1. Background
Leishmaniasis is an important public health problem all over the world. Leishmaniasis, malaria, schistosomiasis, filariasis, trypanosomiasis and tuberculosis, are considered by the World Health Organization (WHO) to be six of the most important tropical diseases (1). Leishmaniasis has been reported in 88 countries around the world and the prevalence of the disease is estimated to be approximately 12 million annually, in addition, approximately 350 million people are at risk of catching the disease (1, 2).
Visceral leishmaniasis (Kala-azar) is characterized by the presence of; fever, splenomegaly, hepatomegaly, swollen lymph nodes and weight loss, depending on the pathogenicity of the Leishmania species and the host’s immune response against the parasite (3, 4). This disease can lead to death in 90% of cases, if the patient is left without any treatment. The drugs which are currently used as treatment for leishmaniasis cannot be handled easily due to a number of problems including; high toxicity, and various side effects (4). Therefore, efforts to introduce a new protein for vaccine production are currently being considered. Up to now, various antigens such as TSA, and P4 LACK, have been evaluated for vaccine development (5-9). In this regard, a new molecule common in Leishmania species, is the 11KD protein of the membrane kinetoplastid surface or KMP-11 (10). The KMP-11 is highly conserved and has common characteristics in most of the Kinetoplastidae family (11). In the beginning, these molecules were called associated proteins of lipophosphogly can and then called kinetoplastid membrane protein-11 (12, 13). KMP-11 was isolated from Leishmania donovani and previously characterized by Jardim et al. in 1995 (12). Some findings suggest that the KMP-11 protein may be involved with mobility in both the parasite and in binding to the host cell. So, it could be considered as a candidate for vaccine production (14).
2. Objectives
The aim of this present study was to increase our knowledge concerning molecular cloning and expression of the KMP-11 gene of L. infantum. In the present study the KMP-11 gene of L. infantum encoding KMP-11proteinswas cloned into a pcDNA3 expression vector, and transfected into Chinese hamster ovary (CHO) eukaryote cells. The main objective of the study was to provide an introduction for DNA vaccine production against visceral leishmaniasis.
3. Material and Methods
3.1. Parasite
Promastigotes of L. infantum (MHOM/TN/80/IPI1) were prepared from the Pasteur Institute of Iran (Tehran, Iran). Promastigotes were cultured in a RPMI1640 medium and supplemented with 10% FCS (Fetal Calf Serum, Sigma-Aldrich, Germany). This culture was then incubated at 24±1˚ C.
3.2. Genomic DNA Extraction
Genomic DNA was extracted from the promastigotes of Leishmania by a DNA genomic extraction kit (Bioneer, Hamburg, Germany), this process was carried out according to the manufacturer’s protocol. DNA concentration and quality was assessed by both UV absorbance and electrophoresis on the 1% agarose gel.
3.3. DNA Amplification
The PCR reaction was performed in a 25µl volume containing; 5 µl of DNA genomic as a template, 1µl of each primer, 2µl of a PCR master mix, and 16µl of distilled water. Sense and antisense oligonucleotide primers were designed based on DNA sequences of KMP-11 that were recorded in GenBank. Forward primer: AAGCTTATGGCCACCACGTACGAGGAG and reverse primer:GAATTCTTACTTGGATGGGTACTGCGCAGC. The forward primer contains a HindIII restriction site and the reverse primer contains an EcoRI restriction site 6 nucleotides. The PCR procedure included: 94ºC for two minutes as initial denaturation, 35 cycles at 94º C for 30 seconds, 59º C for 30 seconds, 72º C for 30 seconds and then 72º C for seven minutes as a final extension. Finally, the PCR products were analyzed by electrophoresis on a 2% agarose gel. KMP-11 gene fragments were purified from the gel using a gel purification kit (Bioneer, Hamburg, Germany).
