1. Background
Pseudomonas aeruginosa is an important cause of nosocomial infections, including pneumonia, burn infection, urinary tract infections, meningitis and bacteremia. The infections can be particularly develop to a severe form in immune deficient patients (1). Antibiotics have been used successfully for several decades, but resistance genes have emerged and disseminated particularly in the last few years (2).
Extended-spectrum β-lactamases (ESBLs) mediate resistance to various broad-spectrum cephalosporins, including cefotaxime, ceftriaxone, ceftazidime, and aztreonam (3). These enzymes originally collected from Klebsiellapneumoniae and Escherichia coli and recently from P. aeruginosa (4-7). Most of the methods for detection of ESBLs are used in bacterial species such as Klebsiella and E. coli lacking chromosomal β-lactamase activity (8, 9), However, the detection of ESBL production in P. aeruginosa has some difficulties, because this bacterium not only has an inducible AmpC enzyme but also has an efflux-mediated resistance and a higher degree of impermeability than Enterobacteriaceae (10, 11). The PER-1 and VEB-1 type ESBLs belong to class A of β- lactamases and is associated with high level of resistance to cephems, monobactams and ceftazidime (12, 13).
2. Objectives
The aim of this study was to determine the antibacterial resistance patterns and prevalence of PER-1 and VEB type ESBLs among P. aeruginosa isolated from patients in west of Iran.
3. Materials and Methods
3.1. Bacterial Isolates
A total of 106 isolates of P. aeruginosa recovered from various clinical specimens (Table 1) in Beasat Teaching Hospital at the Hamadan University of Medical Sciences during a 7-month study in 2009. The identification were carried out by colonial morphology, positive oxidase test, pigment formation; growth test at 42ºC on nutrient agar, Gram staining and motility test.
| Source | Isolates, No. (%) |
|---|---|
| Burn wounds | 55 (51.9) |
| Trashes | 20 (18.9) |
| Urine | 12 (11.3) |
| Blood | 7 (6.6) |
| Feces | 6 (5.7) |
| Sputum | 5 (4.7) |
| CSFa | 1 (0.9) |
| Total | 106 (100) |
Distribution of P. seudomonas aeruginosa by Site of Isolation
3.2. Antibiotic Susceptibility Test
Antibiotic susceptibility of the isolates was determined by standard disk diffusion method (14). The following antibiotics were used: gentamicin (30 µg), aztreonam (30 µg), meropenem (10 µg), imipenem (10 µg), amikacin (30 µg), tobramycin (30 µg), piperacillin (30 µg), ceftazidime (30 µg), ciprofloxacin (5 µg), ofloxacin (5 µg), cefepime (30 µg) and cefotaxime (30 µg) (Himedia, India). P. aeruginosa ATCC 27853 was used as control. The results were interpreted according to the clinical and laboratory standards institute (CLSI) (15).
3.3. Phenotypic Detection of Beta-Lactamase
The isolates were tested for the ESBLs production by using combine disk test (CDT) as CLSI recommendations. CDT were performed on ceftazidime, cefotaxime, cefepime and aztreonam resistant strains by placing disks of ceftazidime, and cefotaxime (30 µg each) at a 20 mm distance from a disk containing ceftazidime-clavulanic acid (30/10 μg), cefotaxime-clavulanic acid (30/10 µg) and cefepime-clavulanic acid (30/10µg) (16). ESBL production was inferred when the cephalosporin inhibitory zones were expanded by the clavulanate.
3.4. PCR Amplification
PCR amplifications were done using specific primers for the β-lactamases PER-1, and VEB-1 genes, as described previously (17). PCR was performed for all ESBL-producers which their resistance to cephalosporins and phenotypic was identified by confirmatory tests. DNA was extracted by the boiling method as previously described (17). The DNA amplification program consisted of an initial denaturation (94 °C, 5 minutes) followed by 35 cycles of denaturation (94 °C, 60 seconds), annealing (50 °C for PER-1 and 55 °C for VEB-1, 60 seconds), extension (72 °C, 45 seconds) and a single final extension for 5 minutes at 72 °C. Reaction mixtures for PCR contained 1.5mM MgCl2, 0.5 mM of each primer, 0.2 mM of dNTPs, 1 U of Taq polymerase, 1X PCR buffer and 2 µL of DNA. Primers PER-F (5-AATTTGGGCTTAGGGCAGAA-3') and PER-R (5'-ATGAATGTCATTATAAAAGC-3') were used for blaPER-1; and VEB-F (5'-CGACTTCCATTTCCCGATGC-3') and VEB-R (5'-GGACTCTGCAACAAAT AC GC-3') were used for blaVEB-1 amplification.
