1. Background
Enterococci are important nosocomial pathogens (1). They cause various infections including urinary tract infections, sepsis, endocarditis, wound infections and bacteremia (2). Amongst enterococci, Enterococcus faecalis is the most common species among clinical enterococcal infections. On the other hand, E. faecium causes 10 - 20 percent of infections yet in comparison to other enterococci species, they show higher antibiotic resistance (1, 3).
Enterococci, not only carry intrinsic resistance against some antibiotic agents, they can also acquire resistance via mutation, transposable elements or conjugative plasmids (1). In addition, the ability of enterococci to form biofilm has also been of growing concern in the recent years. The use of indwelling catheters can intensify biofilm production of enterococci, which would lead to increase in urinary infections in catheterized patients (4). Antibiotic resistance and biofilm formation in bacteria have been under intensive investigation in urinary tract infections and endocarditis, during the recent years (1), in particular the pili and its gene cluster (ebp). This attention, of course, is worthwhile because biofilms can be the source of 80% of chronic infections in humans and their presence can increase antibiotic concentration by up to 1,000 folds (5, 6).
2. Objectives
In this study, we assessed the antibiotic resistance and biofilm formation ability of E. faecalis strains isolated from environmental and clinical samples in Tehran, Iran. In addition, the polymorphisms of ebp genes were determined.
3. Materials and Methods
3.1. Sampling, Isolation and Identification
From October 2009 to April 2010, a total of 90 clinical and hospital environmental isolates of E. faecalis were collected from two main hospitals in Tehran, Iran (Shariati Hospital (SH) and Shahid Rajaie Hospital (SRH)). Overall, 58 of the isolates were from patient samples and 32 from environmental sites. The clinical isolates were collected from blood, wound, suction secretion, sterile body fluid and urine of catheterized patients. Environmental samples were collected from patients’ bathroom, beds and tables and staff’s bathrooms and tables, as well as ventilators and oxygen pumps in the patients’ rooms.
The collected samples from hospitals were cultured on blood agar and incubated at 37˚C for 24 hours. Next, the isolates were cultured on m-Enterococcus agar (Becton Dickinson Co., MD, USA). Enterococcus faecalis strains were detected based on colony morphology, gram staining and polymerase chain reaction (PCR). To amplify genus and species specific gene (sodA) of E. faecalis, PCR was performed as mentioned before (7).
3.2. Antimicrobial Susceptibility
Antimicrobial susceptibility test was performed by the disk diffusion method, according to the Clinical and Laboratory Standards Institute (CLSI) guidelines, for nine antimicrobial agents including: gentamycin (G) (10 µg), ciprofloxacin (CIP) (5µg), ampicillin (AMP) (10 µg), co-trimoxazole (TS) (25 µg), erythromycin (E) (15 µg), tetracycline (T) (30 µg), linezolid (LZD) (30 µg), teicoplanin (TEC) (30 µg) and vancomycin (V) (30 µg) (Mast Diagnostics Ltd, UK).
3.3. Biofilm Formation Assay by the Quantitative Method
Microtiter plate test was used to quantitatively assay biofilm formation as mentioned previously. The cut-off calculation was used to interpret the test results by the formula presented below: Cut-off = (mean (OD) ± 3SD (OD)) + OD (Blank)). Strains with OD of ≤ 0.45, 0.45 - 0.55 and ≥ 0.55 were considered as non-adherent, weakly adherent and strongly adherent, respectively (4, 6, 8, 9).
3.4. Polymerase Chain Reaction Test to Detect Endocarditis and Biofilm-Associated Pili Genes (ebpA, ebpB, ebpC) and Sortase C (srtC)
Amplification of ebpA, ebpB, ebpC and srtC genes was done as described by previous studies (10, 11).
