1. Background
Staphylococcus aureus is a common bacterial pathogen that causes significant mortality and morbidity. A wide spectrum of S. aureus-related infection manifestations exists, ranging from mild infections such as pyoderma to more serious and lethal diseases such as osteomyelitis, necrotizing pneumonia, and infective endocarditis. Nasal carriage of S. aureus strains, including both methicillin-resistant Staphylococcus aureus (MRSA) and multidrug-resistant S. aureus, plays an important role in the pathogenesis of staphylococcal infections, detection and treatment of which might be an important modality in the prevention of infections (1, 2). A diversity of virulence factors work together to create the pathogenicity of S. aureus. These factors include cell surface components that promote adherence to surfaces (e.g., protein A, fibronectin-binding and collagen-binding proteins), and exoproteins that invade or bypass the immune system and are toxic to the host (e.g., enterotoxins, exfoliatins, and Panton-Valentine leukocidin [PVL]) (3).
One major obstacle for the treatment of S. aureus infections is the development of antibiotic resistance in the isolates. This resistance phenomenon originated with penicillin, the first broad-spectrum antibiotic, which was discovered in the 1940s (4). Just a couple of years after introduction of methicillin to battle penicillin-resistant strains, MRSA isolates arose (5). Methicillin-resistant Staphylococcus aureus strains were originally healthcare-associated (HA-MRSA). Community-acquired MRSAs (CA-MRSAs) started being reported in the mid-1990s in individuals with limited or no healthcare-associated risk factors (6) and were shown to have a distinct origin from HA-MRSA (7). In fact, CA-MRSA strains have evolved from the more prevalent methicillin-susceptible Staphylococcus aureus (MSSA) strains in the community (8). Both HA-MRSA and CA-MRSA have the mecA gene, which leads to methicillin resistance. The gene is on a genetic element called the staphylococcal cassette chromosome mec (SCC mec) (9). HA-MRSA mainly has SCC mec types I, II, III, and rarely IV, and has no PVL-encoding genes (9, 10). In contrast, CA-MRSA isolates often have an integrated bacteriophage (phiSLT) carrying the PVL-encoding genes, and are mostly of the SCC mec types IV and V (11).
Panton-Valentine leukocidin is encoded by a bi-cystronic operon with the lukS-PV and lukF-PV genes that account for the high virulence potential of CA-MRSA (12). Panton-Valentine leukocidin is a bi-component pore-forming cytotoxin which destroys leukocytes by creating pores in the mitochondrial membrane (13). Recent lines of evidence suggest that PVL might also boost virulence indirectly by inducing expression of other virulence factors (14). Ever since CA-MRSA strains were reported in Canada (15), laboratories have been making an effort to efficiently identify these strains to improve their surveillance, infection control, and treatment (16). Detection of the lukF-PV and lukS-PV genes by conventional PCR was followed by real-time PCR (17, 18), which significantly reduced the turnaround time.
The prevalence of CA-MRSA differs among communities. PVL-positive CA-MRSA is mostly associated with skin infections but also, to a lesser extent, with necrotizing pneumonia (19). Based on the fact that CA-MRSA has a distinctive antibiotic resistance profile (15), special measures should be taken by clinicians in the regions with a higher prevalence of these bacteria to identify them. For example, empirical antibiotic therapy could be followed (20). On the other hand, for a subset of infections such as those of soft tissues caused by CA-MRSA, wound drainage alone may be the choice instead of antibiotics (21). Thus, knowing the prevalence of CA-MRSA on a regional basis is of high importance. The PVL locus representing both a virulence factor and a stable genetic marker of CA-MRSA useful for molecular diagnosis (8).
2. Objectives
The present study was launched to evaluate a real-time PCR assay for rapid and specific identification of PVL-positive MRSA strains for epidemiological purpose in Shahrekord City, Iran and to determine their susceptibility patterns using phenotypic methods. The TaqMan PCR method was used for the detection of PVL-encoding genes and the amplification of mecA (for detection of methicillin resistance) and nuc (for identification of S. aureus) genes (14).
