1. Background
Pseudomonas aeruginosa is a major microorganism, a cause of nosocomial infections, particularly in burn patients (1). The treatment of infections caused by P. aeruginosa is frequently complicated since the organism is intrinsically resistant to many drug classes and is able to acquire resistance to all the effective antibiotics (2). In many studies, the term multidrug-resistant Pseudomonas aeruginosa (MDRPA) has been used to report isolates with resistant to at least three different classes of antimicrobial agents, mostly aminoglycosides, carbapenems, antipseudomonal penicillins, quinolones, and cephalosporins (3). P. aeroginosa also possesses a large number of virulence factors such as exotoxin A, exoenzyme S, elastase and sialidase, which are powerfully regulated by cell-to-cell signaling systems (4). The major virulence factor produced by most of the P. aeruginosa isolates is exotoxin A (ETA) which plays a major role in the pathogenesis of infections caused by this organism (5). An extracellular neuraminidase is also thought to have a key role in the implantation of the bacterium, but the genetic basis of this process is still unknown (6).
Biofilms are sessile populations of microorganisms which are enclosed by the self-secreted extracellular polysaccharide matrix, or slime. Biofilms act as efficient barriers against antimicrobial agents (7). Previous studies have shown that MDRPA are widespread among Iranian hospitals (8). Despite the high incidence of P. aeruginosa infection in burn patients, there is currently little information on the distribution of the virulence factors and the ability of biofilm production among the isolates of P. aeruginosa in burn patients in Iran.
2. Objectives
The aim of this study was to determine the antibiotic susceptibility pattern, the prevalence of MDRPA, evaluate the nan1 (gene encoding for neuraminidase) and exoA (gene encoding for exotoxin A) genes and the prevalence of biofilm formation in clinical isolates of P. aeruginosa in Shahid Motahari Burn hospital in Tehran, Iran
3. Patients and Methods
3.1. Bacterial Isolated and Identification Test
A total of 144 isolates of P. aeruginosa were collected from wound infections of burn patients admitted to Shahid Motahari Burn Hospital in Tehran, Iran, between March and July 2013. Each isolate was identified according to the standard bacteriological methods including colony morphology, Gram staining, oxidase test, pyocyanin pigment production, growth at 44˚C, catalase, and oxidative-fermentative (OF) tests (9).
3.2. Antimicrobial Susceptibility Testing
Antimicrobial susceptibility was determined according to the Clinical and Laboratory Standards Institute (CLSI) guideline (10), using the Kirby Bauer disk diffusion assay on Mueller-Hinton agar. The susceptibility profiles were determined for seven antibiotics including piperacillin (PIP, 100 µg), cefotaxime (CTX, 30 µg), ceftazidime (CAZ, 30 µg), gentamicin (GE N, 10 µg), amikacin (AMK, 30 µg), ciprofloxacin (CIP, 5 µg), and imipenem (IPM, 10 µg) (Mast Diagnostics, Mast Group Ltd, Merseyside, UK). Pseudomonas aeruginosa ATCC 27835 was used as quality control in each antimicrobial susceptibility assay. The results were interpreted as susceptible or resistance according to the criteria recommended by the CLSI and the manufacture’s protocols (Mast Companies, UK).
3.3. Polymerase Chain Reaction Amplifications
The exoA and nan1 genes were amplified by PCR using a specific set of primers (Table 1). Bacterial DNA for the PCR analysis was prepared using the boiling method (11). The amplifications of the exoA and nan1 genes were performed in a 25 μL reaction mixture containing 2.5 µL PCR buffer, 0.5 μL dNTPs, 0.5 μL of each primer, 1.5 μL MgCl2 and 0.2 μL Taq DNA polymerase (CinnaGen, Iran). Ultra-pure water was then added to make up a final volume to 25 µL. The amplification was performed as follows: initial denaturation step at 94˚C for two minutes (one cycle), followed by 30 cycles consisting of, denaturation at 94˚C for two minutes, annealing at 68˚C for one minutes, extension at 72˚C for one minute and final extension at 72˚C for seven minutes.