3.5. Restriction Fragment Length Polymorphism of Amplified Genes of ebp Operon
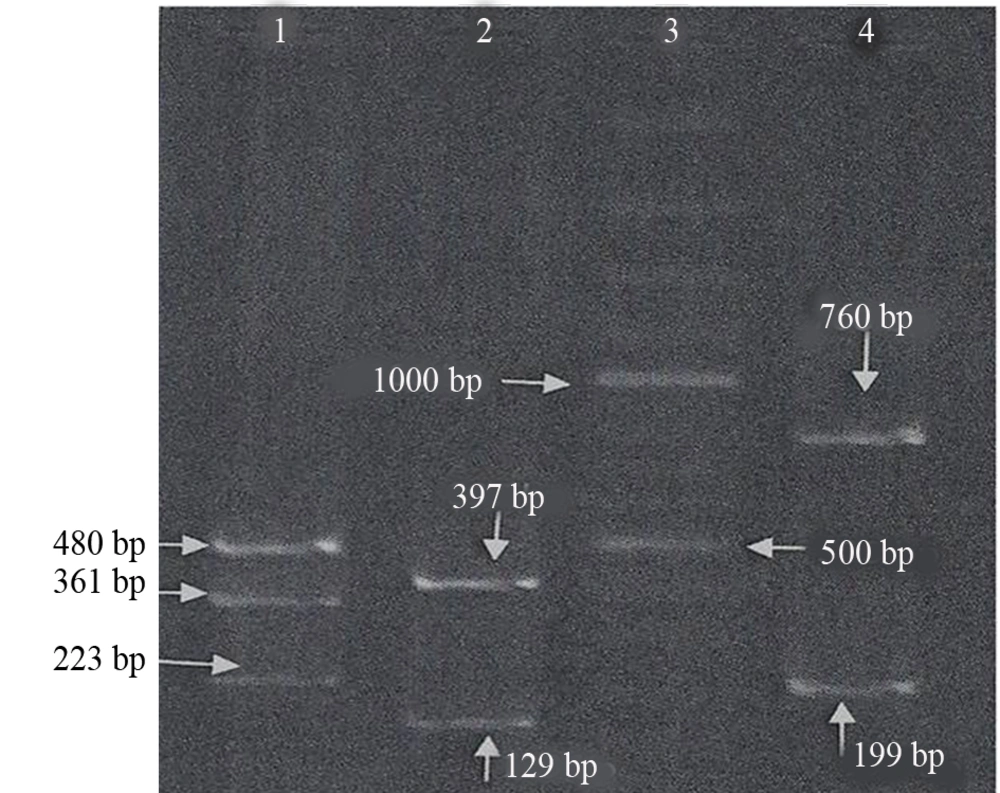
For amplified ebpA, ebpB and ebpC, gene products were digested by HaeIII, EcoRI and HindIII restriction enzymes, respectively. The results showed that ebpA gene digestion by HaeIII resulted in three fragments with a size of 223 bp, 480 bp and 361 bp. EcoRI digestion of ebpB produced 397 bp and 129 bp fragments. The 963 bp ebpC gene was divided to two fragments with a length of 760 bp and 199 bp by Hind III.
4. Results
4.1. Bacterial Isolates and Identification of Enterococcus faecalis Strains
Out of a total of 80 patient isolates of enterococci, 58 were E. faecalis. From the two hospitals, 49 and 9 isolates were from SH and SRH, respectively. Overall, hospital environmental samples were obtained from 250 different locations. The distribution of isolates obtained from patients and hospital enviroments are shown in Table 1.
| Source of Isolates | Values |
|---|---|
| Clinical Isolates | |
| Urinary culture | 16 (27) |
| Blood culture | 14 (24) |
| Wound culture | 11 (19) |
| Catheter | 8 (14) |
| Suction secretion | 5 (9) |
| Body sterile fluid | 4 (7) |
| Total | 58 (100) |
| Environmental Isolates | |
| Staffs’ bathroom | 7 (21.9) |
| Patients’ bathroom | 5 (15.6) |
| Staffs’ table | 5 (15.6) |
| Patients’ table | 3 (9.4) |
| Patients’ bed | 5 (15.6) |
| Ventilator | 4 (12.5) |
| Oxygen pump | 3 (9.4) |
| Total | 32 (100) |
a The values are presented as No. (%).
