm formation, particularly in MDR and XDR strains, highlights the potential for biofilm-targeted therapies to complement traditional antimicrobial strategies.