3.4. Ligation and Transformation
Ligation of the KMP-11 gene into the pTZ57R plasmid was performed using an InsTA cloneTM PCR product cloning kit (Fermentas®, Germany). Ligation reaction was prepared in a 30µl volume containing; 5µl of template (DNA extraction product), 3µl of pTZ57R plasmid, 5µl of 5x ligation buffer, 1µl of T4 ligase and15µl of distilled water. This reaction was first incubated at 22˚C for an hour and then at 4˚C for 16 hours. The competent cells were prepared from the TOP10 strain of Escherichia coli bacteria using the calcium chloride method. E. coli 100µl, was cultured with 5ml of new Luria-Bertani and then incubated at 37˚C for 14hours. The next day, 200µl of these cells were cultured in 10ml of LB broth and incubated at 37˚C for three hours. After incubation, the culture was centrifuged at 7000 rpm and 4˚C for 10 minutes.
The supernatant was discharged and then 3-4ml of cold calcium chloride (100mM) was added to the pellet and mixed gently. The suspension was centrifuged at 7000rpm and 4˚C for 10 minutes. The supernatant was discharged and 250µl of cold calcium chloride added to the pellet and mixed gently before being incubated on ice pieces for one hour. Following this, 10µl of the ligation product was added to this suspension and incubated on ice pieces for 30 minutes. Then, the suspension for heat shock was incubated at 42˚C for 90 seconds and then immediately transferred onto ice. Subsequently 500µl of LB broth, without any antibiotic, was added and incubated at 37˚C for one hour. These cells were cultured on plates of Luria-Bertani (LB) agar medium containing; 100mg/mL of ampicillin, 200mg/mL of IPTG and 20mg/mL of X-Gal. The plates were incubated at 37˚C for 16 hours.
Recombinant plasmids (pTKMP-11) were extracted from white colonies according to the protocol for the plasmid extraction kit (Bioneer, Hamburg, Germany). The recombinant plasmid (pTKMP11), pTZ57R (negative control) and pcDNA3(expression vector) were digested by EcoRI and HindIII enzymes. The double digestion reactions were prepared in the same way for each in 220µl volumes containing 80µl of the plasmid extraction product, 88µl of water, 44µl of tango buffer (Fermentas®, Germany), 4µl of EcoRΙ and 4µl of HindΙΙΙ enzymes. These reactions were incubated at 37° C for 16 hours. The products of digestion were analyzed by electrophoresis on 0.8% agarose gel. The bands belonging to the KMP-11 fragment and digested expression plasmids (pcDNA3) were purified from the agarose gel by a gel purification kit (Bioneer, Hamburg, Germany). KMP-11fragments were purified from the gel and sent to the GenFanaVaran®Company (Iran, Tehran) for sequencing.
3.5. Subcloning in pcDNA3 Expression Vector
The KMP-11genewas ligated into digested pcDNA3 expression vectors using T4DNA ligase enzymes. Recombinant plasmid pcKMP-11 was transformed into competent E.coli cells and recovered in LB broth medium free of antibiotic at 37˚C for one to three hours. Then, it was sub-cultured on new LB agar medium and incubated at 37˚C for 16 hours. Recombinant plasmids (pcKMP-11) were extracted from E. coli by a Bioneer plasmid extraction kit according to the manufacturer’s instructions. The extracted plasmids from the white colony were digested by EcoRI and HindIII enzymes and electrophoresis occurred on 0.8% agarose gel.
3.6. Expression of Recombinant Protein
In order to carry out transfecting, CHO eukaryote cells were used as host cells for recombinant expression plasmids (pTKMP-11). The CHO cells were cultured under sterile conditions in DMEM (Gibco) medium supplemented with 5% FCS under 37ºC with 5% CO2 conditions.
3.7. RNA Extraction and Reverse Transcriptase Procedure
For gene transcription confirmation at the level of mRNA, a RT-PCR kitwas used. The total RNA isolated from the cells by RNXTM-plus Isolation RNA kit (Fermentas, Germany). RTPCR was performed according to the manufacturer’s protocol using a RT PCR kit (Fermentas, Germany) then the result was electrophoresis on 2% agarose gel.
