1. Background
The increasing clinical frequency of antibiotic-resistant pathogens isolating from hospitals is one of the important health care issues all over the world. Studying of resistance determinants with molecular and genetic methods have a critical role in understanding, controlling, and spreading of resistant pathogens. Aminoglycosides are the most frequently used antibiotic agents in the treatment of infections by both Gram-negative and Gram-positive bacteria. This group of antibiotics acts by binding to the ribosomes of bacteria and in turn interferes with protein synthesis (1-3).
The main strategy of resistance against aminoglycosides in clinical bacterial isolates is the production of modifying enzymes, such as Aminoglycoside Phosphoryltransferase (APH), Aminoglycoside Acetyltransferase (AAC), and Aminoglycoside Nucleotidyltransferase (ANT) (4, 5). The APHs and ANTs are the bisubstrate enzymes, which facilitate the transfer of the γ-phosphate and nucleotide monophosphate, respectively, from a nucleotide substrate to the hydroxyl groups of aminoglycoside antibiotics, while AACs acetylate amino groups derived from Acetyl Coenzyme A (acetyl-CoA) (6). Moreover, aac (6′)-I confers resistance to tobramycin and amikacin, aac (6′)-II and ant (2″)-I inactivate tobramycin and gentamicin, and amikacin is the substrate of aph (3′)-VI (7).
The 16S rRNA methylase confers high-level resistance to 4, 6-substituted deoxystreptamines, including, arbekacin, amikacin, kanamycin, tobramycin and gentamicin, by posttranscriptional methylation of 16S rRNA leading to loss of affinity for aminoglycosides (8). Despite the existence of more than 85 aminoglycoside-modifying enzymes, only a limited number of them, ANT(2′′)-I, AAC(6′)-I, AAC(3)-I, AAC(3)-II, AAC(3)-III, AAC(3)-IV and AAC(3)-VI, appear to have been undergoing selection process and causing the majority of aminoglycoside resistance (9).
2. Objectives
In different studies, the phenotypic and genotypic resistance of Enterococcus and Salmonella isolates against aminoglycosides has been reported from Iran and all over the world (10-17). This study aimed to investigate the occurrence of aminoglycoside resistance and the prevalence of the resistance-modifying enzyme genes, acc(3)-Ia, aac(6′)-Ib, acc(3)-IIa, armA and rat in Enterococcus and Salmonella strains isolated from two hospitals in Iran (Imam Khomeni , Ilam and Milad, Tehran, Respectively).
3. Materials and Methods
All of the culture media were purchased from Himedia laboratories (India), and Polymerase Chain Reaction (PCR) materials were also purchased from CinnaGen Company (Iran). Gentamicin (10 µmol/L), amikacin (30 µmol/L), kanamycin (30 µmol/L), and tobramycin (10 µmol/L) were obtained from Himedia, and netilmicin (30 µmol/L) was obtained from Mast laboratories (Mast Diagnostic, Merseyside, UK).
3.1. Bacterial Isolates
Enterococcus and Salmonella isolates were collected from Imam Khomeni and Milad hospitals of Ilam and Tehran, Respectively) during a 6-month period (2010). Antibiotic susceptibility was determined by the disk diffusion method. Moreover, antimicrobial susceptibility was determined using the disk diffusion method (Kirby-bauer) against gentamicin (10 µmol/L), amikacin (30 µmol/L), kanamycin (30 µmol/L), and tobramycin (10 µmol/L), according to the guidelines of the Clinical and Laboratory Standards Institute (CLSI).
3.2. Polymerase Chain Reaction Studies
Detection of aminoglycoside resistance genes, including acc(3)-Ia, aac(6′)-Ib, acc(3)-IIa, and 16S rRNA methylase (armA and rat) were detected by PCR. A fresh bacterial colony was suspended in 100 µL of sterile distilled water and boiled at 100ºC for 10 minutes. After centrifugation, 3 mL of supernatant was used for PCR assays with the primers described in Table 1. Amplification of DNA was performed in a thermal cycler (Eppendrof, Germany). Polymerase chain reaction elongation times and temperature conditions were indicated in Tables 1 and 2. Moreover, PCR products were electrophoresed in 1.5% agarose gels and visualized under UV light. The PCR product of 16S rRNA methylase was then purified with the QIA quick PCR purification kit (QIAGEN, Inc., Chatsworth, CA, USA) and sequenced on both strands with an ABI PRISM 3100 (Applied Biosystems) genetic analyzer (18-20). The sequences were compared with those in GenBank nucleotide database.
