1. Background
Urinary tract infection (UTI) is one of the most frequent types of infections in community and hospital settings, affecting about half of the world’s population during their lives. Although Escherichia coli and other Enterobacteriacae are the most important causes of UTIs, such infections could be caused by different bacteria (1). Recently, several investigators have evaluated the presence of Gram positive species, such as coagulase negative staphylococci, Staphylococcus aureus, Streptococcus group B, and enterococci, which play a clinically significant role in UTIs. It is well established that S. aureus is the cause of UTI among patients with urinary tract catheterization (2). Urinary tract infection associated with S. aureus causes symptoms that are undistinguishable from those caused by E. coli.
Although the existence of S. aureus in the urine is often secondary to staphylococcal bacteremia originating at other sites (e.g. in cases of endocarditis), it could be an important primary urinary pathogen and causes ascending urinary tract colonization and infection (1, 2). The most important factor contributing to the successful wide distribution of this pathogen is believed to be the remarkable ability to acquire resistance to antimicrobial agents, to the extent that, nowadays, one of the major public health concerns regarding S. aureus infections is the increasing prevalence of methicillin-resistant S. aureus (MRSA) (3-5). Resistance to methicillin results from the mecA gene, which is located on a mobile genetic element with 21 to 67 kbp of size, called staphylococcal cassette chromosome (SCC) element (6, 7).
Based on the literature, although mortality rates associated with UTI is usually low, antimicrobial therapy should not be overlooked. Increased antibiotic resistance has significantly limited the choice of therapeutic options available for treatment of staphylococcal infections and presents a particularly difficult challenge in these contexts (2). Due to rapid dissemination of MRSA and multi drug resistance (MDR), therapeutic options are limited when staphylococcal UTI is encountered; therefore, UTI caused by S. aureus requires special management (1, 2).
Various techniques have been developed to determine the type of S. aureus strains, including multilocus sequence typing (MLST), coa typing, pulsed-field gel electrophoresis (PFGE), staphylococcal cassette chromosome mecA (SCCmec), spa typing, the mec-associated hypervariable region (dru), and the accessory gene regulator (agr) (8-10). SCCmec typing classified SCCmec elements based on their structural differences. So far, 11 different types of SCCmec I-XI have been classified (7). The spa typing, as an effective, cheap, and easy technique for typing S. aureus isolates, is a prevalent typing technique for S. aureus isolates. This method is based on the number of tandem repeats and the sequence variation in region X of the protein A gene (11).
2. Objectives
Given that urinary S. aureus may be the source of staphylococcal bacteremia, information on the distribution of molecular types and antibiotic susceptibility pattern of S. aureus in patients whose urine is persistently colonized with this organism is very important. The present study was conducted to determine SCCmec and spa types of S. aureus strains isolated from patients with UTI. The researchers also assessed susceptibility of S. aureus strains isolated from these patients to the commonly used antibiotics.
3. Methods
3.1. Ethics Statement
The current study was approved by the ethics committee of Shahid Beheshti University of Medical Sciences, Tehran (IR.SBMU. MSP.REC.1396.147).
3.2. Study Design and Collection of Urine Samples
The present study was conducted on 863 urine samples collected from patients complaining of UTI at 5 medical centers between February 2015 and January 2016. The 5 hospitals were located in the northern (A), central (B), southern (C), eastern (D), and western (E) areas of Tehran (capital city of Iran). A total of 90 S. aureus isolates were obtained from urine samples of patients with UTI. The patients were instructed to collect proper urine sample after cleaning the genital area. The first and the last portion of the urine stream were discarded and the mid-stream was collected into an appropriate sterile bottle. Urine samples from children and newborns were obtained using an aseptically placed collection bag. All the urine samples were transported to the laboratory in thermal boxes under refrigeration and were processed immediately. Positive urine cultures for S. aureus of male and female participants of all ages with colony count of ≥ 105 colony-forming units (CFU)/mL were included in the study. Positive urine cultures with ≤ 105 CFU/mL were excluded (2, 5).
3.3. Isolation and Colony Count
Homogenized urine samples without centrifugation were inoculated using a disposable loop (1 µL) onto plates containing cysteine lactose electrolyte-deficient (CLED) agar. The plates were incubated under aerobic conditions at 37°C for 24 to 48 hours and the number of colonies was counted (5).
3.4. Phenotypic Identification of S. aureus
Isolation and identification of S. aureus isolates were carried out making use of Gram stain microscopy and following conventional biochemical methods, such as colony morphology on blood agar, growth on mannitol salt agar, and production of catalase, coagulase, and DNase. Screening for MRSA was carried using a cefoxitin disc (30 µg) on Mueller Hinton agar (Merck, Germany) plates supplemented with 4% NaCl (12).
