1. Background
Staphylococcus aureus is one of the well known pathogens that can live in a wide variety of environments. It also has an inherent ability to form biofilms on biotic and a-biotic surfaces (1). Moreover, bacteremia caused by S. aureus and CoNS is a serious infection associated with high morbidity and mortality and refers to the development of resistance virtually to all antibiotics including vancomycin. The capacity to cause infection is probably due to the organism’s capacity to colonize and survive in host during infection process (2).
Staphylococcus aureus is a major cause of illness and death and imposes serious economic costs on patients and hospitals. It can infect a wide variety of human tissues, resulting in different clinical appearances that vary in severity from slight purulent infections such as impetigo to more serious conditions like infective endocarditis (3). The rates of S. aureus infections, both in community and hospital-acquired strains, are increasing steadily. Treatment of these infections is becoming more difficult because of the increasing prevalence of oxacillin and multidrug-resistant S. aureus isolates (4).
Methicillin-resistant S. aureus (MRSA) is proposed as a main pathogen since it is resistant to many antibiotics and responsible for nosocomial infections worldwide. It is alarming that S. aureus strains carrying resistance genes have been isolated from various samples (5, 6). There are rising reports regarding CoNS as a cause of disease in immunosuppressive patients and increased prevalence of multiple drug resistant strains (7). Disinfectants and antiseptics are widely used in hospitals and other health care centers as a type of topical and hard-surface practices. Especially, they are a necessary portion of infection control programs and help prevent nosocomial infections (8, 9). The quaternary ammonium compounds (qacs) and small multidrug-resistance (smr) genes are responsible for resistance to antiseptics (10). On the other hand, one of the resistance mechanisms to disinfectants and antiseptics in S. aureus is mediated by two gene families named qac and smr (11, 12).
Three determinants including qacA/B, qacC, and smr genes cause resistance to organic cations by means of multidrug efflux pumps dependent on the proton motive force. The qacA gene causes resistance to some different organic cations, including benzalkonium, ethidium bromide, cetrimide, and Chlorhexidine. This gen is located both on chromosome and pSK1 plasmid family. The qacB gene gives resistance mostly to monovalent organic cations and to a lower extent to certain divalent compounds. It is located on pSK23 plasmid. The qacA gene is closely related to the qacB gen (5, 6). Resistance to ethidium bromide and quaternary ammonium compounds can be acquired by the qacC gene. It is usually located on conjugative plasmids in S. aureus and other Staphylococci clinical isolates (13, 14).
2. Objectives
The objective of the present study was to characterize the antibiotic resistance and also determine the prevalence of QACs genes in Methicillin-sensitive S. aureus (MSSA) and MRSA isolated from nares of nurses, and also in MSCoNS and MRCoNS from clinical samples. The association between the presence of mecA gene and antiseptic resistance genes was also investigated in Staphylococci isolates.
3. Methods
3.1. Study Design
In this cross-sectional study, samples were obtained from nurse’s nose and clinical specimens. The study included a total of 120 Staphylococcal strains isolated from Kashani and Hajar teaching hospitals in the southwest of Iran during the years 2012 to 2013. Staphylococcal strains included 60 S. aureus strains isolated from anterior nares of the nurse’s nose, and 60 CoNS strains isolated from clinical specimens (wound, soft tissues, blood cultures, intravascular catheter tips, and midstream urine).
3.2. Bacterial Identification
Each nasal swab was inoculated in a trypticase soy broth (TSB) medium (Merck, Darmstadt, Germany.co) and incubated for 24 hours at 37°C. TSB mediums were subcultured on to mannitol salt agar (Merck, Germany.co) and incubated for 24 hours. Colonies with Staphylococcal morphology were identified by biochemical tests such as catalase and coagulase and confirmed by 16S rRNA genes. Tube coagulase negative strains were reported as CoNS. Staphylococci isolates were stored at -70°C in TSB mediums supplemented with 30% glycerol.
3.3. Susceptibility Testing
Antibiotic susceptibility testing was performed by disk diffusion method according to the CLSI. All S. aureus isolates were tested by oxacillin (1 µg), Gentamycin (10 µg), Ticoplanin (30 µg), Linezolid (30 µg), quinopristin-dalfopristin (15 µg), Tigecycline (15 µg), Rifampin (5 µg), and Vancomycin (30 µg) and all CoNS isolates were examined by Bacitracin (0.4 U), Novobiocin (30 µg), and Vancomycin (30 µg) (MAST Diagnostics, Merseyside, UK).