For the nan1 gene, an initial denaturation (95˚C, three minutes), followed by 30 cycles of denaturation (94˚C, 30 seconds), annealing (55˚C, 30 seconds) and extension (72˚C, 1 minute and 30 seconds), and a single final extension (72˚C, 5 minutes) were performed. The PCR products were visualized by electrophoresis using 1% agarose gel after staining with ethidium bromide.
3.4. Biofilm Formation
The P. aeruginosa isolates were analyzed for their ability to produce biofilm using microtiter dish biofilm formation assay (14). In this method, the P. aeruginosa isolated were grown overnight at 37˚C in tryptic soy broth (TSB) containing 0.25% glucose. The cultures were diluted 1:100 in TSB medium. Sterile flat-bottomed 96-well poly styrene microtiter plates were inoculated with 125 µL of the bacterial suspension and incubated for 24 hours at 37˚C without agitation. The wells were washed three times with 300 µL distilled water, dried in an inverted position at room temperature and finally stained with 125 µL of 0.1% crystal violet solution in water for about 10 - 15 minutes. After staining, the wells were washed three times with distilled water. The wells were destained with 125 μL of 30% acetic acid in water. A new sterile flat-bottomed 96-well poly styrene microtiter plate was inoculated with 125 µL destaining solution in each well. The absorbance of the destaining solution was measured at 570 nm using an ELISA reader (Stat Fax-2100). Each test was performed in triplicate. As control, uninoculated medium was used. Based on the optical density of the samples (ODi) and on the average of the optical density of the negative control (ODc), the samples were classified as strong (4xODc < ODi), mod rate (2xODc < ODi ≤ 4xODc), weak (ODc < ODi ≤ 2xODc), or non-producer of biofilm (ODi < ODc).
Statistical Package for Social Sciences (SPSS) software (SPSS Inc. No. 16) was used for statistical analyses. Fischer exact test or χ2 test was used for the analysis of the categorical data. P value < 0.05 was considered statistically significant.
4. Results
The antibiotic susceptibility patterns of the P. aeruginosa isolates are shown in Table 2. A high rate of resistance was seen against ciprofloxacin (93.7%), aztreonam (86.8%), piperacillin (85.4%), ceftazidime (82.6%), amikacin (82%), and imipenem (79.2%). The least resistance was seen to gentamicin (11.1%). A total of 2 (1.4%) isolates were resistant to all the tested antibiotics. In total, 93.1% of the isolates showed resistance against at least three different classes of antimicrobial agents and were identified as MDRPA.
| Antibiotics | Sensitive | Intermediate | Resistant |
|---|---|---|---|
| Amikacin | 12 (8.3) | 14 (9.7) | 118 (82) |
| Imipenem | 12 (8.3) | 18 (12.5) | 114 (79.2) |
| Ciprofloxacin | 5 (3.5) | 4 (2.8) | 135 (93.7) |
| Ceftazidime | 24 (16.7) | 1 (0.7) | 119 (82.6) |
| Gentamicin | 117 (81.3) | 11 (7.6) | 16 (11.1) |
| Aztronam | 11 (7.6) | 8 (5.6) | 125 (86.8) |
| Piperacillin | 16 (11.1) | 5 (3.5) | 123 (85.4) |
a The values are presented as No. (%).
Figure 1 shows the amplification for the presence of the exoA gene. Figure 2 shows the amplification for the presence of the nan1 gene. The frequencies of the presence of virulence genes in all the studied isolates were as follows: exoA and nan1 in 75% and 11.8% of the isolates, respectively. There was no correlation between the distribution of exoA and nan1 and MDRPA (P > 0.05).
