1. Background
Pseudomonas aeruginosa is one of the leading antibiotic-resistant bacterium, which plays an important role in the inception of nosocomial infections worldwide (1). In most areas of the world, resistance of P. aeruginosa to β-Lactam compounds is increasing (2). Recently, by carbapenems, infection due to resistant species is controlled better. In recent decades, P. aeruginosa is reported to be resistant against these compounds (3, 4). Although different reasons such as generation of chromosomal Amp-c, loss of purine D2 or efflux pumps can be considered for this resistance, the most important reason is generation of carbapeneme hydrolyzing enzymes (5).
Metallo-beta-lactamase-generative P. aeruginosa species were reported for the first time in Japan in 1991. However, these species are reported worldwide as well as Iran, in which isolation of multidrug-resistant P. aeruginosa species has reached a critical point (6). Metallo-beta-lactamases (MBLs) such as VIM, IMP, and SPM, which belong to group B of Ambler classification, cause hydrolysis of a variety of beta lactams except monobactam (aztreonam) (7). Among beta lactamases, MBLs are unique in requiring the presence of zinc ion in the active site of the enzyme. Their activity is inhibited by chelating agents such as Ethylene Diamine Tetra Acetic Acid (EDTA), Sodium Mercapto Acetic Acid (SMA), and Dipicolinc Acid, but beta lactams inhibitors such as clavulanic acid, sulbactam, and tazobactam have no effects on them (8-10).
Based on epidemiological studies conducted worldwide, it is proved that the prevalence of genes coding MBL in P. aeruginosa species is different in various geographic zones and even in various hospitals (11-13). Therefore, according to the clinical importance of organisms generating MBL, it is necessary to identify and control them in hospitals for therapeutic purposes (13).
2. Objectives
One of the major clinical problems related to P. aeruginosa is attributed to MBL. Given that there was no information about MBL genes in Zahedan hospitals, the current study was designed to investigate Metallo-beta-Lactamase VIM-1, SPM-1, and IMP-1 genes among clinical P. aeruginosa species isolated in Zahedan, Iran by phenotypic (E test) and genotypic (Polymerase Chain Reaction) methods.
3. Materials and Methods
3.1. Bacterial Isolates
Samples included 191 species of P. aeruginosa isolated from Zahedan hospitals (Emam Ali, Boali, Nabi Akram, and Tamin-e-Ejtemaei) from October 2012 to March 2013. The isolates were identified using microbiological and biochemical methods.
3.2. Antibiotic Resistance Investigation