3. Materials and Methods
3.1. Bacterial Isolates and Bacteriologic Methods
In total, 284 Staphylococcus isolates were collected from Hajar and Kashani, the two main Shahrekord University Hospitals. The isolates were selected randomly from routine clinical specimens such as deep and superficial wounds, blood, tracheal, urine, CSF, and venous catheter. No two isolates were collected from the same patient. Staphylococcus isolates were identified based on colonial morphology on blood agar plates (Merck, Germany), Gram stain characteristics, and the catalase test (22). The S. aureus isolates were identified with catalase, coagulase, mannitol fermentations, and DNase tests.
3.2. DNA Extraction
The Promega Wizard MagneSil bead kit (Promega, USA) was used for the extraction of DNA from bacteria, following the instructions provided by the company. The method takes less than 30 minutes.
3.3. Real-Time PCR for Detection of nuc, mecA, and PVL-Encoding Genes
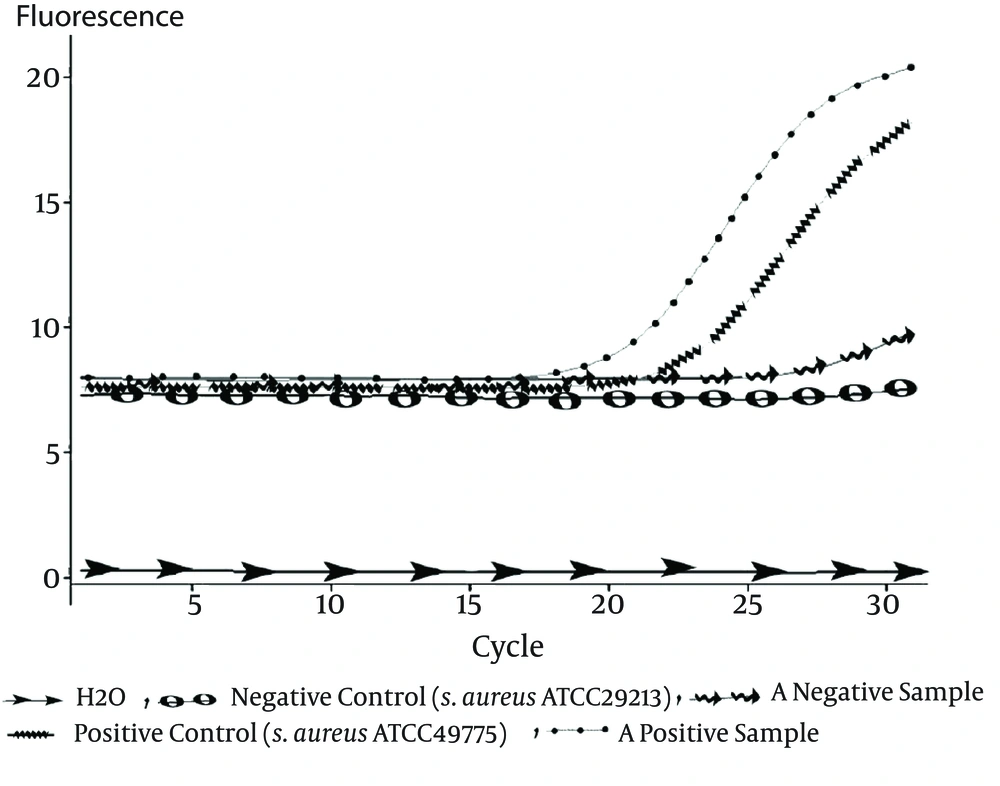
A TaqMan real-time PCR technique in a Rotor Gene 3000 real-time PCR system (Qiagen- Netherlands) was developed to detect nuc (the gene which identifies S. aureus), mecA for identifying methicillin-resistant S. aureus strains (MRSA), and PVL-encoding genes. The primers and probes used in this study are listed in Table 1 (14). Each PCR reaction mixture (12.5 μL) contained 6.25 μL of 2X master mix. Ampliqon, USA), 0.5 μL of each of the primers (10 PM. Methabion, Germany), 0.5 μL of the mecA probe (5 PM). Methabion, Germany), 1 μL MgCl2 (50 mM. Cinnagene, Iran), 1.25 μL H2O, and 2.5 μL DNA. Thermal cycling was performed under the following conditions: 2 minutes at 95°C, followed by 30 cycles of 95°C for 30 seconds and 58°C for 1 minute (23). Reference strains for negative and positive controls are listed in Table 2.