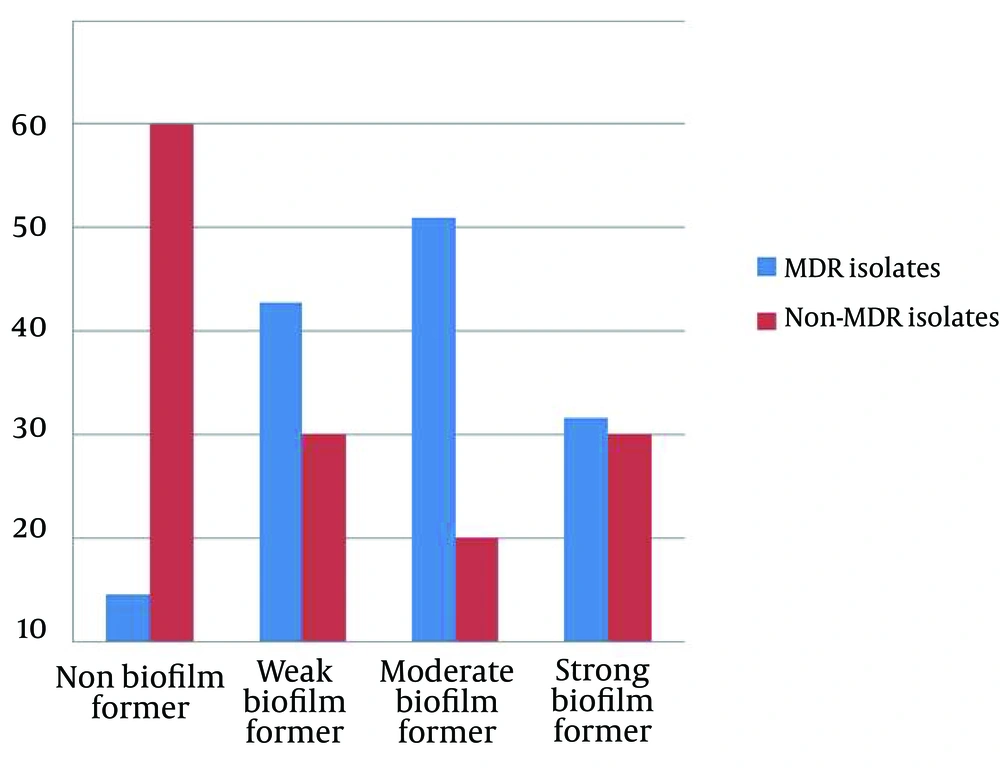
Quantitative biofilm determination using the microtiter assay revealed that 133 isolates (92.4%) produced biofilm and the remaining 11 isolates were non-biofilm producers. The relation between biofilm formation and MDRPA is shown in Figure 3. The statistical analysis to examine the link between antibiotic resistance and biofilm formation showed that the biofilm production in MDRPA isolates was significantly higher than that in the non–MDRPA isolates (P < 0.001). Table 3 shows the comparative frequency (as percentages) of virulence factors among the MDRPA and non-MDRPA isolates. The proportion of MDRPA isolates containing two (of three) virulence factors was higher than the proportion of non-MDRPA isolates with two virulence factors (P < 0.01) (Table 3).
| No. of Virulence Factors | MDRPA | Non-MDRPA |
|---|---|---|
| 0 | 5 (3.7) | 1 (10.0) |
| 1 | 22 (16.4) | 4 (40.0) |
| 2 | 96 (71.6 | 3 (30.0) |
| 3 | 11 (8.2) | 2 (20.0) |
| Total | 134 | 10 |
a Abbreviations: MDRPA, multidrug-resistant Pseudomonas aeruginosa; non-MDRPA, non-multidrug-resistant Pseudomonas aeruginosa.
b The values are presented Vas No. (%).