Methicillin-resistant Staphylococci (MRS) strains were known by multiplication on Muller-Hinton agar and using oxacillin disc (1 µg) diffusion method. The inhibition zone size was interpreted according to the CLSI criteria (15). The MRSA strains were also confirmed through the presence of mecA gene by PCR technique that is so far considered as the gold standard method for the detection of methicillin resistance.
MICs of Chlorhexidine gluconate (Sigma Aldrich, St Louis, USA) (ranging from 1 to 64 μg/mL, by serial twofold dilutions) were determined in triplicate by the broth micro dilution method according to the CLSI guideline (15). The lowest concentration totally inhibiting growth after 24 hours incubation at 37°C was considered the MIC. S. aureus ATCC 13353 and S. aureus ATCC 25923 were used as positive and negative controls, respectively.
3.4. Qac genes Detection
Total genomic DNA of the microorganisms in this study was extracted using boiling method. The qacA/B, qacC, and smr resistance genes were tested in all isolates by multiplex PCR. The qacA and qacB genes were considered together because simple PCRs cannot discriminate between them. To detect qacA/B, qacC, and smr genes, the multiplex PCR conditions were as follows: 1 cycle of initial denaturation at 95°C and 35 cycles of denaturation (at 94°C for 40 seconds), annealing (at 54°C for 50 s), extension (at 72°C for 50 seconds), and one final extension (at 72°C for 5 minutes).
The detection of qacA/B, qacC, and smr genes was performed by multiplex PCR with the following sets of primers: 5′- GCA GAA AGT GCA GAG TTG G-3′ and 5′- CCA GTC CAA TCA TGC CTG -3′ for qacA/B (product size, 361 bp), and 5′- GGC TTT TCA AAA TTT ATA CCA TCC T -3′ and 5′- ATG CGA TGT TCC GAA AAT GT -3′ for qacC (product size, 246 bp), and 5′- GCC ATA AGT ACT GAA GTT ATT GGA-3′ and 5′- GAC TAC GGT TGT TAA GAC TAA ACC T -3′ for smr (product size 195 bp), as previously described (16, 17).
3.5. Statistical Analysis
Statistical analyses were performed using SPSS software version 18.0 for Windows (SPSS Inc., Chicago, IL, USA). The association of categorical variables was determined by Chi Square or Fisher’s exact tests. P < 0.05 was considered statistically significant.
4. Results
Of total 60 S. aureus isolates included in this study, 51 (85%) isolates were MRSA and, of 60 CoNS, 7 (11.66%) isolates were MRCoNS. The frequency of antimicrobial resistance in the isolated MRSA strains was 17 (33%) for rifampin, 15 (29.5%) for Tigecycline, 12 (23.5%) for Ticoplanin, and 7 (13.7%) for Gentamycin. All MRSA isolates showed susceptibility to vancomycin, Linezolid, and quinopristin-dalfopristin. As well as, only 2 isolates (1.3%) of MRCoNS strains were resistant to Vancomycin. The MICs of Chlorhexidine were 1 µg/mL for 42 (35%) isolates, 2 µg/mL for 22 (18.3%) isolates, 4 µg/mL for 40 (33.3%) isolates, 8 µg/mL for 14 (11.6%) isolates, and 16 µg/mL for 2 (1.6%) isolates. The association between the presence of mecA gene and Chlorhexidine MIC is shown in Table 1.
| Species (no. of Isolates) | Chlorhexidine MIC Level, µg/mL | |||||
|---|---|---|---|---|---|---|
| MIC Range 1 - 2 | MIC Range 2 - 4 | MIC Range 4 - 16 | ||||
| Positive | P Value | Positive | P Value | Positive | P Value | |
| MRSA (51) | 1 (1.96) | 0.15 | 34 (66.6) | 0.002 | 16 (31.3) | 0.05 |
| MSSA (9) | 8 (88.8) | 1 (11.11) | 0 | |||
| MRCoNS (7) | 3 (42.8) | < 0.001 | 4 (57.1) | < 0.001 | 0 | NS |
| MSCoNS (53) | 52 (98.1) | 1 (1.88) | 0 | |||
| Total | 64 | 40 | 16 | 120 | ||
Abbreviation: NS, non-significant.
aValues are expressed as No.(%).