4. Results
4.1. Antimicrobial Susceptibility Test
Table 2 shows the antimicrobial susceptibility pattern of P. aeruginosa strains. Ciprofloxacin (91.5%), imipenem
(84.9%) and meropenem (82.1%) were the most active antimicrobial agents followed by ofloxacin (67.9% susceptibility). Aztreonam, a monobactam, and amikacin showed antibiotic activity against 66% of the strains. Susceptibility to the cephalosporins was reduced to 37.7% for ceftazidime, followed by cefotaxime (9.4%) and Cefepime (0.9%).
| Antimicrobials | Resistance, No. (%) | Sensitive, No. (%) | Intermediate, No. (%) |
|---|---|---|---|
| Amikacin | 32 (30.2) | 70 (66) | 4 (3.8) |
| Ceftazidime | 54 (51) | 40 (37.7) | 12(11.3) |
| Aztreonam | 29 (27.4) | 70 (66) | 7 (6.6) |
| Cefepime | 103 (97.2) | 1 (0.9) | 2 (1.9) |
| cefotaxime | 53 (50) | 10 (9.4) | 43 (40.6) |
| Ciprofloxacin | 5 (4.7) | 97 (91.5) | 4 (3.8) |
| Gentamicin | 39 (36.8) | 55 (51.9) | 12 (11.3) |
| Imipenem | 8 (7.5) | 90 (84.9) | 8 (7.5) |
| Meropenem | 14 (13.2) | 87 (82.1) | 5 (4.7) |
| Ofloxacin | 31 (29.2) | 72 (67.9) | 3 (2.8) |
| Piperacillin | 99 (93.4) | 7 (6.6) | 0 |
| Tobramycin | 39 (36.8) | 67 (63.2) | 0 |
Antimicrobial Susceptibility Pattern of P. aeruginosa Strains
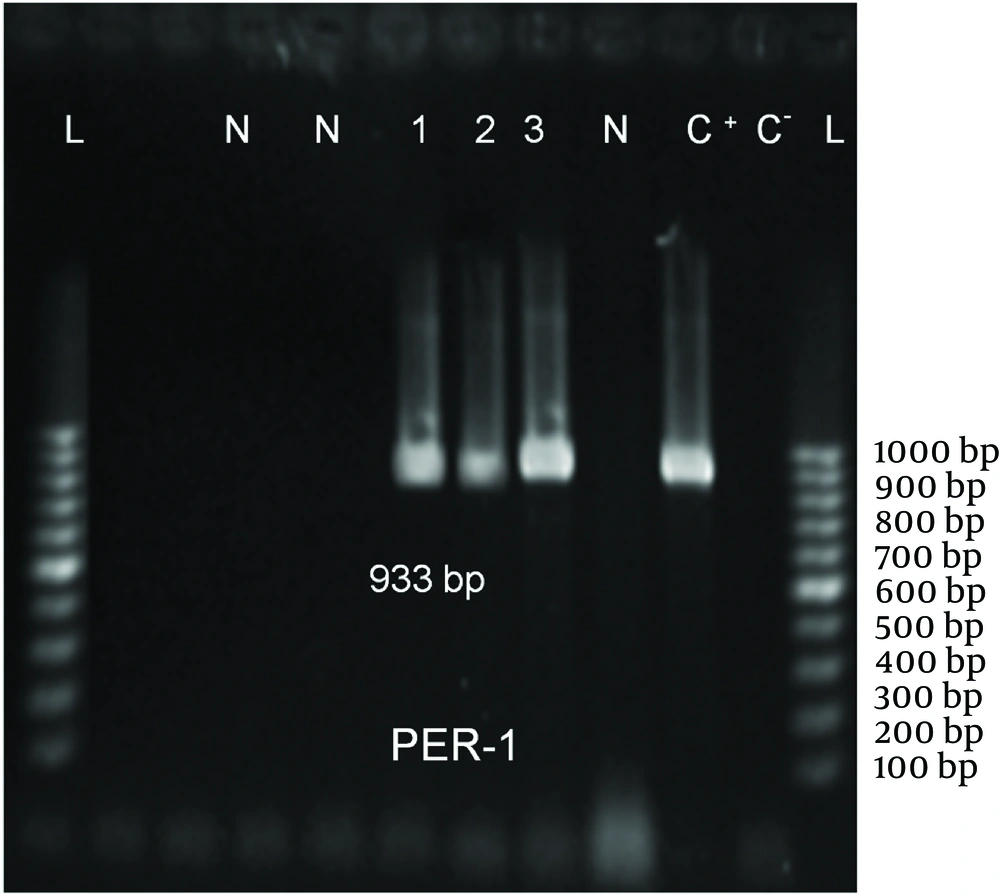
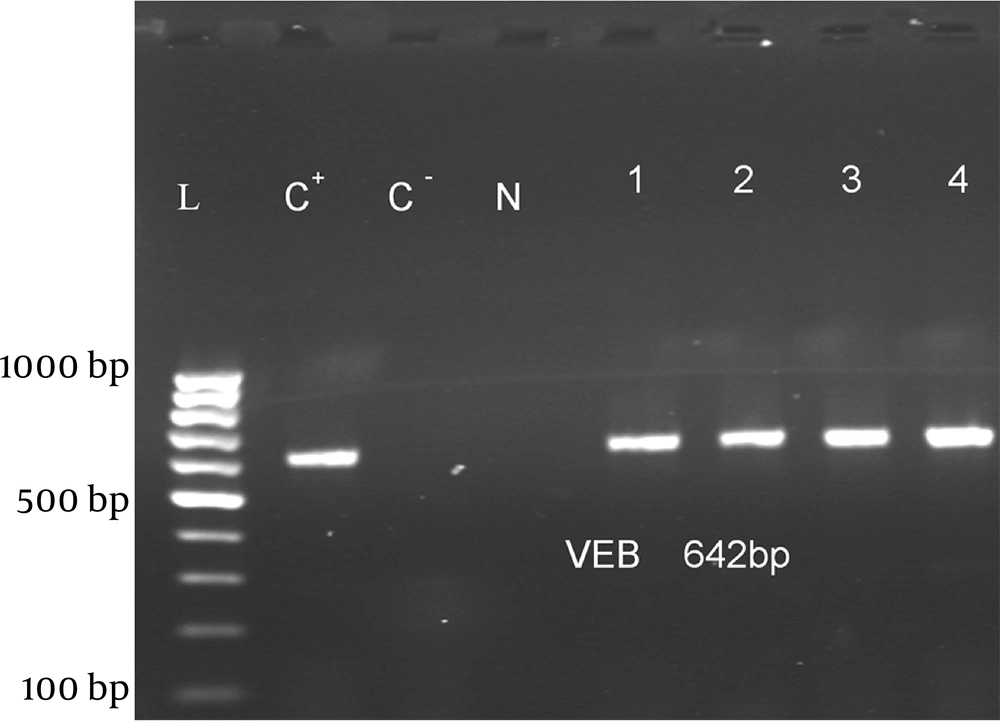
ESBL Production: Among 106 isolates, 94 (88.7%) were multidrug resistant and 60(58.3%) were putative ESBL producers using phenotypic confirmatory tests (Table 3). Sixteen (26.6%), 9 (15%) and 3 (5%) strains among 60 ESBL-producing strains had blaPER-1 (Figure 1), blaVEB-1 (Figure 2), and blaPER-1- blaVEB related genes, respectively (Table 4).
| ESBL Positive (n=60) | ESBL Negative (n=46) | |||||
|---|---|---|---|---|---|---|
| Antimicrobials | Resistant, No. (%) | Susceptible, No. (%) | Intermediate, No. (%) | Resistant, No. (%) | Susceptible, No. (%) | Intermediate, No. (%) |
| Amikacin | 29 (48.3) | 30 (50) | 1 (1.7) | 4 (9.3) | 37 (86) | 2 (4.7) |
| Ceftazidime | 40 (66.7) | 18 (30) | 2 (3.3) | 13 (28.3) | 24 (55.2) | 6 (13) |
| Aztreonam | 25 (41.7) | 32 (53.3) | 3 (5) | 3 (6.97) | 39 (90.69) | 1 (2.32) |
