1. Background
Members of the Echovirus genus belonging to the Picronaviridae family are among the most common disease-causing viruses in humans, including echoviruses, coxsackieviruses, polioviruses and other echoviruses (1). More than 200 serotypes of echoviruses have been recognized (2). Echoviruses infect several mammals, humans, non-human primates, cattle and pigs (1). Echovirus infection is associated with significant morbidity and mortality in humans, especially in neonates (3). Molecular biology methods based on RT-PCR have recently been developed for echovirus typing (4), which may result in a more conclusive diagnosis compared to the neutralization of virus isolates in cell culture using standardized antisera (5). The neutralization method has been replaced by the sequence of VP1 region as the gold standard of echovirus typing (5). Meanwhile, the typing strategy, based on the partial sequencing of 5’ non-coding region (5’-NCR), has been developed to predict the type of echovirus, which is concordant with the VP1 region dependent analysis (6).
Simian echoviruses were originally isolated from primary cell cultures or captivated and wild-caught non-human primates in 1950s and 1960s (5, 7, 8). Some simian echoviruses are classified in the species of human echovirus A (HEV-A) on the basis of their sequence relationships to the human echoviruses in HEV-A, such as simian echovirus 46 and human echovirus 92 (7, 9). Human echovirus 92 (EV92), newly typed in 2008, was first detected in diarrheal rhesus macaques and pigtailed macaques (9). EV92 was detected more frequently than other types of echoviruses in simian diarrhea (9). However, there are limited information about the epidemiology of EV92 in the Southwest China, where most of the primates were harbored.
2. Objectives
From June 2008, Macaca mulatta, captured from the wildlife of the southwest China were affected by acute diarrhea and/or acute flaccid paralysis. To investigate the causative agents, primer pairs were designed based on the 5’-NCR of EV92 to detect the echovirus using RT-PCR method.
3. Materials and Methods
3.1. Stool Samples
Fresh stool samples (246) of diarrhoeal M. mulatta were collected for the echovirus detection. Samples were stored at -70℃ until used.
3.2. RT-PCR Detection
All the samples were suspended in phosphate-buffered saline (PBS), pH 7.4 with 0.01 % diethyl pyrocarbonate (DEPC), at a proportion of 10% (w/v). Total RNA was extracted from each sample by Trizol (Invitrogen, America) in accordance with the manufacturer’s directions. cDNA was transcribed from the total RNA (~3 ug) using an AMV Reverse Transcriptase XL for RT-PCR (Takara, Japan) according to the manufacturer’s directions. The reverse transcription primer was P2: 5’-CTGTGTGTGTAGAATCTGTTC-3’. RT-PCR was performed using the following primers: P1: 5’-TTTAAAACAGCCTGTGGGTTG-3’ and P2. The reverse transcription protocol was performed at 30ºC for 10 minutes,42ºC for 30 minutes and 99ºC for 5 minutes. The resulting cDNA (2 µl) was amplified by PCR at 94ºC for 2 minutes, followed by 94ºC for 30 seconds, 50ºC for 30 seconds and 72ºC for 1 minute, and the reaction was repeated for 29 cycles. The PCR products were detected by electrophoresis on agarose gel containing 0.5 μg/mL ethidium bromide. The expected DNA band, specific for the EV92, was extracted from the gel, purified with the AxyPrep DNA Gel Extraction kit (Axygen, USA) and cloned into the pMD 18 T-Vector (TaKaRa, Japan). Both strands of each inserted DNA amplicon were sequenced in a DNA analyzer (Applied Biosystems 3730 DNA Analyzer; Invitrogen, USA). Standard precautions were used for all procedures to reduce the possibility of sample contamination and RNA degradation.
3.3. Sequence Analysis
Sequences were aligned using Clustal X 1.8 or MegAlign program in the DNASTAR software package. Phylogenetic tree was constructed using the Mega 4 software, (http://www.megasoftware.net/) using the neighbor-joining method. A maximum-likelihood tree that showed the similar clustering is not shown.
4. Results
The sequence analysis suggested that one specimen from a four years old monkey was positive for EV92 RNA, which belongs to the human echovirus A group (HEV-A). The nucleotide sequence contained 1105 nucleotides, including the 5’ NCR, VP4 region and partial VP2 region, which shared the highest similarity (85%) with human echovirus 92 (GenBank No. EF667344.1). The coding region encoded a potential 134 -amino acid poly-protein.