The association between the presence of qac genes and a Chlorhexidine MIC of > 4 µg/mL was statistically significant (P < 0.0001). It is considerable that among two strains with the highest MIC of Chlorhexidine gluconate (MIC ≥ 8 µg/mL), one of them was qac negative and the other strain carried qacC and smr genes at the same time. Clinical strains with reduced susceptibility to Chlorhexidine (MIC ≥ 2 µg/mL) had a strong association with the presence of qacA/B and smr determinants (P < .0001). As well as, among the tested bacteria with various Chlorhexidine MICs, there were some strains of Staphylococci isolates that carried the two genes at the same time (Table 2).
| Qac Genes | Frequency (%) | Chlorhexidine MIC Level, µg/mL, % | |||||
|---|---|---|---|---|---|---|---|
| 1 | 2 | 4 | 8 | 16 | P Value | ||
| qacA/B | 6 (5) | 2 (33.3) | 1 (16.7) | 3(50) | 0 | 0 | |
| qacC | 8 (6.7) | 1 (12.5) | 0 | 7 (87.5) | 0 | 0 | |
| smr | 19 (15.8) | 1 (5.2) | 1 (5.2) | 17 (89.4) | 0 | 0 | |
| qacA/B + smr | 5 (4.2) | 0 | 0 | 5 (100) | 0 | 0 | < 0.0001 |
| qacA/B + qacC | 4 (3.3) | 0 | 0 | 2 (50) | 2 (50) | 0 | |
| qacC + smr | 14 (11.7) | 0 | 0 | 1 (7.15) | 12 (85.7) | 1 (7.15) | |
| Negative | 64 (53.3) | 38 (59.4) | 20 (31.2) | 5 (7.8) | 0 | 1 (1.56) | |
| Total | 120 (100) | 42 (35) | 22 (18.3) | 40 (33.35) | 14 (11.7) | 2 (1.6) | |
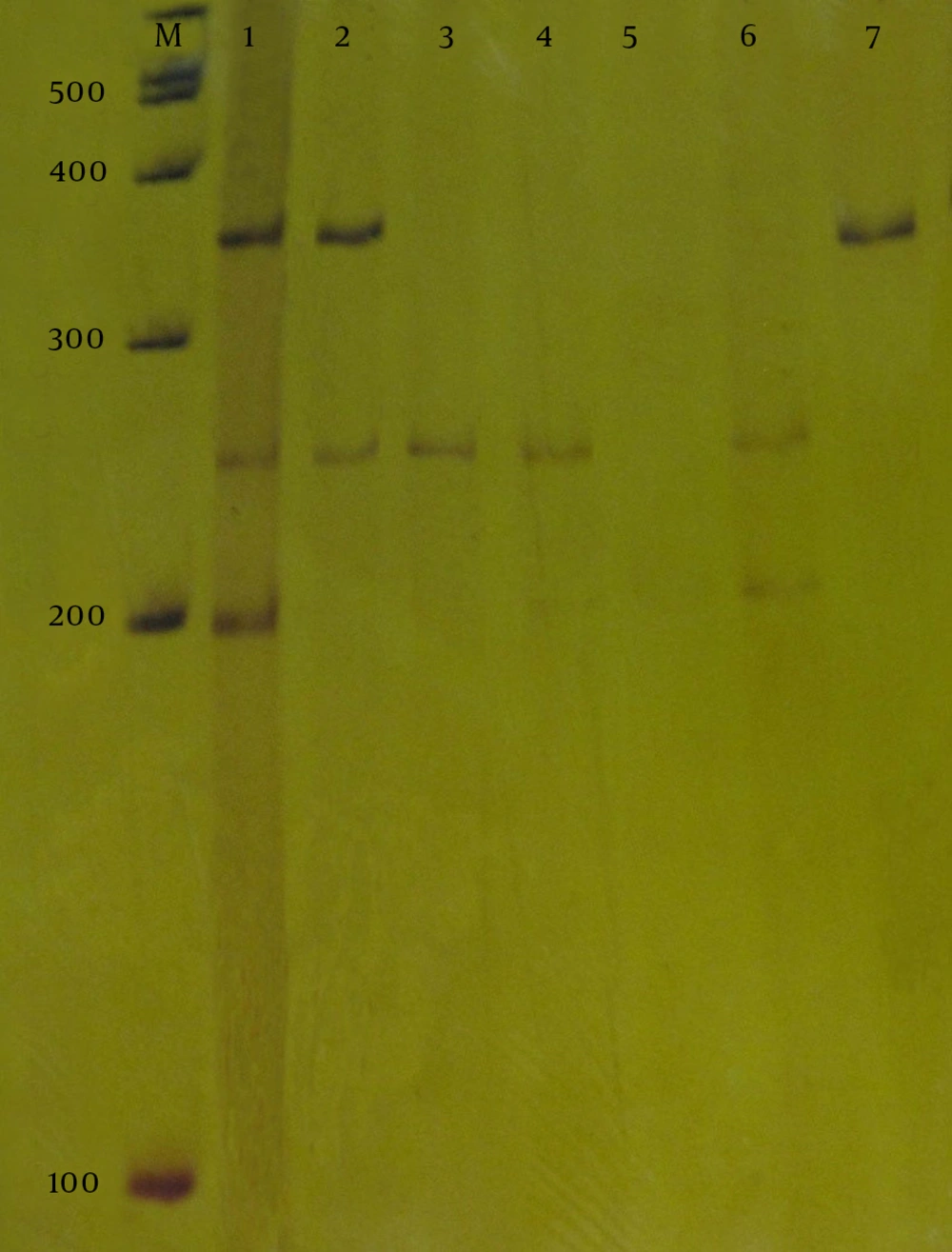
The qacA/B gen was identified in 15 (12.5%) isolates from which, 6 (40%) were qacA/B alone, 5 (33.3%) with both smr and qacA/B, and 4 (26.7%) with both qacA/B and qacC at the same time. The qacC was identified in 26 (21.7%) isolates from which, 8 (30.8%) were qacC alone, 4 (15.4%) plus qacA/B, and 14 (53.8%) plus smr at the same time. The smr was identified in 38 (31.7%) isolates from which, 19 (50%) smr alone, 5 (13.1%) plus qacA/B, and 14 (36.9%) plus qacC at the same time (Figure 1). Also, 41 out of 120 (34.1%) isolated strains were qac-negative. Isolates with qac genes (N = 56) had significantly a higher mean and wider range of Chlorhexidine MICs. The results of this study also showed that methicillin resistant Staphylococci were more often resistant to Chlorhexidine than methicillin sensitive isolates (Table 3).