| Primer or Probe Name Reaction Concn, pM | Sequence (5´ → 3´) | 5´Reporter Dye | Reaction |
|---|---|---|---|
| nuc | |||
| nuc For | CAAAGCATCAAAAAGGTGTAGAGA | NA | 10 |
| nuc Rev | TTCAATTTTCTTTGCATTTTCTACCA | NA | 10 |
| nuc Probe | TTTTCGTAAATGCACTTGCTTCAGGACCA | FAMα | 5 |
| mecA | |||
| mecA For | GGCAATATTACCGCACCTCA | NA | 10 |
| mecA Rev | GTCTGCCACTTTCTCCTTGT | NA | 10 |
| mecA Probe | AGATCTTATGCAAACTTAATTGGCAAATCC | FAMα | 5 |
| PVL | |||
| PVL For | ACACACTATGGCAATAGTTATTT | NA | 10 |
| PVL Rev | AAAGCAATGCAATTGATGTA | NA | 10 |
| PVL Probe | ATTTGTAAACAGAAATTACACAGTTAAATATGA | FAMα | 5 |
aα Reporter dye quenched with 3- TAMRA quencher.
| Reference Strains | Positive | Negative |
|---|---|---|
| nucgene | S. aureus [ATCC 43300] | S. epidermidis [ATCC 12228] |
| mecA gene | S. aureus [ATCC 43300] | S. aureus [ATCC 29213] |
| PVL encoding genes | S. aureus [ATCC49775] | S. aureus [ATCC 29213] |
3.4. Antimicrobial Susceptibility Testing
Susceptibility to a range of antimicrobial agents was determined by the disk diffusion method (Kirby-Bauer) for 21 PVL-positive MRSA isolates (24). The following antibiotics were used: oxacillin (1 μg), ciprofloxacin (5 μg), ampicillin (10 μg), clindamycin (2 μg), cefazolin (30 μg), trimethoprim/sulfamethoxazole (1.25/23.75 μg), cefoxitin (30 μg), penicillin (10 U), tetracycline (30 μg), gentamicin (10 μg), erythromycin (15 μg), and ofloxacin (5 μg) disks (MAST Diagnostics, Merseyside, U.K.).
As the gold standard for the detection of vancomycin resistance, the E-test method was used to determine vancomycin minimum inhibitory concentrations (MICs) (25). Muller-Hinton agar plates (Merck, Germany), supplemented with 0.85% NaCl, were inoculated by streaking the standardized inoculums (bacterial suspension with 0.5 McFarland standard) with a sterile swab. Vancomycin E-test strips (AB Biodisk, Solna, Sweden) were placed on the plates, followed by an incubation at 37°C for 16 - 20 hours in ambient air. The MIC for each isolate was read at the intersection point of the zone of growth inhibition with the graduated strip (vancomycin: resistant: ≥ 32 μg/mL; intermediate: 8 μg/mL < MIC < 16 μg/mL; and susceptible: ≤ 4 μg/mL) Enterococcus faecalis ATCC 29212 and Enterococcus faecalis A256 were used as the vancomycin susceptible control and resistant control, respectively.
3.5. Statistical Analysis
Results were analyzed using SPSS statistical software version 13 (SPSS Inc., SPSS/PC+, Chicago, IL, USA). We used the Chi-square test or Fisher exact test to determine the associations of PVL positivity with the presence of the mecA gene and the type of infection. A P value of less than 0.05 was considered to be statistically significant. The risk ratio with a 95% confidence interval (CI) was calculated by comparing the risks of PVL positivity in isolates with and without each type of the studied infection.
4. Results
4.1. Detection of Staphylococcus Isolates
In total, 284 Staphylococcus isolates selected from clinical samples were identified based on colonial morphology on blood agar plates, Gram stain characteristics, and the catalase test.
4.2. Detection of nuc Gene Among Referred Isolates
Out of the 284 isolates, 196 (69%) were positive for the presence of the nuc gene.
