1. Background
Helicobacter pylori is a Gram-negative, curved, microaerophilic bacillus, which is colonized in the human gastric mucosa and induces chronic infection. Helicobacter pylori infection is accompanied with atrophic gastritis, peptic and duodenal ulcers, gastric adenocarcinoma, and gastric mucosa-associated lymphoid tissue (MALT) lymphoma (1, 2). Helicobacter pylori was recently defined as a group I carcinogen (3). According to statistics, 60% of the world’s population is infected with H. pylori, ranging from 20-30% in developed countries to 70% - 80% in developing countries (4-7).
Cytotoxin-associated gene A (CagA) protein is produced in most H. pylori strains and is recognized as an immunogenic protein. In fact, CagA, as a 120 - 145 kDa protein, is able to induce multiple alterations in cellular signaling pathways, leading to the designation of CagA as a “bacterial oncoprotein” (8). Overall, H. pylori CagA+ strains have been associated with gastric ulcers, duodenal ulcers, and gastric cancer (9).
Helicobacter pylori infection can be identified through invasive and non-invasive methods. Serological tests are non-invasive assays, performed on clinical samples. Serological detection of H. pylori infection via anti-CagA enzyme-linked immunosorbent assay (ELISA) and CagA immunoblotting is the only available non-invasive test for determining the strain virulence potential and possible risk of disease. Considering the availability of various serological assays, reliability of CagA serology as a predictive test for determining the CagA genotype of infecting strain is of great significance (10).
2. Objectives
Diagnosis of H. pylori antigen in the stool is a non-invasive method, which may be used for the detection of H. pylori. In this regard, Farjadi et al. identified the antigenic region of CagA (arCagA) protein through bioinformatics and produced a recombinant protein. The results showed the remarkable antigenic properties of the purified protein. With this background in mind, the aim of the present study was to evaluate an assay using arCagA gene for the detection of H. pylori infection in stool samples.
3. Methods
3.1. Clinical Samples
In total, 110 adult dyspeptic patients within the age range of 21 - 78 years (mean age: 50 years), who were referred to Vali-Asr hospital (a teaching hospital in Arak, Iran) from April to November 2012, were included in the study; the subjects had not received any particular treatment for H. pylori infection. Stool samples were collected from the participants and stored at -20°C. Helicobacter pylori infection was confirmed, using the ELISA kit for the measurement of CagA antigen in stool samples (Dimeditec, Germany), according to the manufacturer’s instructions. This study was approved by the ethics committee of Arak University of Medical Sciences (code: 91-138-12).
3.2. Cloning of H. pylori arCagA Gene
All DNA manipulations were carried out according to the standard protocols, described by Sambrook (11). The region of antigenic proteins was determined, based on the results reported by Farjadi et al. (12) (Accession No.: Fj798973) (12). The arCagA gene was amplified from the H. pylori genome via polymerase chain reaction (PCR) method, using the following primers: (5′-TATGGATCCACAATAACGCTC-3′) as the forward primer and (5′-ATTCTCGAGTTCATCAAAAGATT-3′) as the reverse primer. The primers contained a BamHI endonuclease site at the 5´ end of the forward primer and a XhoI endonuclease site at the 5′end of the reverse primer. After PCR purification, the PCR product was inserted into the pET32 cloning vector (Novagen, USA).
3.3. Expression and Purification of the Recombinant CagA Protein
Escherichia coli BL21 (DE3) pLysS was transformed into pET32a-arCagA and grown in a nutrient broth (Merck, Germany), supplemented with ampicillin (100 μg/mL) and chloramphenicol (35 μg/mL) at 37°C with agitation. For the expression of the recombinant protein, 500 μL of the culture was added to 50 mL of the nutrient broth [per liter containing 10 g of the yeast extract (Difco, USA), 20 g of Bacto tryptone broth (Difco, USA), 0.2% (mass/vol) glucose, 10 g of NaCl, 1 g of KCl, 0.5 g of MgCl2, 0.5 g of CaCl2, 100 μg/mL of ampicillin, and 35 μg/mL of chloramphenicol] and incubated at 37°C (200 rpm); then, it was shaken with vigorous agitation until the optical density reached 0.6 at 600 nm.