Line M ladder 100 bp (Fermentas), Line 1 positive control (S. aureus ATCC 13353), line 2 isolates harboring qacA/B (361 bp) and qacC (247 bp), line 3 and 4 isolates harboring qacC (247 bp), line 5 isolates negative for qacs gene, line 6 isolates harboring smr (195 bp) and qacC (247 bp), line 7 isolates harboring qacA/B (361 bp).
| Species (No. of Isolates) | qacA/B | qacC | smr | |||||||||
|---|---|---|---|---|---|---|---|---|---|---|---|---|
| Positive | P Value | OR | 95% CI | Positive | P Value | OR | 95% CI | Positive | P Value | OR | 95% CI | |
| MRSA (51) | 4 (7.8) | 0.38 | 0.92 | 0.85 - 0.99 | 7 (13.7) | 0.83 | 1.2 | 0.13 - 11 | 17 (33.3) | 0.041 | 0.66 | 0.54 - 0.8 |
| MSSA (9) | 0 | 1 (11.11) | 0 | |||||||||
| MRCoNS (7) | 1 (14.28) | 0.086 | 8.6 | 0.47 - 157 | 0 | NS | NS | 2 (28.57) | < 0.001 | 0.71 | 0.44 - 1.1 | |
| MSCoNS (53) | 1 (1.88) | 0 | 0 | |||||||||
Abbreviations: CI, confidence interval; NS, non-significant; OR, odds ratio.
aValues are expressed as No.(%).
5. Discussion
Staphylococcus aureus is a flexible pathogen, which has been known as a common cause of various community-acquired and nosocomial infections (18). Hospital staff nasal carriers of MRSA can increase the risk of outbreaks of MRSA at Hospitals because they are potential reservoirs for the spread of MRSA strains among patients. Nasal colonization of MRSA and MRCoNS is different all over the world. For example, a Korean study reported a rate of MRCoNS colonization of non-healthcare workers (13%) that is similar to the rate found in our study (19). The unnecessary use of disinfectants at hospitals results in the permanence of Staphylococcal isolates, as well as, the augmented prevalence of qac genes in such strains (20).