4. Results
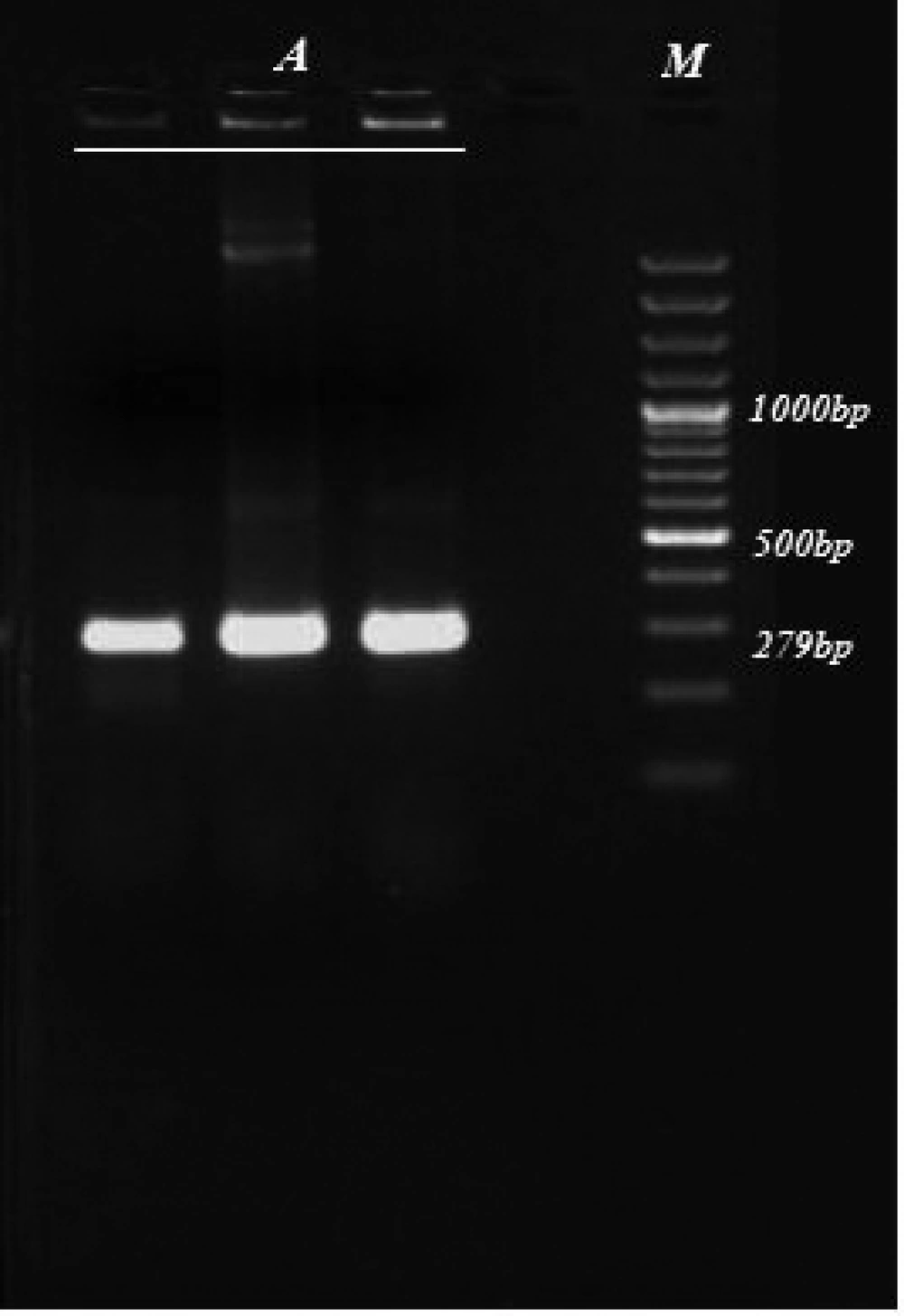
The size of the amplified KMP-11gene of L. infantum in our study was similar to earlier predictions. A 279 bp DNA fragment was identified following agarose gel electrophoresis of the PCR products ( Figure 1 ). The PCR products were ligated into a pTZ57R/T standard cloning vector. The plasmid (pTKMP-11) was transformed successfully into E. coli. The recombinant plasmids from the white colony were sequenced and compared with sequences in the GenBank.The KMP-11gene sequence was 97-99% identical with that of other organisms ( Table 1 and Table 2 ).