4.3. Frequency of mecA Gene Among Referred Isolates
Of the 196 isolates of S. aureus, 96 (49%) were mecA positive (MRSA strains) and 100 were susceptible to methicillin, with no presence of the mecA gene (MSSA strains).
4.4. Frequency of PVL Genes Among Referred Isolates
Of the 100 MSSA strains, 3 (3%) contained PVL genes; of the 96 MRSA isolates, 18 (18.8%) were positive for PVL genes. Thus, the highest prevalence of S. aureus carrying PVL-encoding genes was found within the MRSA strains. In this study, there was a significant relation between the presence of the mecA gene and that of PVL-encoding genes (P = 0.001) (Figure 1).
4.5. Distribution of PVL Genes in Staphylococcal Disease
Of the 196 strains of S. aureus isolated from different types of staphylococcal infections, 21 (10.7%) were PVL positive, 10 (47.6%) of which were associated with tracheal samples (pneumonia) (risk ratio: 6.21; 95% CI: 2.94 - 13.11; P = 0.00027). PVL genes were not detected in isolates associated with eye infection, nasal swab, urine infection, and intravenous catheter (Table 3).
| No. of Strains | PVL-Positivea | Risk Ratio (95% CI)b | P Value | |
|---|---|---|---|---|
| Blood | 35 | 2 (5.7) | 0.48 (0.12, 1.98) | 0.38 |
| CSF | 13 | 2(15.4) | 1.48 (0.38, 5.68) | 0.63 |
| Wound Infection | 54 | 4 (7.4) | 0.62 (0.21, 1.75) | 0.26 |
| Eye Infection | 6 | 0 | 0 | 1 |
| Tracheal | 25 | 10 (40) | 6.21 (2.94, 13.11) | 0.000027 |
| Urine | 16 | 0 | 0 | 0.22 |
| Swab Nasal | 17 | 0 | 0 | 0.22 |
| Urinary Catheter | 5 | 1 (20) | 1.91 (0.31, 11.57) | 0.43 |
| intravenous catheter | 5 | 0 | 0 | 1 |
| Peritoneal Fluid | 20 | 2 (10) | 0.92 (0.23, 3.68) | 1 |
| Total | 196 | 21 (10.7) | 0 | 0 |
aData are presented as No. (%).
bRisk ratio is the ratio of the risk of PVL positivity in the presence of a particular type of infection to the absence of that type of infection.
4.6. Antimicrobial Susceptibility Patterns
The antimicrobial susceptibility patterns differed among the PVL-positive isolates. Eighteen (85.7%) were resistant to oxacillin, which were confirmed by the presence of the mecA gene. One isolate was phenotypically oxacillin sensitive but carried the mecA gene. The number of isolates resistant to different antibiotics was obtained as follows: 11 isolates (52.38%) were resistant to ciprofloxacin, 16 (76.19%) resistant to ampicillin, 13 (61.9%) resistant to clindamycin, 14 (66.66%) resistant to cefazolin, 13 (61.9%) resistant to trimethoprim/sulfamethoxazole, 14 (66.66%) resistant to cefoxitin, 21 (100%) resistant to penicillin, 15 (71.42%) resistant to tetracycline, 11 (52.38%) resistant to gentamicin, 10 (47.61%) resistant to erythromycin, and 11 (52.38%) resistant to ofloxacin. The MICs of the 21 PVL-positive MRSA strains against vancomycin showed that all of the S. aureus strains were susceptible to vancomycin (MIC ≤ 4 μg/mL). Eight PVL-positive MRSA strains were resistant to all of the tested antibiotics in this study except vancomycin.