5. Discussion
Pseudomonas aeruginosa remains one of the most important opportunistic causes of nosocomial infections and it has developed resistance to a range of antimicrobial agents in burn centers (1). We carried out this study to determine the antibiotic resistance pattern, the prevalence of MDRPA, exoA and nan1 genes, and biofilm formation in P. aeruginosa isolated from burn patient of Shahid Motahari Burn Hospital in Iran. According to the results, there was a high frequency (> 75%) of resistance against all the tested antibiotics, except for gentamicin. These results indicated a severe antimicrobial resistance among P. aeruginosa in Motahari Hospitals in Tehran which might be due to the unsuitable use of antibiotics in this setting. In a study in 2013 at the Burn Centre of Guilan in north of Iran by Nikokar et al. (15) the percentage of resistance to tested antibiotics was as follows: imipenem 97.5%, amikacin 90%, piperacillin 87.5%, gentamicin 67.5%, ciprofloxacin 65%, and ceftazidime 57.5%. Interestingly, our results showed that the percentage of resistance to gentamicin was lower in comparison with previous reports published from our country (16, 17).
Over the recent years, several reports confirmed an increasing multidrug resistance among P. aeruginosa isolated from burn wound infections in Iranian hospitals (18, 19). Our study showed that the frequency of MDRPA was 93.1%. This high frequency might be due to the prolonged hospital stay and intensive use of antibiotics. MDR in P. aeruginosa can be mediated by means of some mechanisms including the production multidrug efflux systems, enzyme production, or outer membrane protein (porin) loss and target mutations (20). Carbapenems are the effective antibiotic against MDR isolates. However, the increasing frequency of carbapenem-resistant P. aeruginosa has recently become a worldwide challenge (21). Our results showed high resistance to imipenem. Previously, resistance to imipenem in Tehran was reported to be within the range of 16% - 100% (22).
In P. aeruginosa infections, Biofilm production has been measured as an important determinant of pathogenicity (23). Our data identified that 92.4% of the P. aeruginosa isolates produced biofilm. In a study by Jabalameli et al. (24) in Iran, 96.9% of the isolates produced biofilm, which is in correlation with our results. The study of the relationship between biofilm formation and antibiotic resistance/susceptibility revealed that the MDR isolates displayed significant biofilm production as compared to susceptible isolates, probably due to the delayed penetration of antimicrobial agents inside the bacterial cell. Our results are consistent with Abidi et al. (25) reports. Fluoroquinolones are effective antibiotics against biofilm-forming bacteria (26); but, our results showed high resistance to ciprofloxacin (fluoroquinolones), since ciprofloxacin is one of the most currently prescribed classes of antibiotics in burn centers in Iran (22).
The virulence of P. aeruginosa depends on several factors. ETA is one of the most toxic factors secreted by P. aeruginosa. ETA is encoded by the exoA gene (5). In this study, the exoA gene was found in 75% of the isolates by PCR tests. Khan and Cerniglia reported that ETA gene was detected in 97% of P. aeruginosa isolates by PCR (12). The nan1 gene can play an important role in the pathogenesis of P. aeruginosa infections (6). The nan1 gene was found in 11.8% of the isolates. Similar to our results, Mitov et al. (13) found that the prevalence of nan1 among P. aeruginosa clinical isolates was 21.3%. The low prevalence of this factor among the isolates from burn infections may show that the role of this gene in burn infections is less important. In this study, there was no correlation between the distribution of exoA and nan1 and being MDRPA. In contrast with this study, Mitov et al. (13) found that the percentage of three genes including pilB, nan1 and exoU manifested a significantly higher spread (P < 0.001) among MDRPA compared with non-MDRPA isolates.
In conclusion, we presented a significantly high spread of biofilm formation among MDRPA isolates for the first time in Iran. However, there was no correlation between the distribution of the nan1 and exoA genes with MDRPA. The simultaneous determination of virulence factors and antimicrobial resistance is the contemporary approach for the examination of the microbiological aspects of infections caused by P. aeruginosa. Finally, our work revealed a significant difference between the MDRPA containing two virulence factors (of three) and the non-MDRPA. However, MDRPA isolates which possess virulent phenotypes remain a controversial issue and further work is necessary.