| Genes | Primer Sequence | M. W, bp | Reference |
|---|---|---|---|
| aac (3)-Ia | 484 | This study | |
| Forward | 5′-ATGGGCATC ATTCGCACA-3′ | ||
| Reverse | 5′-TCTCGGCTTGAACGAATTGT-3′ | ||
| acc (6′)-Ib | 524 | This study | |
| Forward | 5′-ATGACTGAGCATGACCTTG-3′ | ||
| Reverse | 5′-AAG GGT TAG GCA ACA CTG-3′ | ||
| aac (3)-IIa | 749 | This study | |
| Forward | 5′-CGGAAGGCAATAACGGAG- 3′ | ||
| Reverse | 5′-TCGAACAGGTAGCACTGAG -3′ | ||
| armA | 591 | AY220558 (1) | |
| Forward | 5′-AGGTTGTTTCCATTTCTGAG-3′ | ||
| Reverse | 5′-TCTCTTCCATTCCCTTCTCC-3′ | ||
| rat | 585 | AB103506 (1) | |
| Forward | 5′-CCCAAACAGACCGTAGAGGC-3′ | ||
| Reverse | 5′-CTCAAACTCGGCGGGCAAGC-3′ |
a All of genes manufactured by CinnaGen.
| Cycle Counts | Temperature and Time |
|---|---|
| 1 | |
| Primary denaturation | 94ºC, 5 min |
| 35 | |
| Denaturation | 94ºC, 45 s |
| Annealing | |
| aac(3)-Ia | 55ºC, 30 s |
| aac(6′)-Ib | 53ºC, 30 s |
| aac(3)-IIa | 55ºC, 30 s |
| armA | 53ºC, 30 s |
| rat | 56ºC, 30 s |
| Primary extension | 72ºC, 1 min |
| 1 | |
| Final extension | 72ºC, 10 min |
4. Results
From a total of 140 and 79 isolates of Enterococcus and Salmonella collected presumptively, 117 and 77 isolates were identified and included as Enterococcus and Salmonella, respectively. Isolates were preserved in Tripticase Soy Broth (TSB) containing 20% glycerol. The isolates were obtained with high percentages from specimens of urine, ulcer and sputum, with 87.2%, 6%, and 3.4%, respectively, after that 0.9% of the positive specimens were taken from the catheter, blood, ascites, and abscess (Table 3). Moreover, 46.2% (n = 54) and 53.8% (n = 63) of Enterococcus were from male and females, respectively. Ninety-three out of 117 strains of Enterococcus were resistant to aminoglycosides. Resistant strains of Enterococcus have the highest prevalence of resistance against both kanamycin and amikacin (68.4% and 80% of the cases, respectively), and after that tobramycin with 42.7% (50 cases) and gentamicin with 28.2% (33 cases), respectively. Salmonella isolates have the highest rate of resistance against kanamycin (6.9%), and minor percentages of resistance against other 3 antibiotics. Ninety-three isolates of Enterococcus and 26 isolates of Salmonella at least were resistant against one of the aminoglycosides using the disk diffusion method.
4.1. Polymerase Chain Reaction for Detection of Aminoglycoside Resistance Genes
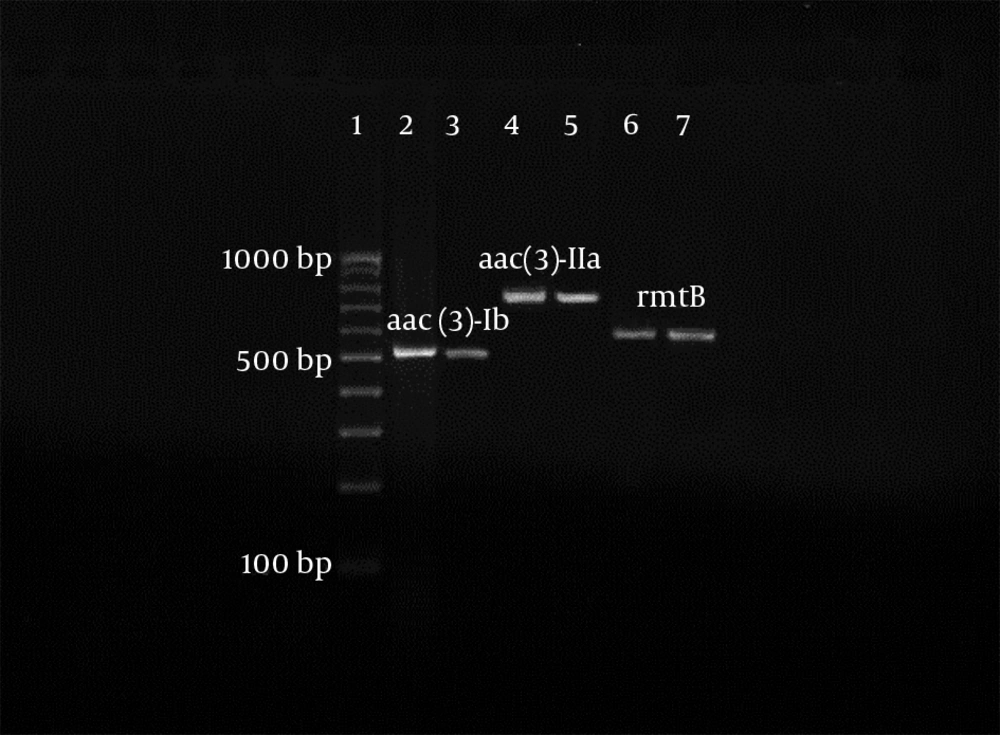