In order to evaluate the MIC of Chlorhexidine, determine the presence of qac genes, and investigate the association between the presence of mecA gene and Chlorhexidine MIC, 60 isolates of S. aureus and 60 isolates of CoNS were studied. The Chlorhexidine MIC defined in this study was equal to or higher than 4 μg/mL. According to previous studies (16-18), the Chlorhexidine MIC for MRSA strains was typically equal to or higher than 4 μg/mL. Therefore, our finding was in accordance with those of the above-mentioned studies. The correlation between the presence of qac genes (qacA/B, qacC, and smr) and a Chlorhexidine MIC of ≥ 4 µg/mL was statistically significant (P ≤ 0.0001). It is noteworthy that 2 (33.3%) out of 6 strains with qacA/B gene had Chlorhexidine MIC = 1 µg/mL. As well as 1 (12.5%) out of 8 strains with qacC and 1 (5.2%) out of 19 strains with smr genes had Chlorhexidine MIC = 1 µg/mL. The presence of smr gene had a strong relationship with the presence of mecA gene in S. aureus (p ≤ 0.041) and CoNS (P ≤ 0.001). The only one out of 14 qacC plus smr-carrying strains showed a MIC value of 16 µg/mL. It is necessary to mention that among 64 strains which did not have qac genes, one strain also showed an MIC value of 16 µg/mL. In fact, we observed a significant increase in Chlorhexidine MICs associated with the qacA/B, qac, and smr genes. In a Canadian study on MRSA isolates, there was no significant increase in MICs with the qac and smr genes (21); while our data showed an increase in the MIC value of Chlorhexidine in the presence of qacA/B, qacC, smr, and mecA genes. The increased MIC values for Chlorhexidine (MIC of > 2 µg/ml) had a strong relationship with the presence of mecA in S. aureus (P ≤ 0.002) and CoNS (p ≤ 0.001). In summary, our data showed that 85% of MRSA and 11.6% of MRCoNS isolates carried qac genes, which were in association with reduced susceptibility to Chlorhexidine.
The QACs resistant genes such as qacA/B, qacC, and smr have been isolated in various environments from clinical CoNS isolates and different Staphylococcal species (13, 16). Whereas the current antiseptic MICs related to qac gene positive Staphylococci will not allow survival at in-use concentrations, even moderately increased resistance may allow persistence when residual disinfectants are present (16, 22). The over-use of Chlorhexidine as a decontaminant could result in the appearance of MRSA isolates carrying qacA/B resistance genes (23). While hospital environments act as a source of qac genes, long-term care facilities and nurses may play a contributory role in the transmission of antiseptic-resistant Staphylococci between hospital sections and patients (24). Many studies have been performed in order to understand the incidence and possible genetic link of antiseptic and antibiotic resistance genes in Staphylococci. Similar to the findings of other studies (20, 25), in our study the existence of mecA gene in these isolates was related to the presence of qac genes. Our study demonstrated that the raised ratio of qac gene positive Staphylococci strains offers the co-selection of these genes due to the increased contact with MRSA-infected patients. Decreased susceptibility to antiseptic (Chlorhexidine) could be associated with qac genes, which is also consistent with the reports of clinical isolates in above studies.
In this study, the frequency of qacA/B, qacC, and smr genes was 7.8%, 13.7%, and 33.3%, respectively while in the study of Noguchi et al. in Japan (26), the frequency of qac genes was 45.9% that is higher than that in our study; but the frequency of smr gene was 5.3% that is much lower than the value obtained in our study. In another study by Miyazaki et al. conducted on 74 Brazilian MRSA isolates (27), the rate of qacA/B was that higher than the ratio in our study. Lee et al. in Switzerland identified qacA/B in 91% of the MRSA isolates (28). In a study by Longtin et al. in Canada (21), only 2% of the MRSA strains possessed the qacA/B while smr gene was detected in 7% of strains. In Iran, a study by Hasanvand et al. (29) showed that the frequency of qacA/B antiseptic gene in MRSA isolates was 9% that is in agreement with our study. On the other hand, the smr gene in the same study was not detected in both MRSA and MSSA isolates while the frequency of the smr gene in this study was 17 (33.3%) in MRSA and 2 (28.57%) in MRCoNS isolates. The mecA gene is located on a mobile genomic island Staphylococcal cassette chromosome (SCC), which not only serves as a vehicle for the genetic exchange of genes among Staphylococci, but also as a carrier for virulence and additional drug-resistant genes (17, 30). Genetic linkage between qac genes and mecA genes conferring resistance to methicillin on the same Staphylococcal plasmids has also been reported elsewhere (31).
Application of antiseptics might be chosen for strains resistant to antibiotics and help them maintain in healthcare settings. The presence of an association between mecA and qacA/B, qacC, and smr may promote survival of MRSA and MRCoNS in the hospital environments. These Staphylococci may be a hazard for infection control because of their persistence in places with low amounts of antiseptic residues. In conclusion, our study showed Chlorhexidine resistance is commonly found in MRSA isolates from nurse’s nose and clinical specimens. This study had limitations such as coordination problems in nasal swab sampling from staff and clinical specimens found in different units of hospitals.
The increase of antiseptic-resistant bacteria is one of the most significant problems and a serious threat to public health. Understanding the selection of gene transfer that causes the distribution of resistance genes is very important for long-term strategies in order to treat microbial diseases.
5.1. Ethical Considerations
Ethical issues (including plagiarism, informed consent, misconduct, data fabrication and/or falsification, double publication and/or submission, redundancy, etc.) have been completely observed by the authors.