3.5. Genomic DNA Extraction
The researchers extracted the whole genomic DNA using the QIAamp DNA Mini Kit (Qiagen GmbH, Hilden, Germany), according to the manufacturer’s instruction with the modification of adding Lysostaphin (15 µg/mL) (Sigma-Aldrich, USA) used for cell wall lysis. DNA purity was determined using a spectrophotometer.
3.6. Genotypic Identification of S. aureus and MRSA Screening
In addition to the tests described above, presumptive S. aureus strains were submitted to genotypic identification using the polymerase chain reaction (PCR) for nucA genes to achieve definitive identification (13). Methicillin-resistant S. aureus isolates were confirmed by the amplification of mecA gene using PCR (14). Confirmed MRSA isolates were stored in tryptic soy broth (TSB; Merck, Germany) containing 20% glycerol at -70°C for molecular investigation.
3.7. Antimicrobial Susceptibility Testing (AST)
The susceptibilities of S. aureus isolates to the antimicrobial agents ampicillin (AP 10 µg), ciprofloxacin (CIP 5 µg), tetracycline (T 30 µg), erythromycin (E 15 µg), amikacin (AK 30 µg), clindamycin (CD 2 µg), gentamicin (GM 10 µg), and trimethoprim- sulfamethoxazole (TS 2.5 µg) were determined following the Kirby-Bauer disc diffusion method, according to the guidelines of the clinical and laboratory standards institute (CLSI) (12). All the antibiotic disks were prepared form MAST (Mast Diagnostics, Group Ltd, Merseyside UK). The minimum inhibitory concentration (MIC) for vancomycin was determined using E-test strips (AB BIODISK, Sweden) method. Multidrug resistance phenotype was defined as resistance of S. aureus to 3 or more unique antimicrobial categories in addition to beta-lactams (14). The reference strain, S. aureus ATCC25923, was included as quality control in parallel in each run.
3.8. Polymerase Chain Reaction Assays for Detection of Toxin Encoding Genes
All S. aureus isolates were tested for the presence of lukS-PV-lukF-PV (pvl genes) and toxic shock syndrome toxin (tst) gene using degenerate primers listed in Table 1.
| Target | Primer | Primer Sequence (5’ → 3’) | Product Size, bp | Reference |
|---|---|---|---|---|
| nucA | F | GCGATTGATGGTGATACGGTT | 270 | (8) |
| R | AGCCAAGCCTTGACGAACTAAAGC | |||
| mecA | F | AGAAGATGGTATGTGGAAGTTAG | 583 | (6) |
| R | ATGTATGTGCGATTGTATTGC | |||
| luk-PV | F | TTCACTATTTGTAAAAGTGTCAGACCCACT | 180 | (3) |
| R | TACTAATGAATTTTTTTATCGTAAGCCCTT | |||
| tsst-1 | F | TTATCGTAAGCCCTTTGTTG | 398 | (13) |
| R | TAAAGGTAGTTCTATTGGAGTAGG | |||
| spa | F | TAAAGACGATCCTTCGGTGAGC | Variable | (11) |
| R | CAGCAGTAGTGCCGTTTGCTT | |||
| SCCmec | Fβ | ATTGCCTTGATAATAGCCYTCT | 937 | (7) |
| R α3 | TAAAGGCATCAATGCACAAACACT | |||
| F ccrC | CGTCTATTACAAGATGTTAAGGATAAT | 518 | ||
| R ccrC | CCTTTATAGACTGGATTATTCAAAATAT | |||
| F 1272 | GCCACTCATAACATATGGAA | 415 | ||
| R 1272 | CATCCGAGTGAAACCCAAA | |||
| F 5RmecA | TATACCAAACCCGACAACTAC | 359 | ||
| R 5R431 | CGGCTACAGTGATAACATCC | |||
| F | ATCATTAGGTAAAATGTCTGGACATGATCCA | 433 | ||
| R | GCATCAAGTGTATTGGATAGCAAAAGC |
3.9. SCCmec Typing
The identified S. aureus strains were subjected to multiplex PCR for detection of SCCmec types, as described previously (7). The SCCmec types were identified through comparing the banding patterns of S. aureus to strain ATCC 10442 (SCCmec type I), strain N315 (SCCmec type II), strain 85/2082 (SCCmec type III), strain MW2 (SCCmec type IVa), and strain WIS (SCCmec type V), as reference strains. Primers for SCCmec typing are presented in Table 1.