4.2. Antibiotic Resistance of Isolates
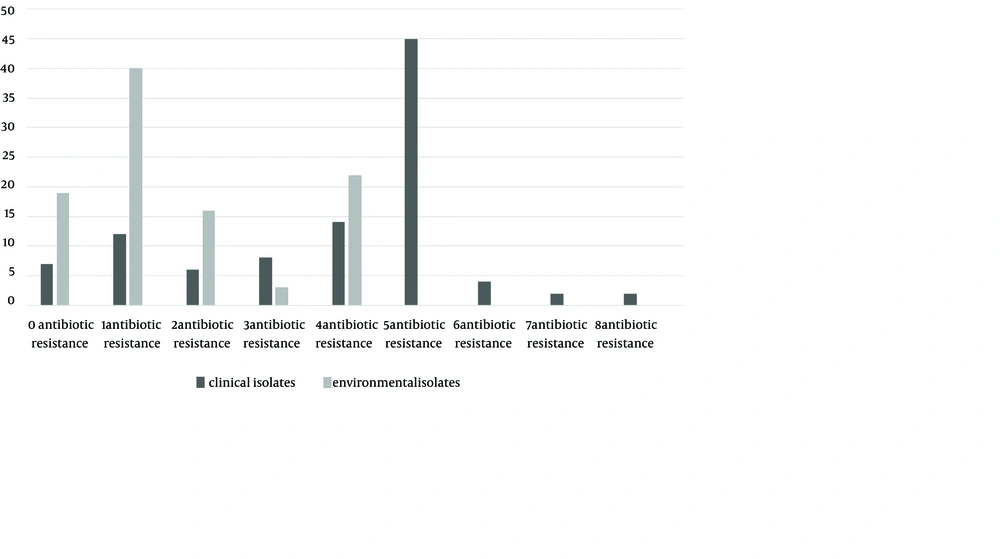
Among the clinical isolates, 86% of isolates were resistant to tetracycline (86%). The resistance observed against teicoplanin (3%), vancomycin (9%), linezolid (9%) and ampicillin (9%) was low. Among environmental isolates only 25% were sensitive to tetracycline with no resistance against vancomycin, teicoplanin, linezolid and ampicillin. Overall, 20 different antibiotic resistance patterns were obtained for 58 patient isolates and nine for environmental isolates (Figure 1).
4.3. Identification of Virulence and Biofilm Formation Genes
Polymerase chain reaction product sizes for ebpA (1064 bp), ebpB (530 bp) and ebpC (964 bp) were found in 50 (86%), 55 (95%) and 56 (97%) of the patient isolates and 29 (91%), 30 (94%) and 29 (91%) of environmental isolates, respectively. The PCR product of srtC gene was a 500 bp fragment. The presence of this gene was confirmed in all patient and 30 (94%) of the environmental isolates. Distribution of the mentioned genes among E. faecalis strains in this study is shown in Table 2.
| The number of genes | Clinical Isolates | Environmental Isolates |
|---|---|---|
| 4 genes | 47 (81) | 25 (78) |
| 3 genes | 9 (16) | 6 (19) |
| 2 genes | 2 (3) | 0 (0) |
| 1 gene | 0 (0) | 0 (0) |
| None of the genes | 0 (0) | 1 (3) |
| Total | 58 (100) | 32 (100) |
a The values are presented as No. (%).
4.4. Results of Biofilm Formation Assay by the Quantitative Method
Quantitative assay of biofilm formation was conducted three times for each strain. Amongst the 58 studied patient isolates 36, 6 and 16 isolates were strongly adherent, weakly adherent and non-adherent, respectively. Amongst 32 environmental isolates, 23, 4 and 5 were strongly adherent, weakly adherent and non-adherent.
4.5. Results of Antibiotic Sensitivity Test and Biofilm Formation Assay
The antibiotic resistance and the ability of biofilm formation of all 90 isolates of E. faecalis were examined. The samples were divided to two groups; biofilm positive isolates with strongly and weakly adherent and biofilm negative isolates with non-adherent strains. The results are shown in Table 3.
| Antibiotics | Clinical Isolates | Environmental Isolates | ||
|---|---|---|---|---|
| Biofilm Positive | Biofilm Negative | Biofilm Positive | Biofilm Negative | |
| Vancomycin | 5 | 19 | 0 | 0 |
| Teicoplanin | 0 | 13 | 0 | 0 |
| Linezolid | 10 | 6 | 0 | 0 |
| Tetracycline | 95 | 62 | 81 | 40 |
| Erythromycin | 67 | 50 | 48 | 20 |
| Cotrimoxazole | 76 | 50 | 26 | 20 |
| Ampicillin | 0 | 31 | 0 | 0 |
| Ciprofloxacin | 63 | 63 | 26 | 0 |
| Gentamicin | 44 | 44 | 30 | 0 |
a The values are presented as percent.