| Cefepime | 58 (96.7) | 0 | 2 (3.3) | 42 (97.67) | 1 (2.32) | 0 |
| cefotaxime | 41 (68.3) | 6 (10) | 13 (21.7) | 12 (27.90) | 3 (6.97) | 28 (65.11) |
| Ciprofloxacin | 5 (8.3) | 50 (83.3) | 5 (8.3) | 0 | 42 (97.67) | 1 (2.32) |
| Gentamicin | 32 (53.3) | 28 (46.7) | 0 | 7 (16.27) | 36 (83.72) | 0 |
| Imipenem | 7 (11.7) | 45 (75) | 8 (13.3) | 1 (2.32) | 41 (95.34) | 1 (2.32) |
| Meropenem | 13 (21.7) | 44 (73.3) | 3 (5) | 2 (4.65) | 39 (90.69) | 2 (4.65) |
| Ofloxacin | 30 (50) | 29 (48.3) | 1 (1.7) | 4 (9.30) | 37 (86.04) | 2 (4.65) |
| Piperacillin | 54 (90) | 6 (10) | 0 | 41 (95.34) | 2 (4.65) | 0 |
| Tobramycin | 31 (51.6) | 28 (46.7) | 1 (1.7) | 4 (9.30) | 39 (90.69) | 0 |
Susceptibility Pattern to Antimicrobial Agents in ESBLs and non ESBLs-producing P. aeruginosa Strains
| PER-1:16 (26.6%) | VEB: 9 (15%) | |||||
|---|---|---|---|---|---|---|
| Antimicrobials | Resistant, No. (%) | Susceptible, No. (%) | Intermediate, No. (%) | Resistant, No. (%) | Susceptible, No. (%) | Intermediate, No. (%) |
| Amikacin | 10 (62.5) | 5 (31.25) | 1(6.25) | 5 (55.5) | 2 (22.2) | 2 (22.2) |
| Ceftazidime | 12 (75) | 4 (25) | 0 | 9 (100) | 0 | 0 |
| Aztreonam | 10 (62.5) | 4 (25) | 2 (12.5) | 9 (100) | 0 | 0 |
| Cefepime | 16 (100) | 0 | 0 | 9 (100) | 0 | 0 |
| cefotaxime | 87.5 (14) | 0 | 2 (12.5) | 9 (100) | 0 | 0 |
| Ciprofloxacin | 2 (12.5) | 13 ( 81.25) | 1 (6.25) | 2 (22.2) | 6 (66.6) | 1 (11.11) |
| Gentamicin | 10(62.5) | 6 ( 37.5 ) | 0 | 6 (66.6) | 3 (33.3) | 0 |
| Imipenem | 5 (31.25) | 5 (31.25) | 6 (37.5) | 1 (11.11) | 6 (66.6) | 2 (22.2) |
| Meropenem | 7 ( 43.75) | 8 (50) | 1 (6.25) | 6 (66.6) | 2 (22.2) | 1 (11.11) |
| Ofloxacin | 11 (68.8) | 5 (31.25) | 0 | 8 (88.8) | 1 (11.11) | 0 |
| Piperacillin | 16 (100) | 0 | 0 | 9 (100) | 0 | 0 |
| Tobramycin | 10 (62.5) | 6 ( 37.5 ) | 0 | 6 (66.6) | 3 (33.3) | 0 |
Association of Antimicrobial Resistance Pattern and ESBL Genotypes
5. Discussion