Isopropyl β-D-1-thiogalactopyranoside (IPTG) with a concentration of 1 mM was added to induce the expression of CagA protein for 4 hours. The expressed protein was purified via Ni-NTA agarose resin affinity chromatography, according to the manufacturer's instructions (QIAGEN, USA). The quality and quantity of the purified recombinant arCagA protein were analyzed by sodium dodecyl sulphate-polyacrylamide gel electrophoresis (SDS-PAGE 12%) and Bradford methods, respectively (13).
3.4. Production of Mouse Anti-Serum Against arCagA Protein
Six rats were used for the production of anti-arCagA antibody. The purified protein was injected into the rats at three doses. In the first dose, 30 µg of protein, along with complete Freund’s adjuvant, was subcutaneously injected. The second and third doses of the protein (30 µg) were injected, along with incomplete Freund’s adjuvant. Finally, the animal’s blood was collected and its serum was separated and kept at -20°C. Before immunization, 500 μL of blood was collected and used as the negative control.
3.5. Western Blot Analysis
The Western blot assay was used to examine the presence of CagA protein in the stool samples, collected from patients with H. pylori infection. The Laemmli method was applied for SDS-PAGE of the fecal samples (14). Then, the protein bands were transferred to a polyvinylidene difluoride (PVDF) membrane. After blockage with 3% bovine serum albumin (BSA) solution, PVDF membrane was incubated with a primary antibody (mouse anti-arCagA) for 1 hour.
After reaction with the primary antibody, the membranes were washed three times with tris-buffered saline (TBS) and incubated with horseradish peroxidase-conjugated goat anti-mouse IgG (Bioscience, USA). After rinsing, the membrane was exposed to diaminobenzidine (DAB; Sigma, USA) solution. The membrane was washed after the appearance of the desired band and stored in a dark place (15). The sensitivity and specificity of the Western blot technique for the detection of CagA protein were determined in the stool samples of patients with H. pylori infection (95% confidence interval).
4. Results
4.1. Cloning of arCagA Gene
The 1245 bp fragment of the gene, produced through PCR amplification, was cloned in the pET32a vector. The sequencing of the cloned fragment showed that its nucleotide sequence is the same as the desired sequence with antigenic properties.
4.2. Production and Purification of arCagA Protein
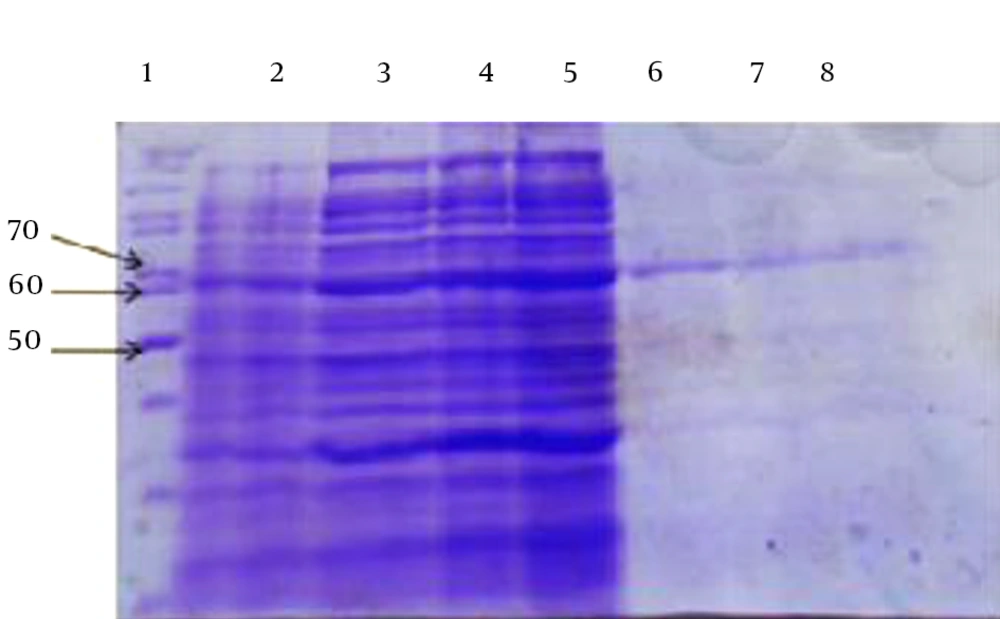
Production of the protein was initiated by adding IPTG after transferring the recombinant vectors into E. coli BL21 (DE3) pLysS competent cells; the produced protein weighed 65 kDa. After bulk culturing of the bacteria, cellular precipitation was disrupted by sonication technique and centrifuged. Then, the precipitation and supernatant were examined through SDS-PAGE to detect the recombinant protein.