All of 119 of both Enterococcus and Salmonella isolates, which were resistant to aminoglycosides in disk diffusion were tested for the detection of acc(3)-Ia, aac(6′)-Ib, acc(3)-IIa, 16S rRNA methylase (armA and rat). Also, 72.04% (67), 66.7% (62), and 36.6% (34) of the resistant strains of Enterococcus had the aac(3′)-Ia, aac(3′)-IIa, and aac(6′)-Ib genes, respectively. All of the resistant isolates of Enterococcus were negative for both armA and rat genes, and none of the Salmonella isolates have the studied aminoglycoside genes, because of this fact other data relating to Salmonella isolates were not reported in later sections of article. Mono and multi-aminoglycoside gene resistance patterns were indicated in Enterococcus isolates in Table 4. Furthermore, a sample of electrophoresis profile of the PCR products of aminoglycoside resistant enterococci was indicated in Figure 1.
| Antibiotic Resistance Phenotype | Percentage of Resistance | Gene | |||||
|---|---|---|---|---|---|---|---|
| aac (3)-Ia | aac (3)-IIa | aac (6′)-Ib | IIa and Ib | armA | rat | ||
| GM | 33 (28.2) | 21 (63.6) | 25 (75.8) | 8 (25.2) | 7 (21.2) | 0 | 0 |
| K | 80 (68.4) | 59 (73.8) | 53 (66.2) | 30 (37.5) | 17 (21.2) | 0 | 0 |
| AK | 80 (68.4) | 53 (66.2) | 56 (70) | 22 (27.5) | 15 (18.8) | 0 | 0 |
| TOB | 50 (42.7) | 32 (64) | 40 (80) | 14 (28) | 12 (24) | 0 | 0 |
| GM and K | 33 (28.2) | 21 (63.6) | 25 (75.8) | 8 (24.2) | 7 (21.2) | 0 | 0 |
| GM and AK | 33 (28.2) | 21 (63.6) | 25 (75.8) | 8 (24.2) | 7 (21.2) | 0 | 0 |
| GM and TOB | 33 (28.2) | 21 (63.6) | 25 (75.8) | 8 (24.2) | 7 (21.2) | 0 | 0 |
| GM and K and AK | 33 (28.2) | 21 (63.6) | 25 (75.8) | 8 (24.2) | 7 (21.2) | 0 | 0 |
| GM & K & AK & TOB | 33 (28.2) | 21 (63.6) | 25 (75.8) | 8 (24.2) | 7 (21.2) | 0 | 0 |
| K and AK and TOB | 46 (39.3) | 30 (65.2) | 36 (78.3) | 12 (26.1) | 10 (21.7) | 0 | 0 |
| AK and TOB | 48 (41.3) | 31 (64.6) | 38 (79.2) | 13 (27.1) | 11 (22.9) | 0 | 0 |
| K and TOB | 47 (40.2) | 30 (63.8) | 37 (78.7) | 12 (25.5) | 10 (21.3) | ||
| K and AK | 68 (58.1) | 48 (70.6) | 48 (70.6) | 21 (30.9) | 14 (20.6) | 0 | 0 |
aAbbreviations: GM, gentamicin; K, kanamycin; AK, amikacin; TOB, tobramycin.
bPercentage of resistant strains and percentage of genes have been calculated considering total count of isolates and total resistant strains, respectively.
cAll of the mono and multi-aminoglycoside gene resistance patterns in Salmonella isolates were zero or without any result.
5. Discussion
There is no doubt that screening of resistance genes are very useful tools in epidemiologic tracing of genes between bacterial species, prediction of future outbreaks in different regions, treatment strategies and many other subjects in the treatment and control of bacterial diseases. Based on internet resources, we conducted first report of aminoglycoside resistance genes from Iranian clinical isolates of Enterococcus strains. We could not found 16sRNA modifying gene in studied isolates of Enterococcus and Salmonella. Enzymatic modification of aminoglycosides including phosphorylation, adenylation and acetylation of amine and hydroxyl groups are from most common methods of antibiotic resistance in bacteria (21).