P. aeruginosa has a high resistance to antibiotics and is a common cause of morbidity and mortality in hospitalized and immunocompromised patients (18). Treatment of P. aeruginosa infections is complicated by the inhered and acquired resistance to the most of commonly used antimicrobial agents (19). The results from this study showed the high resistance of P. aeruginosa to most of used antimicrobial agents. It was also demonstrated that the prevalence of antibiotic resistance of the isolates was very high in comparison to other studies and most of P. aeruginosa isolates (88.7%) were multi-drug resistant (resistant to ≥ 3 different antibiotic classes) (20-25).
The prevalence of ESBL- producing P. aeruginosa isolates in this study was also higher than other investigations (5, 20, 26, 27). Among 60 ESBL-positive strains, 16 (26.6%), 9 (15%) and 3 (5%) contained PER-1, VEB-1 and PER-1-VEB-1 genes, respectively. These results indicated that the prevalence of VEB-1 gene in our area, is higher than Turkey and Korea, but the prevalence of VEB-1 and PER-1 genes, is lower than in Thailand (94.44% blaVEB-1) and Italy (34.61% blaPER-1) (5, 13, 26). Data on the prevalence of ESBL- producing P. aeruginosa strains in our area is limited. In the study performed by Shahcheraghi and colleges on P. aeruginosa isolates in Tehran, the rate of blaVEB and blaPER ESBLs were reported 24% and 17%, respectively, that was similar to our results (28). The high prevalence of PER-1 and VEB-1 indicated the high resistance to penicillins, ceftazidime and cefotaxime, as reported by other studies (26, 28).
This is the first report about the presence of these enzymes in P. aeruginosa isolates from west of Iran. It has shown that ESBL production in strains of P. aeruginosa can greatly complicate the clinical management of infection if advanced care is not taken. However, further studies are needed to determine other ESBL- types in P.aeruginosa strains in this area and their role in resistance to other antibiotic classes. The results of this study emphasizes on the need for a surveillance network to monitor the trends and emerge of new resistance mechanism in P. aeruginosa from different geographic regions. Therefore, the improvement in antibiotic prescription policies and infection control programs are of high necessity to prevent the spread of such resistant infectious agents.