5. Discussion
Panton-Valentine leukocidin-positive S. aureus isolates producing leukocidal toxins are frequently recovered from deep skin and soft tissue infections, such as cutaneous abscesses and severe necrotizing pneumonia, suggesting that PVL is a major virulence factor. In addition, PVL is mostly associated with CA-MRSA infections. Panton-Valentine leukocidin expression enhances MRSA pathogenicity and is a critical determinant in the choice of suitable antibiotics. Therefore, investigating the prevalence of the PVL marker among MRSA strains, which are a major health issue, is of high importance (6, 26). In the present study, a series of samples collected from the clinical setting were examined by TaqMan real-time PCR in order to identify PVL-positive S. aureus isolates. The results of this study showed that only a small proportion (10.7%) of isolates harbored the PVL-encoding genes. These findings resemble those obtained by related studies performed in other parts of the world (3, 27). To explain this observation, it has been suggested that only some of the S. aureus strains are vulnerable to infection by PVL-converting phages. This hypothesis has been verified by several studies including one that indicated that the bacteriophage SLT infected only 3% of clinical PVL-negative S. aureus strains to produce PVL-positive strains (28). In addition, it has been shown that different strains of S. aureus have different PVL-carrying phages (8).
Our results showed that out of the 100 MSSA strains, 3 (3%) carried PVL-encoding genes. The prevalence data for some studies have been 26%, 16.4%, 27.3%, 12%, and 14% in Nepal, Algeria, Bangladesh, Greece, and Romania, respectively (29-33). Out of the 96 MRSA isolates in our study, 18 were luk-PV positive. Thus, the prevalence of PVL among MRSA isolates is 18.8% in this geographical region. Previous studies in Iran have reported the prevalence to be 7.23% in Ahvaz, Southwest (34), 5.47% in Shiraz, South (35), and 24.16% in Tehran, capital of Iran (36). The prevalence has been reported differently in different regions: under 20% in France, UK, Austria, and Turkey (3, 27, 37), 20% - 50% in Romania, Nepal, Canada, and Greece (14, 31-33), and over 50% in Tunisian, Texas, and Australia (38-40). These differences, of course, may also reflect the type of assay used for detecting the genes.
Our results showed a significant difference between MRSA and MSSA populations in term of carrying the PVL locus. As expected, luk-PV genes are more likely to be present among mecA positive MRSA strains than mecA negative ones (8). In contrast, in a study in Bangladesh, luk-PV genes were found with greater frequency among the MSSA strains (30). In this study, PVL-positive Staphylococcus strains were mostly methicillin-resistant (85.7%), all were susceptible to vancomycin, and the majority of the isolates were resistant to beta-lactam antibiotics. Similar results have been obtained in two other studies (3, 6). When we categorized the isolates according to the type of infection, a statistically significant association was found between PVL genes and pneumonia. However, no such association could be observed for the isolates from eye infection, urine, intravenous catheter, nasal carriers, urinary catheter, CSF, peritoneal fluid, and blood (Table 3). The results of our study substantiate and extend previous findings that S. aureus strains isolated from in-patients affected by necrotizing pneumonia, which led to death in most cases, are mostly positive for the PVL-encoding genes (41). Lung infections also showed a high prevalence of these genes in this study. By contrast, other studies have reported higher frequency of PVL-positive S. aureus in other types of infections. In one study, out of 172 S. aureus isolates, selected among samples referred to the French reference centre for Staphylococcus Toxaemia during a period of 4 years, PVL genes were mostly detected in skin infection-related S. aureus strains (93% and 55% of furunculosis and cellulitis strains, respectively) (16). PVL-producing S. aureus isolates were mostly associated with necrotizing skin infections at a hospital in France (42, 43).
Given the fact that the prevalence of CA-MRSA infections and resultant mortalities is globally increasing (17, 44), applying simple and rapid methods of screening for the identification of PVL-containing CA-MRSA isolates of S. aureus seems to be crucial as the first essential step toward controlling the spread of the pathogen. The present study investigated the prevalence of PVL-positive MRSA in the region of Shahrekord City, Iran. So far, several teams have reported successful real-time PCR assays for the detection of the PVL genes, either alone or in combination with other marker genes including mecA, spa, or nuc (14, 17, 18). Real-time PCR facilitates monitoring of the reaction and there is no need for post-PCR processing, which saves resources and time. Real-time PCR assays are well-suited for diagnostic purposes as they are easy to perform, have high sensitivity and greater specificity, and provide an opportunity for automation (14).
In conclusion, the prevalence of PVL-containing MRSA isolates, found to be 18.8% in this study, warrants further detailed scrutiny to prevent possible future endemics in the studied hospitals as well as other hospitals in the region.