Among aminoglycoside modifying enzymes, aac (6')-Ib gene is more common and leads to tobramycin, kanamycin and amikacin resistance. This gene firstly identified in Klebsiella pneumonia isolates in 1986 (22). Although aminoglycoside modifying enzymes are the most common and major methods of resistance, 16S rRNA methylase-producing Gram-negative bacteria have been increasingly reported in recent years (23). This type of resistance is mediated by plasmids and is a new resistance mechanism in Enterobacteriaceae and non-fermentative Gram-negative bacilli. Comprehensive study of resistance patterns of isolates from different resources including environment, animals, and humans could help us to direct our treatment and control policies of infectious diseases, considering this point more than 95% of Escherichia coli and enterococci isolates from common buzzards in one study showed that they are resistant to common antimicrobials such as streptomycin, tetracycline, ampicillin, chloramphenicol, gentamicin, kanamycin, etc. Because of the absence of direct antibiotic pressure in the world, this fact clearly indicates direct distribution of resistance genes from human life to other environments, which in turn helps to recycle such problems (24).
The witness for this claim is the frequent studies in which the presence of resistance, high-level aminoglycoside resistance, and other antibiotic resistance genes have been indicated in human, animals and food resources (25-28). Overall, there are scarce reports on aminoglycoside resistance genes in Salmonella species (13). There are little studies on resistance patterns of Enterococcus from Iran, and many of them have been focused on vancomycin and High-Level Gentamicin Resistant (HLGR) (14, 15, 29); in one of the studies, the aac(6')-Ie-aph(2")-Ia gene were identified in 83% of Enterococcus faecalis and 100% of E. faecium HLGR isolates, respectively (29). Moniri et al. (16) in their study showed that aac (6′)/aph (2") and esp, genes were found in E. fecalis isolates 68.4%, and 61.1%, respectively. They proposed a meaningful relationship between biofilm formation and possession of Enterococcal Surface Protein (ESP) and aac (6′)/aph (2") genes. Vaziri et al. (30) in their study of the prevalence of aminoglycoside-modifying enzyme genes (aac (6')-I, aac (6')-II, ant (20)-I, aph (3')-VI) in 250 clinical Pseudomonas aeruginosa isolates found that aac (6')-II (36%) was the most frequently identified gene in phenotypic resistant isolate. Because of widespread distribution of aminoglycoside resistance, related common genes with new resistance mechanisms were evaluated in this study.
This study aimed to determine the prevalence of aminoglycoside resistance genes (acc (3)-Ia, aac (6′), acc (3)-IIaIb, armA, rat) in Enterococcus and Salmonella isolates from clinical samples of Milad (a 1000 bed Tehran hospital) and Ilam hospitals. Totally, 117 and 77 isolates of Enterococcus and Salmonella were collected and studied for the presence of mentioned genes. RmtB and ArmA genes are not identified in our studied isolates, these enzymes are belonging to a group of resistance genes calling 16S rRNA-methylases (16S-RMTases) that are plasmid-mediated and until now 5 main types of ArmA, RmtA, RmtB, RmtC, and RmtD have been identified all over the world. RmtB has been reported from various Gram-negative bacterial species including K. pneumoniae, and K. oxytoca, isolated in Japan, South Korea, and Taiwan. Until now, 16S-RMTases have been reported from at least 30 countries or regions, and worldwide distribution of these genes is considered in some studies as an alerting hint to health-care officials (31), In china E. coli clinical isolates were reported highly resistant to gentamicin and etimicin, and the aminoglycoside-modifying enzyme, AAC(3)-II, seems to bea main type of E. coli strains resistance to aminoglycoside agents (32). Results of the current study similar to many other researches emphasizes on implementation of concise programs, including molecular methods from officials toward special regulations on resistance pattern of bacteria in hospital environments especially on apparently noncommon bacteria, such as Salmonella and Enterococci. Results of this study showed that in Enterococcus isolates, acetylation genes had an important role in resistance to aminoglycosides and because of a high frequency of aminoglycoside resistance in clinical Enterococcus isolates, the following suggestions have been recommended to health care professionals: 1- Prescription of aminoglycosides only after antibiogram, or prescription of amikacin, tobramycin and netilmycin in empirical treatment cases. 2- Evaluation of efflux pumps of MexAB-OprM, and aminoglycoside resistance genes in these strains. 3- Annual or 6 month reports and tracking of antibiograms with precise and suitable sensitive tests, especially on hospital acquired infection cases.