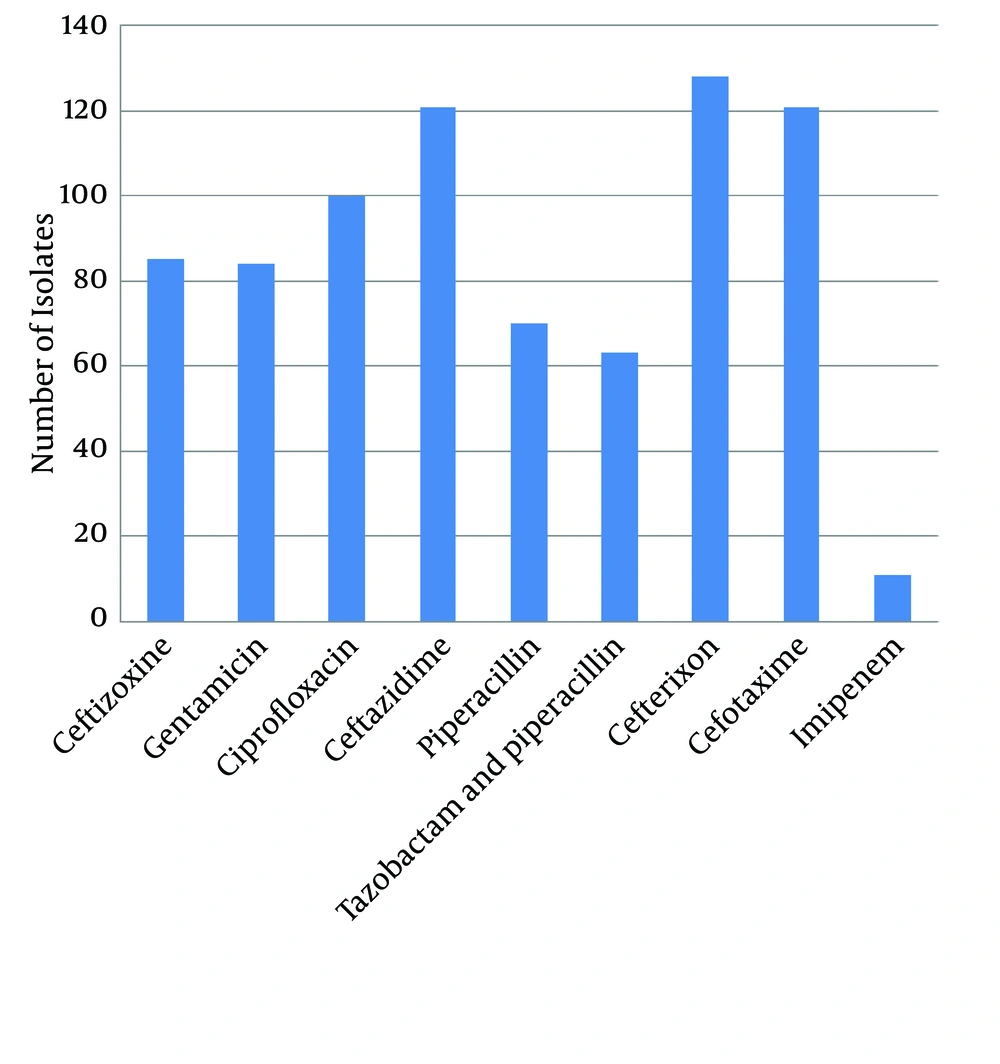
Measurement of antibiotic resistance was conducted by disk diffusion method for the following antibiotics: ceftizoxime (ZOX), gentamicin (GM), ciprofloxacin (CP), ceftazidime (CAZ), piperacillin (PLP), tazobactam and piperacillin (TZP), cefteriaxon (CRO), cefotaxime (CTX), and imipenem (IPM) (Mastdiscs, Britain) according to the standards of Clinical and Laboratory Standards Institute (CLSI).
3.3. Susceptibility Testing
Minimum Inhibitory Concentration (MIC) of imipenem resistant isolates was measured by E-test, according to CLSI protocol. Pseudomonas aeruginosa ATCC 27853 was used as a control reference strain. In order to detect MBL generative isolates, E-test tapes were used (AB Biodisk, Solna, Sweden) according to the manufacturer’s instructions. These tapes have two parts. In one part (IP), there is a gradient of different imipenem ratios (4 - 256 µg/mL) and in the other one (IPI), a gradient of different imipenem ratios plus a constant concentration of EDTA (1 - 64 µg/mL) (10, 14). To interpret E-test tapes, it should be considered that if MIC of IP/IPI ratio is equal or greater than eight, (MBL) enzyme is generated (10).
3.4. Investigation of bla-VIM-1, bla-IMP-1, and bla-SPM-1 Genes by PCR
Imipenem resistant isolates were used to investigate bla-VIM-1, bla-IMP-1, and bla-SPM-1 genes by PCR. For DNA extraction, boiling method was applied. First, three to five colonies were extracted from fresh culture medium and then suspension was prepared using 200 mL of distilled water boiled at 100˚C for 10 minutes. Next, it was centrifuged at 12000 rpm for 10 minutes. The supernatant containing DNA was transferred to new Eppendorf tubes for PCR in order to amplify MBL genes, bla-VIM-1, bla-IMP-1, and bla-SPM-1. The following primers (Sinaclone, Iran) were used to amplify MBL genes (15):
VIM-1:
Forward: 5′-AGTGGTGAGTATCCGACAG-3′
Reverse: 5′-ATGAAAGTGCGTGGAGAC-3′
IMP-1:
Forward: 5′- ACCGCAGCAGAGTCTTTGCC-3′
Reverse: 5′-ACAACCAGTTTTGCCTTACC-3′
SPM-1:
Forward: 5′- GCGTTTTGTTTGTTGCTC -3′
Reverse: 5′-TTGGGGATGTGAGACTAC-3′
To perform PCR, 1 µL of the extracted DNA was added to PCR Master Mix with the final volume of 25 µL containing 1.5 Mm of MgCl2, 0.24 Mm of dNTPs, 5 pm of each primer, and one unit of Taq polymerase enzyme. PCR was controlled by P. aeruginosa species carrying out SPM-1, VIM-1, and IPM-1 genes. In order to control the molecular weight, 100 base pair DNA ladder was applied and electrophoresed on 1% agarose gel and then stained using ethidium bromide. Result was observed by transilluminator under ultraviolet (UV) light.