a Accession number JF422108.1 were compared with 16 sequences of other organisms submitted to GenBank
| Protein ID | Amino Acids | Country | |
|---|---|---|---|
| L. infantum | AEK80413.1 | 92 | Iran |
| L. infantum | XP_001469032 | 92 | England |
| L. infantum | CAA64883 | 92 | Spain |
| L. major | XP_843328 | 92 | USA |
| L. donovani | AAB33127 | 92 | England |
| L. donovani | S53442 | 92 | Canada |
| L. tropica | CAA03902 | 92 | Spain |
| L. panamensis | AAC61837 | 92 | Columbia |
| L. amasonensis | AAG32958 | 92 | Brazil |
| L. guyanensis | AAB94115 | 92 | Columbia |
| Tryponosoma rangeli | ABA42053 | 92 | Columbia |
| Crithidiasp | ABA42050 | 92 | Columbia |
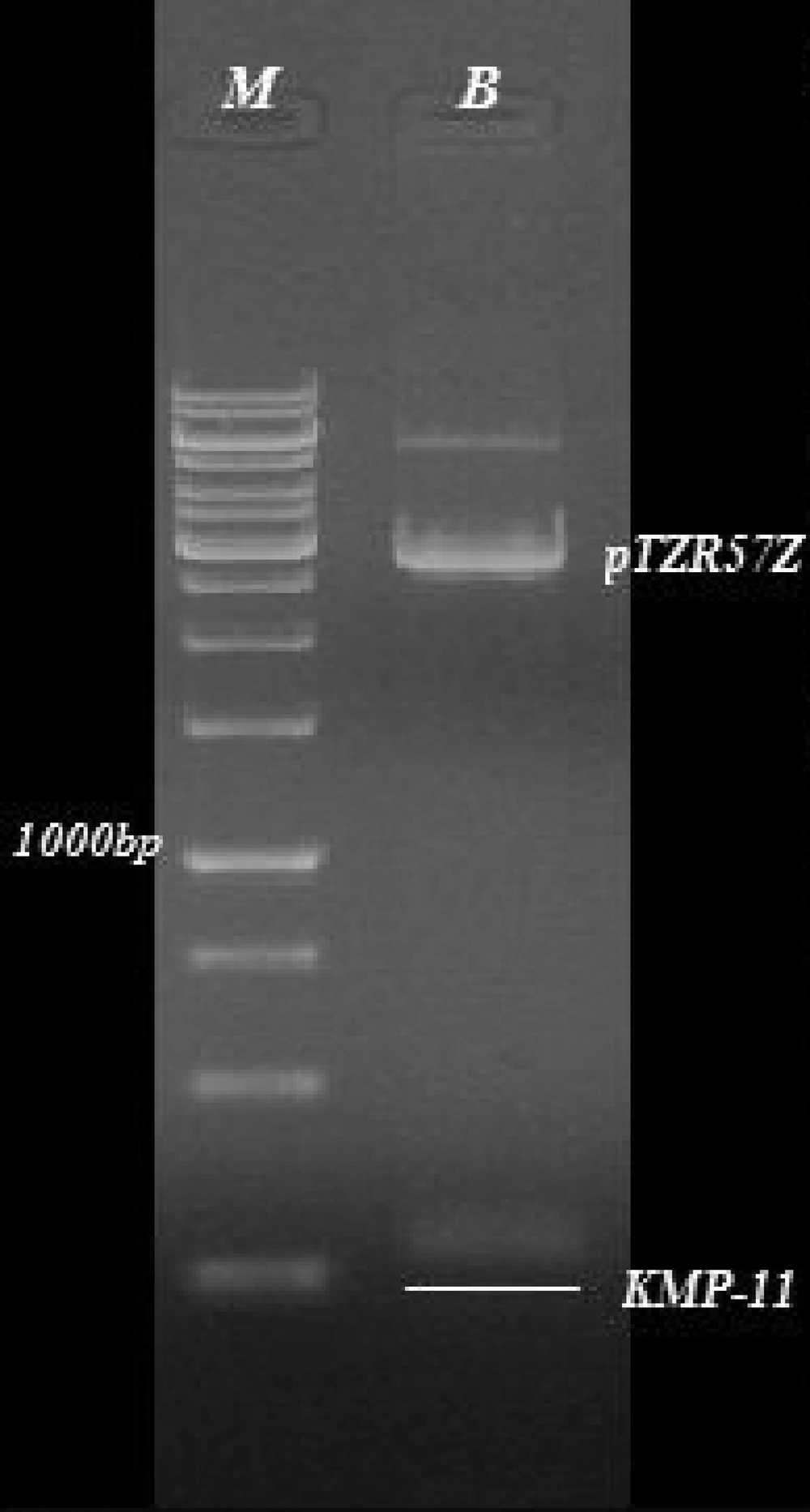
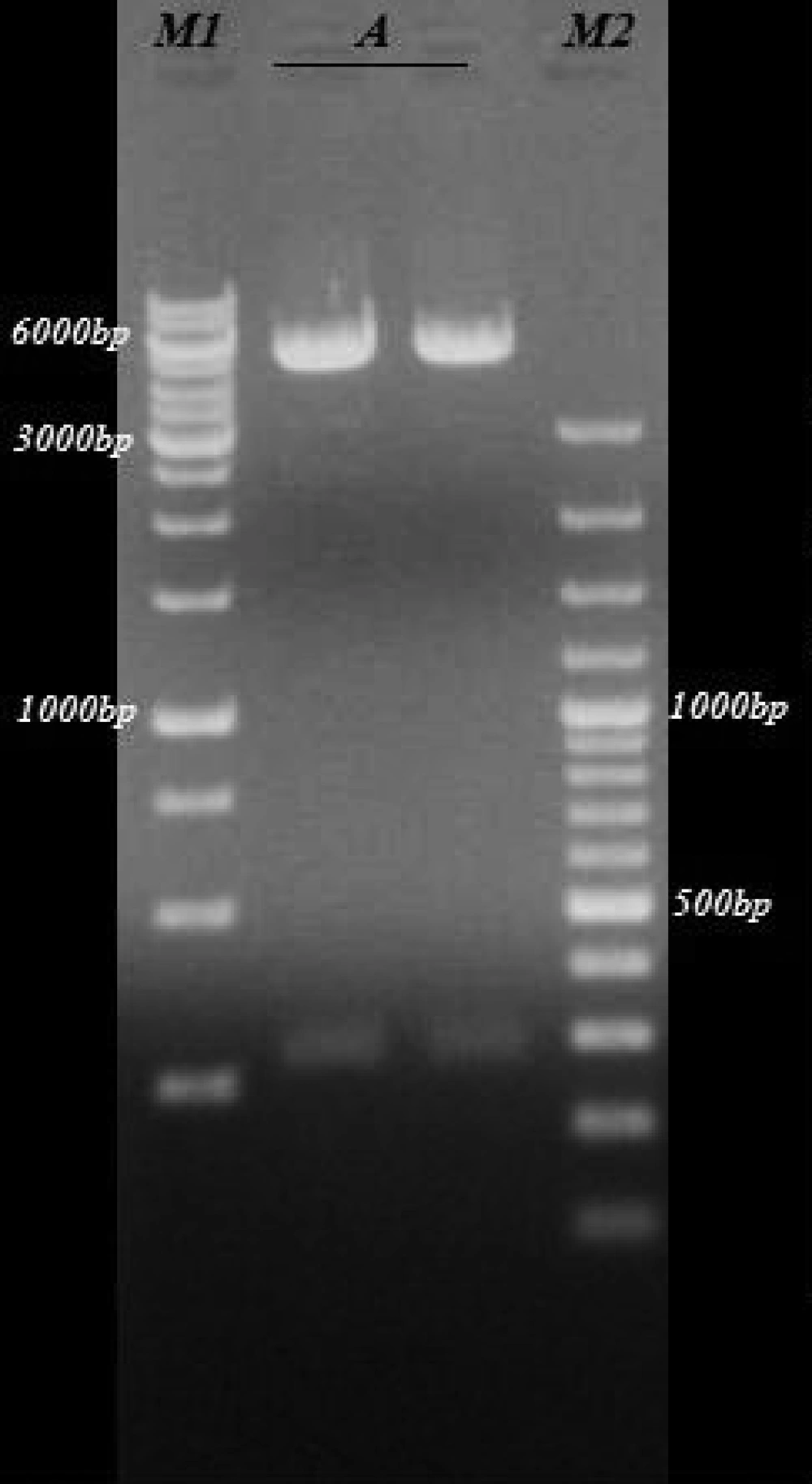
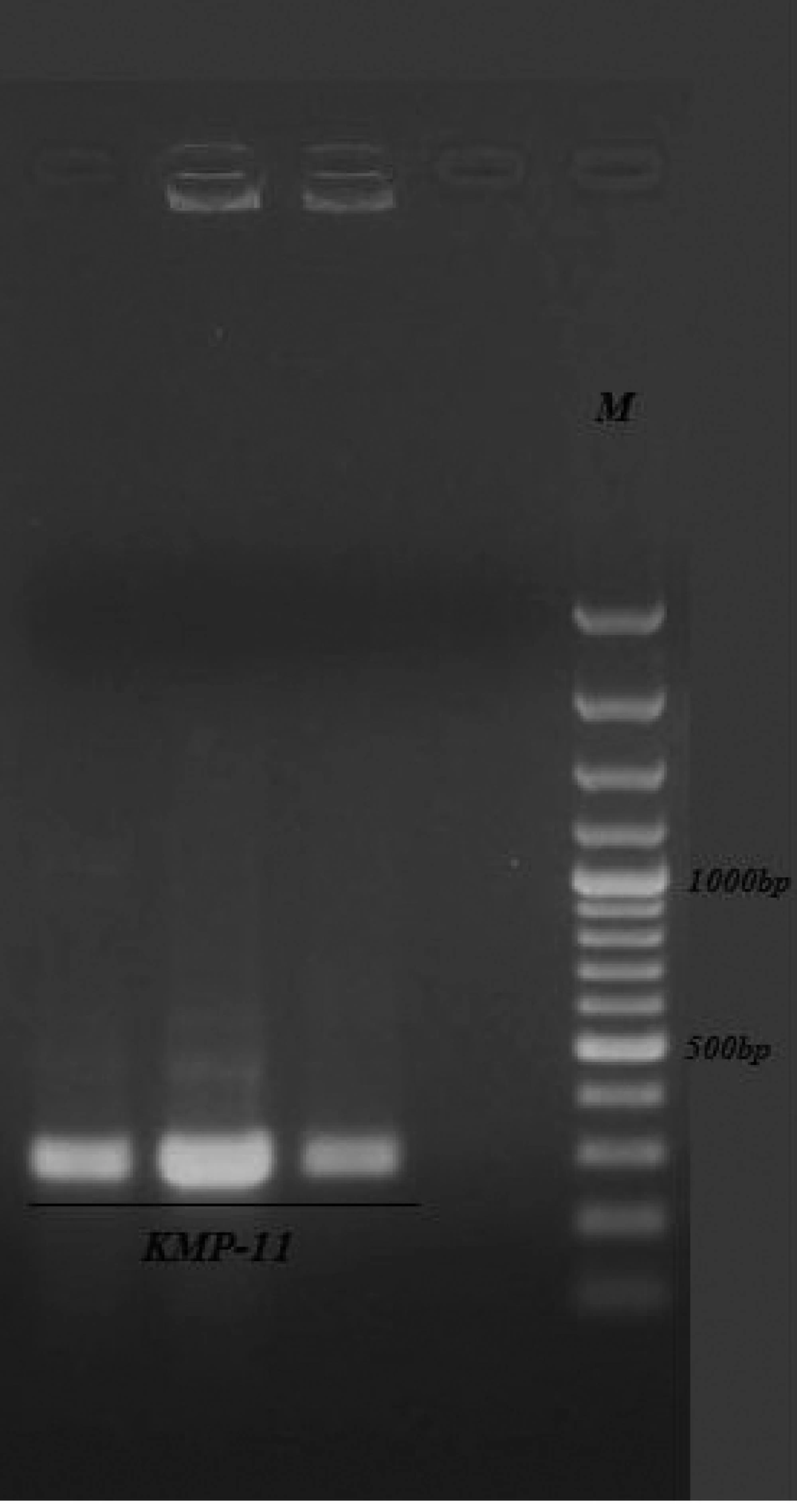
The sequence of the KMP-11gene from L. infantum in our study was submitted to GenBank with the accession number: JF422108.1. The extracted plasmids from the white colony were digested by EcoRI and HindIII enzymes and delectrophoresis on 0.8% agarose gel ( Figure 2 ). At the next step, this gene was successfully cloned into pcDNA3 expressing eukaryote vectors and transformed into E. coli bacteria. The extracted expression plasmids digested by EcoRI and HindIII enzymes are shown in Figure 3 . The expression plasmids pTKMP-11, were transfected successfully into the CHO cells and following RNA extraction, they were confirmed by RTPCR methods, finally the results were electrophoresis on 2% agarose gel ( Figure 4 ).