3.10. spa Typing
Determining different spa types was performed by PCR, according to the method described by Harmsen et al. (11). The PCR products were subjected for DNA sequencing for both strands by Macrogen (Seoul, South Korea). The obtained sequences were edited using the Chromas software (Version 1.45, Australia). The edited sequences were assigned to particular spa types, according to the guidelines described in the spa typing website (http://www.spaserver.ridom.de). Primers for spa typing are presented in Table 1. The efficiency of amplicon was monitored through electrophoresis on 1% agarose gel (Invitrogen, Carlsbad, CA, USA) prepared in TAE buffer. The PCR product was stained with ethidium bromide and photographed under UV transillumination (UVItec, Cambridge, UK).
4. Results
A total of 90 S. aureus isolates were analyzed in the present study. The overall prevalence of patients with S. aureus complaining of UTI was 10.4%. Overall, 79 S. aureus isolates were obtained from female patients and 11 isolates from male patients. Participants’ mean age was 38 years (median 45.8 years, ranging from 2 to 68 years) and the age distribution was as follows: 10 patients were ≤ 20 years old, 65 patients 21 to 45 years old, and 15 patients ≥ 60 years old. According to the findings, 55 isolates (61.1%) were confirmed as MRSA using genotypic and phenotypic screening and 35 isolates (38.9%) were Methicillin-Sensitive S. aureus (MSSA).
As for the susceptibility to the tested antibiotics, the following resistance patterns were observed: ampicillin (94.4%), ciprofloxacin (75.6%), gentamicin (73.3%), erythromycin (72.2%), tetracycline (68.9%), clindamycin (60%), amikacin (57.8%), and trimethoprim- sulfamethoxazole (31.1%). Distribution of resistance rate among MRSA and MSSA isolates is presented in Table 2. Fortunately, all the strains were susceptible to vancomycin and inhibited at similar MIC50 and MIC90 of 1 µg/mL.
| Antibiotics | MRSA (N = 55) | MSSA (N = 35) | Total Resistance | ||
|---|---|---|---|---|---|
| Resistance | Sensitive | Resistance | Sensitive | ||
| Ampicillin | 55 (100) | 0 (0) | 30 (85.7) | 5 (14.3) | 85 (94.4) |
| Ciprofloxacin | 52 (94.5) | 3 (5.5) | 16 (45.7) | 19 (54.3) | 68 (75.6) |
| Gentamicin | 54 (98.2) | 1 (1.8) | 12 (34.3) | 23 (65.7) | 66 (73.3) |
| Erythromycin | 50 (90.9) | 5 (9.1) | 15 (42.9) | 20 (57.1) | 65 (72.2) |
| Tetracycline | 48 (87.3) | 7 (12.7) | 14 (40) | 21 (60) | 62 (68.9) |
| Clindamycin | 54 (98.2) | 1 (1.8) | 0 (0) | 35 (100) | 54 (60) |
| Amikacin | 17 (30.9) | 38 (69.1) | 32 (91.4) | 3 (8.6) | 52 (57.8) |
| Trimethoprim-Sulfamethoxazole | 21 (38.2) | 34 (61.8) | 7 (20) | 28 (80) | 28 (31.1) |
aValues are expressed as No. (%).
Based on antimicrobial susceptibility testing, all the isolates were identified as MDR. The resistance profiles among the isolates were as follows; resistance to 6, 4, 5, and 3 antimicrobial drugs, which were common among 41 (45.5%), 25 (27.8%), 23 (25.6%), and 1 (1.1%) isolates, respectively. The major resistance profile detected within the MSSA included resistance to ampicillin, gentamicin, tetracycline, and amikacin, while in MRSA, it included resistance to ampicillin, ciprofloxacin, gentamicin, erythromycin, tetracycline, and clindamycin.
The results obtained in the present study revealed that 25 isolates (27.8%) were positive for pvl-encoding gene and 40 (44.4%) isolates harbored tst-1 encoding gene. Also, 20 MRSA isolates were Panton-Valentine Leukocidin (PVL)-positive (20/55; 36.4%), while 5 MSSA isolates were PVL-positive (5/35; 14.3%); the percentage of MRSA PVL-positive was higher than that of the MSSA strains. All the PVL-positive isolates belonged to S. aureus isolates with resistance to 6 antimicrobial drugs while TST positive isolates were distributed among S. aureus isolates with resistance to 6, 5, and 4 antimicrobial drugs.