Amplification reactions were carried out in a 25 µL volume using one unit of Taq polymerase enzyme (Sinaclone, Iran), 20 ng of template DNA, 1.5 Mm of MgCl2 , 0.24 Mm of dNTPs and 5 pm of each primer in the reaction buffer recommended by the enzyme manufacturer. PCR was performed using Taq polymerase kit (Sinaclone, Iran). The products were amplified with 30 cycles in the following program: denaturation at 95°C for 5 minutes, denaturation at 94°C for 30 seconds, annealing at 55°C for 30 seconds, and extension at 72°C for 30 seconds. Pseudomonas aeruginosa isolate generating SPM-1, VIM-1, and IPM-1was used as positive control. PCR product was electrophoresed on 1% agarose gel.
4. Results
Among the 191 isolated species of P. aeruginosa, the maximum and minimum number of isolated species belonged to ICU (43) and CCU (one), respectively. Furthermore, the species were isolated from different clinical samples; most of them (71 species) belonged to mid-stream urine and the least (one specie) to biopsy samples.
4.1. Antibiotic Resistance Pattern
The results of antibiotic resistance pattern, obtained by Kirby-Bauer method on 191 P. aeruginosa species isolated from different areas of Zahedan hospitals for piperacillin, cefotaxime, ceftizoxime, ceftazidime, piperacillin, imipenem, tazobactam, piperacillin, and ciprofloxacin are shown in Figure 1.
4.2. Metallo-bata-Lactamase Producers Detection
The imipenem-nonsusceptible isolates were evaluated by E-test to detect MBLs. Eleven out of 191 isolates were resistant against imipenem. For these resistant species, E-test was applied and nine out of eleven resistant isolates were identified as MBL producers.
4.3. PCR Detection of Metallo-beta-Lactamase Genes
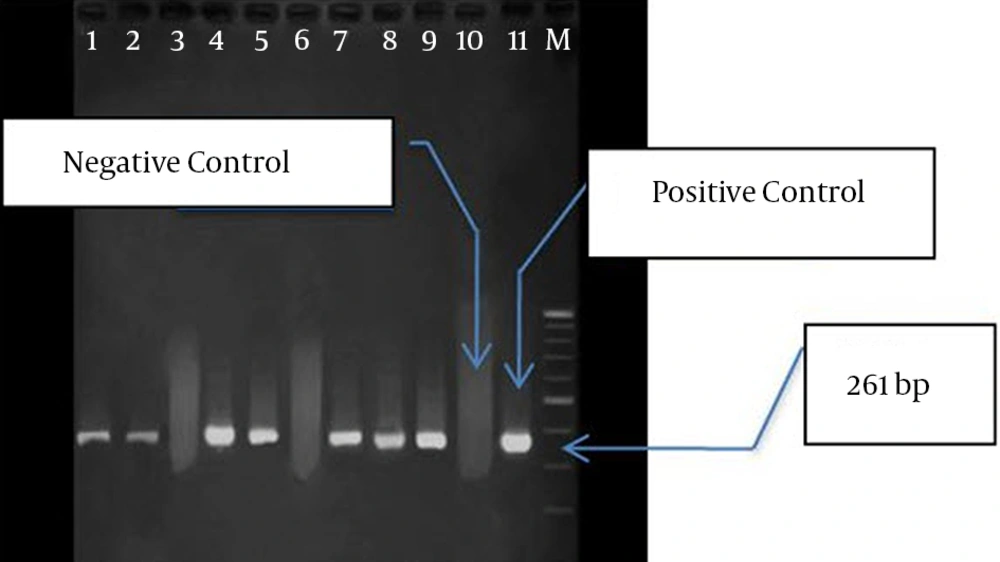
PCR was conducted on nine isolates to investigate the frequency of IPM-1, SPM-1, and VIM-1 genes. In seven isolates, only VIM-1 gene was found, while none of the genes were found in two isolates (Figure 2). Among the isolates containingVIM-1 gene, three samples were isolated from lesions, three from mid-stream urine, and one from endotracheal tube (ETT). Totally, four isolates belonged to ICU and other three isolates to CCU. These seven isolates were resistant against all the studied antibiotics, but they were sensitive to polymyxin B and colistin.