The main part of the desired protein was placed as the inclusion body in the precipitation. The histidine tag (6 × His. tag) at the beginning of the protein was purified, using nickel resin (Figure 1). Then, the protein was dialyzed and the extract appeared as a single band in the desired region of the gel. The concentration of the produced recombinant protein was 1.5 mg/mL for arCagA.
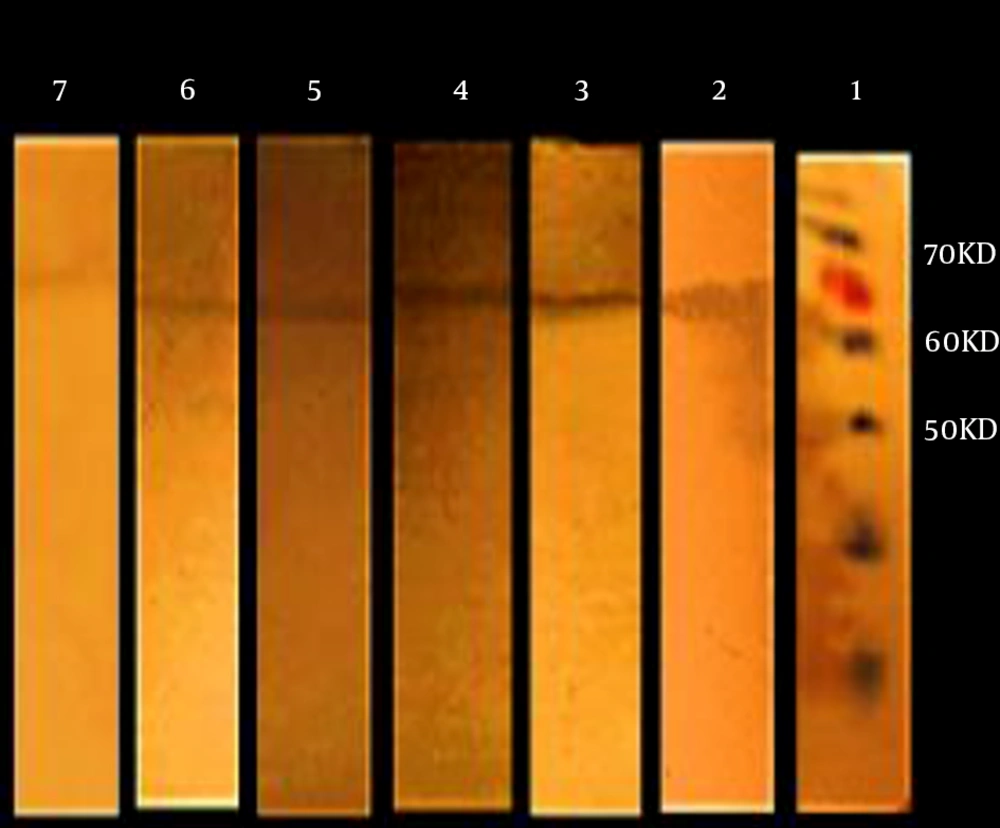
4.3. Western Blot Analysis
Among 113 patients evaluated in the present study, 80 were positive for the new recombinant arCagA antigen, while 33 were negative, based on the results obtained by the commercial ELISA CagA kit. In the present study, 70 cases were positive for the new recombinant arCagA antigen, while 10 were false negative. On the other hand, in the control group, three samples were false positive for arCagA recombinant protein. Figure 2 shows the specific interaction between the samples, and Figure 3 demonstrates normal human samples. The sensitivity and specificity of Western blotting were calculated to be 87.5% and 90%, respectively, as shown in Table 1. The measurements are based on the literature, using a 2 × 2 table (16).