5. Discussion
To date, various antigens such as TSA, LACK and P4 have been evaluated to assess their potential for DNA or recombinant vaccine development against leishmaniasis (5-9). In addition, the KMP-11protein has been proposed as a candidate for vaccine production against L. infantum (14).The sequence of the L. infantumKMP-11 gene in the present study (accession no: JF422108.1), was 98-99% identical with other Leishmania species. Translation from the sequence of this gene product produces a protein containing 92 amino acids and it has an isoelectric point of 4.8, which is associated with the membrane of the promastigote.
According to the Berberich et al. study (1998), the transcription of the KMP-11 gene results invariable sized mRNA with 0.5kb in Trypanosoma cruzi, 0.8 kb and 1.3 kb in T. brucei and L. infantum, respectively (10). Proteins in most of the kinetoplastids have common characteristics such as weight;11KD molecular and expression at all stages of the parasite's life cycle(14). Immunofluorescence studies suggest that the KMP-11 protein is located mainly along the flagellum and flagellar base of the parasites. Parasites like L. infantum, T. congolense and T. brucei, have been shown to be immunoreactive primarily along the flagellum and flagellar pocket, while that presented at Leishmania donovani’s flagellar location differs, as it exhibited a pattern of fluorescence which was more widely dispersed throughout the cell (10, 11).
At present, the biological function of the KMP-11proteinis not yet fully understood, however, it clearly has three immunological roles; B-cell immunostimulatory, inducer lymphocyte proliferation and response cytotoxic and immunoprotective in animal models (15).The KMP-11 protein could induce the proliferation of T lymphocytes. Tolson et al. (1994) showed that the KMP-11 protein in L. donovani is a potent stimulator of CD4+, CD8-in mice immunized with KMP-11proteins (16). In addition, according to Basu et al. (2005) this protein is an excellent target for T cytotoxic lymphocytes (CD8+) and natural killer cells (NK). It has been revealed that KMP-11 induces significant production of IFN-γ cytokine from T-helper lymphocytes (17). Moreover, Trujillo et al. (1999) showed that the subclasses of immunoglobulins involved in the response fronts of KMP-11 in Leishmania, in order of reactivity are; IgG1, IgG3, IgG2 and IgG4 (18).
Considering the aforementioned findings, KMP-11 has been used previously for immunotherapy and immunization against leishmaniasis. Maranon et al. (2001) immunized transgenic mice with an A2/Kb fusion protein composed of the heat shock proteins HSP70 and KMP-11 in T. cruzi that induces a cytotoxic response against cells expressing the KMP-11in the parasite (19). Moreover, Planelles et al. (2002) have found that the fusion protein HSP70/KMP-11, is capable of acting as a stimulator for mature mouse dendritic cells and the consequent production of interleukin 12 (IL-12) and tumor necrosis factor(TNFα) (20). In addition, Ramirez et al. (2001) showed that immunization of BALB/c mice with an attenuated strain of Toxoplasma gondii expressing the protein Leishmania KMP-11, induces a specific immune response and immunoprotective in such animals (21). The study of Berberich et al. (2003), showed that immunization of mice with murine dendritic cells pulsed with a mixture of recombinant Leishmania antigens, including KMP-11 proteins, are able to control L. major infections (22).
In conclusion, the recombinant plasmid of pTKMP-11 that was successfully constructed in the present study could be used for future DNA vaccine production against leishmaniasis.