4.6. Results of Restriction Fragment Length Polymorphism Analysis of ebp A, B and C Gene Fragments
Out of 90 isolates, 72 had all four genes. Twenty-three including 15 patient and 8 environmental isolates were selected and polymorphism in their ebp gene by RFLP was examined (Figure 2). The results showed no difference in the obtained fragment patterns in all analyzed isolates.
5. Discussion
In this study, strains of E. faecalis isolated from patient and environmental specimens were analyzed. The patient isolates showed higher antibiotic resistance in comparison to environmental isolates, which could be due to higher transferability of antibiotic resistant genes amongst the patient isolates than environmental isolates. More than 60% of the patient isolates were resistant to tetracycline, erythromycin, co-trimoxazole, ciprofloxacin and gentamycin. On the other hand, the environmental isolates showed resistance only against tetracycline. Furthermore, 5% of the patient isolates were resistant to vancomycin (VRE), whereas none showed this resistance amongst environmental isolates. Other investigators have found various E. faecalis resistance rates to vancomycin, up to 11% in Europe (7), or 17% in isolates from hospital staff hands in Nigeria (12). In this study only 7% (5) of the patient isolates were sensitive to all tested antibiotic agents. Out of all E. faecalis examined, 43 were resistant to three or more antibiotics. Isolation of strains with identical antibiotic resistant patterns indicates widespread dissemination of resistance genes among strains as well as distribution of a common clone in different hospital wards.
In accordance with other reports (13, 14), we also found that biofilm formation was higher in isolates with antibiotic resistance to vancomycin, teicoplanin and ampicillin (as shown in Table 3). This may suggest a genetic linkage between the biofilm and these antibiotic genes. This conclusion was, however, not inclusive for all antibiotics. Quantitative assessment of biofilm formation exhibited that 72% and 85% of all patient and environmental isolates could produce biofilms, respectively. This finding was similar to reports by other investigators. In Japan 62.5% (4), Spain 62% (6) and in USA 79% (9) of the E. faecalis clinical samples were found to form biofilms.
Other studies showed a connection between genes and biofilm existence (15). The presence of ebp gene has been shown to be variable. In the present study, 86% of the isolates carried all four genes, which is lower than previous reports (16, 17). This difference could be due to the fact that there is variable genetic make-up amongst the isolates within the ebp gene. Restriction Fragment Length Polymorphism analysis of our amplified product, however, revealed no such difference which suggests complete gene sequencing is required. The lack of variants in RFLP analysis may indicate the dissemination of a single E. faecalis clone within the hospital setting, which may have gone under slight single nucleotide mutations.
In order to identify the effects of ebp genes in biofilm formation by E. faecalis, the ability of biofilm formation of isolates having four ebp and srtC genes were compared with isolates in which the presence of one or more genes had not been confirmed. Eighty-one percent of patient isolates having four genes could produce biofilms. On the other hand, 61% of strains not possessing one or two of the genes did not form biofilms. For environmental isolates the two possibilities mentioned above were 48% and 14%, respectively. It can be concluded that ebp genes play an essential role in biofilm formation, since the absence of one or two genes have a negative role in production of biofilms. The study of Nallapareddy on E. faecalis OG1RF revealed that mutation in ebp A-srt considerably reduced the ability of biofilm formation.
In conclusion the high antibiotic resistance seen in this study may indicate the intrinsic resistance of enterococci; circulation of transposable elements carrying resistant genes in clinical isolates, indiscriminate and uncontrolled usage of antibiotics and the presence of biofilm are reasons for higher antibiotic resistance among the patient isolates when compared to environmental isolates.