| Tests | Positive | Negative | Sensitivity, (%) | Specificity, (%) |
|---|---|---|---|---|
| Total No. | 80 | |||
| Positive (conventional test) | 80 | 0 | 100 | |
| Positive (present study) | 70 | 10 | 87.5 | |
| Negative (conventional test) | 0 | 33 | 100 | |
| Negative (present study) | 3 | 30 | 90 |
5. Discussion
Helicobacter pylori infection can be identified through invasive and non-invasive methods. Selection of a proper method for each patient depends on the cost, availability, infection distribution, and the patient’s place of residence and antibiotic use; currently, non-invasive tests are preferred. According to the literature, stool antigen test is an available, cost-effective method, which seems to be as precise as the urea breath test.
Regardless of previous investigations and treatments, the present method, which seems to be as effective as urea breathe test, can be recommended for the detection of active infections with less potential false negative results (17-21). In addition, it could be a substitute for invasive methods in older patients for whom endoscopy may be intolerable. The high global prevalence of H. pylori infection and its certain role in the development of gastric malignancy have encouraged researchers to study different antigens of this bacterium and investigate the possibility of their application as diagnostic indicators and/or vaccines. Overall, different antigens of this bacterium, such as VacA, NAP, Fla, and urease, have been studied for the design of vaccines and diagnostic kits (22-27).
Sheikhian et al. (28) (2004) extracted a 26 kDa protein, currently known as alkyl hydroperoxide reductase C (AhpC) enzyme, from H. pylori. This protein was found in the blotting pattern of all infected patients and was introduced as a diagnostic tool. Overall, production and purification of proteins are more cost-effective than natural proteins in slow growing, fastidious bacteria such as H. pylori (29). Moreover, Pourakbari et al. (30) recently produced AhpC recombinant protein from E. coli and revealed that the anti-AhpC antibody can be utilized for the detection of this protein in the stool samples of patients. The sensitivity and specificity of the test were 83.3% and 91.7%, respectively, which represent an acceptable performance for the detection of H. pylori infection in children and adults.
Use of the whole recombinant protein, containing the full length of the gene, increases the molecular weight of the produced protein due to the presence of unnecessary epitopes. It also encumbers the process of cloning, expression, and purification and reduces the production of the recombinant protein. Logically, decreased production of the whole protein is not desirable, which is why in most studies, antigenic segments of the protein are selected and produced instead of the entire protein. In a study by Farjadi et al., 12 antigenic regions of H. pylori CagA protein were determined, using bioinformatics and then cloned. It was shown that the antigenicity of the produced protein is similar to the whole CagA protein in patients with H. pylori infection. Therefore, in this study, we evaluated the possibility of using the arCagA of H. pylori for the detection of CagA in the stool samples of patients with H. pylori infection through Western blotting method.
According to the results of the present study, the antibody against antigenic regions of CagA protein (arCagA) can detect the natural form of the protein in stool samples; sensitivity of 87.5% and specificity of 90% highlight this finding. According to the obtained results, this protein exhibits proper antigenic strength and possesses epitopes, similar to its natural form; as a result, it can be used for the design of detection kits.
In the present study, the false negative results were only reported in three samples, gathered from the patients. The reasons for false negative results can be the consumption of proton pump inhibitors or bismuth, watery diarrhea, possible increased use of plant materials in the diet (resulting in stool weight increment), and presence of polysaccharide inhibitors. In addition, the false negative results may arise from problems confronted in western blot technique, such as reduced concentration of the primary antibody, decreased antigen level, proteolytic cleavage, antigen inactivation, and increased transfer time and temperature.
In summary, the results of the present study showed that the antibody against arCagA protein can identify the natural form of the protein in the stool specimens of patients with H. pylori infection. Therefore, the produced recombinant protein has proper antigenicity, and therefore, it can be used as a serological tool for the detection of H. pylori infection and development of diagnostic kits.