5. Discussion
It is a great challenge to treat infections caused by P. aeruginosa, due to its resistance against drugs. Different antibiotics including beta lactames, aminoglycosides and quinolones are applied to treat the infections caused by P. aeruginosa. Antibiotic resistance pattern of these bacteria (P. aeruginosa) is changing quickly. However, studies showed that the resistance of P. aeruginosa against antibiotics fluctuates, especially regarding imipenem (16). Carbapenems and beta lactam antibiotics are using considerably to treat infections due to multidrug-resistant P. aeruginosa since they are resistant against most of the beta lactamases and have great membrane permeability (17). However, in last decades, warnings about an increase in P. aeruginosa resistance against antimicrobial agents are reported. These organisms have various self-protection ways against antibiotics.
One of the most important ways for resistance against imipeneme is generation of MBL enzymes (18, 19). VIM, IMP, and SPM from MBL family are quite prevalent in P. aeruginosa (20). Since various reports indicate the increased resistance of P. aeruginosa isolates against antibiotics (especially imipeneme), proper use of these antibiotics and time-consuming identification of isolates generating MBL should be considered. It leads to successful treatment and prevents propagation of resistant genes, which can be seriously harmful for societies (21). There are different methods to identify MBL-generative species. The present study used E-test tapes. These tapes contain EDTA, which is a MB inhibitor (22, 23).
In the current study, seven out of nine isolates, which were positive in E-test (phenotypic method), contained VIM-1 gene and two isolates were negative in both phenotypic and genotypic (PCR) methods, indicating the presence of other factors such as lack of opr D, which results in membrane permeability change, efflux pumps or chromosomal Ampc beta lactamase. Two isolates, positive in phenotypic method, were reported negative in PCR reaction indicating that other genes are involved in this process. Among the isolates containing VIM-1 gene, four species (57%) were isolated from the patients hospitalized in ICU for approximately a long time and had undergone antibiotic therapy. It indicates the presence and propagation of the above mentioned gene under such circumstances. In the current investigation, IMP-1 and SMP-1 genes were not observed in any of the strains resistant to imipenem. The study by Shahcheraghi et al. on 243 P. aeruginosa species isolated from Imam Khomeini Hospital in Tehran, showed that 15 isolates contained VIM-1 gene (24). Also, Sepehriseresht et al. reported that 94 out of 483 species of P. aeruginosa isolated from the patient lesions were resistant and intermediate resistant species and carried vim1, vim2, ipm1, and ipm2 genes by PCR (25).
The results of the above mentioned researches are nearly consistent with those of the present study indicating that VIM-1 is the predominant MBL in Iran. Another research showed that 34 out of 111 (43%) species isolated carried VIM gene (6, 26) and their results are not consistent with those of the current study. It can be said that the prevalence of MBL in the species isolated from that hospital was more than expected. Since Imam Musa Kazem Hospital is a burn center, a great amount of imipenem is prescribed there and resistance against this antibiotic has reached 94.9%. Therefore, it is not unexpected to find genes driving drug resistance, especially MBL ones located more on dynamic elements (plasmide or transpozone) are propagated more in this center.
The current study showed that fortunately the prevalence of MBL in Zahedan is less than other cities. However it should be noted that according to the studies in most areas of the world, the prevalence is incremental and if preventive measures are not taken, there will be major clinical problems. Since most of integrones containing VIM-1 gene coding aminoglycosides destructing enzyme, the importance of the problem is intensified (27). Therefore diagnostic methods should be routinely used to determine these species in clinical laboratories; also synthesis of more effective antimicrobial compounds with new effecting mechanisms should be noted.