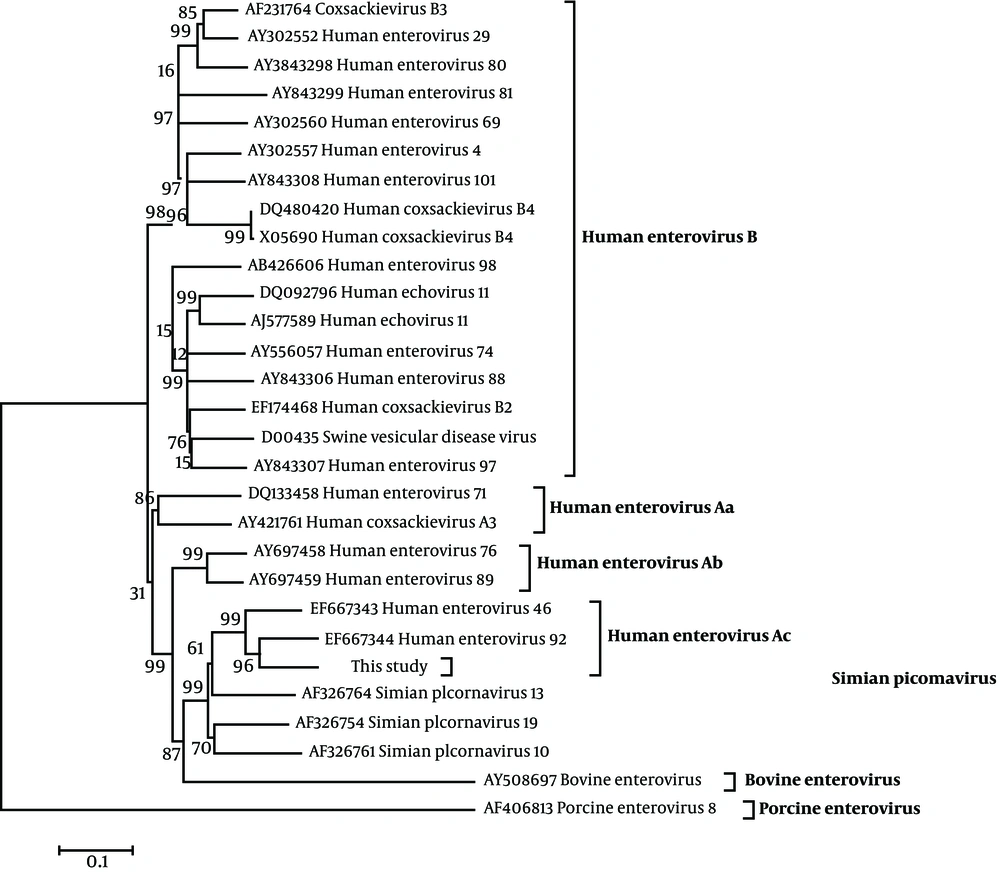
To investigate the evolutionary relationship of the virus isolated from the diarrhoeal monkey, phylogenetic relationships of the sequence were examined using twenty-eight echovirus reference strains (including simian, human, porcine and bovine echovirus original isolates) and the complete genome obtained from GenBank (Figure 1). The prototype of porcine echovirus (GenBank No. AF406813) and bovine echovirus (GenBank No. AY508697) were used as the out-group viruses for construction of the phylogenetic tree. Sequence analysis indicated that the isolate shared high identities with other reported simian echovirus isolates and the highest identity with EV92 (GenBank No. EF667344). Phylogenetic analysis indicated that the EV92 isolated in this study, cluster together with EV92 and SV46 isolates, separated in a US primate center ( 10 ), as shown in Figure 1 Human echovirus Ac subgroup.
The tree was evaluated using the interior branch test method with Mega 4 software. Percent bootstrap support is indicated at each node. The scale bar represents the nucleotide substitution per base. GenBank accession No. and name of the virus are indicated. The isolate identified in this study is marked with a triangle
5. Discussion
Most echovirus infections are asymptomatic or just cause mild illnesses, but some echovirus es such as some strains of EV76, EV89, EV90, EV91 and EV92, can cause clinical illnesses, including acute diarrhea and acute flaccid paralysis ( 9 - 11). In the present study, EV92 was detected in the stool of primates (rhesus macaques and pigtail macaques) possessing gastroenteritis. EV92 caused acute diarrhea in rhesus macaques, the clinical symptoms of which were similar to the previous studies ( 7 , 9 , 11). The phylogenetic relationships among human EV71 echoviruses (Figure 1) (Human echovirus, Aa subgroup), EV76 and EV89 (Figure 1) (Human echovirus, Ab subgroup), and EV92 isolated in this study (Figure 1) (Human echovirus, Ac subgroup), clustered in a unique phylogenetic clustering, suggested an unexplored relationship between human and simian echovirus es.
Previously, studies suggested that some simian echoviruses, SV19, SV46 and EV92 had little distinction between human and simian echovirus stains, and transmission from non-human primates to humans is possible (9, 11). A high prevalence of EV92 (about 56.7 %, 17/30) had been reported in diarrhoeal primates in a US primate center (9, 11).
In conclusion, although only one EV92 isolate was detected in this study (0.4 %, 1/246), EV92 breakout should be promptly alerted in this region of China. EV92 might be a new echovirus in non-human primates, causing acute diarrhea, and the probability of transmission across species should be warned.