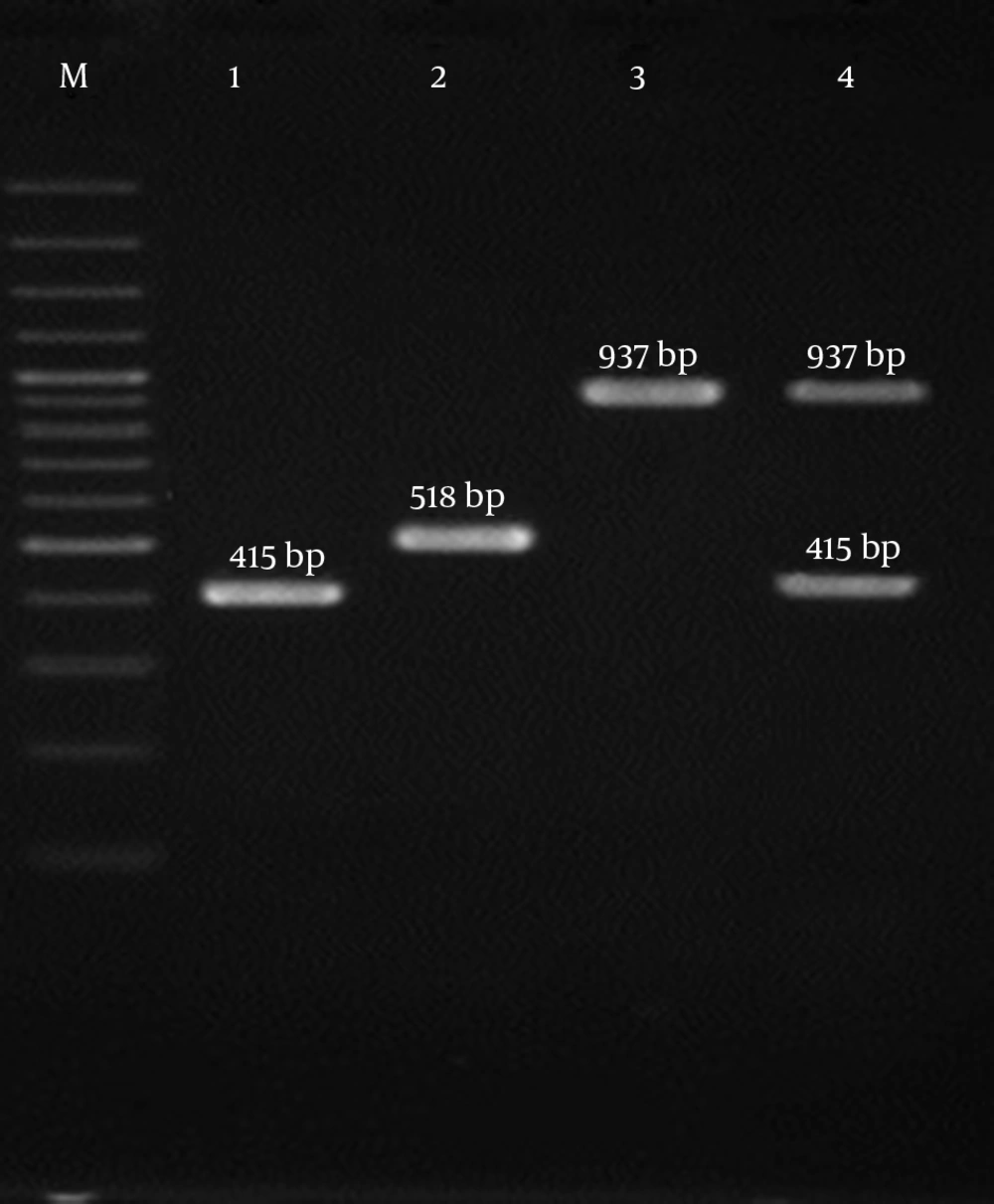
The results showed that all 4 SCCmec types (I - IV) were detected among the isolates investigated (Figure 1). The most prevalent SCCmec type was type III (n = 35; 38.9%), commonly known to be associated with hospital-associated MRSA (HA-MRSA) infections, followed by SCCmec type II (n = 28; 31.1%), SCCmec type IV (n = 26; 28.9%), and SCCmec type I (n = 1; 1.1%). The current analysis revealed that PVL-positive isolates belonged to SCCmec type IV (52%) and SCCmec type II (48%). Moreover, tst encoding genes were found to be distributed among all SCCmec types with a high frequency rate in SCCmec type III (27.8%). All the patients with SCCmec types II and III isolates belonged to the age group of 21 to 45 years old. Patients infected with SCCmec types IV were distributed in all age groups with the majority in the age group of ≥ 60 years.
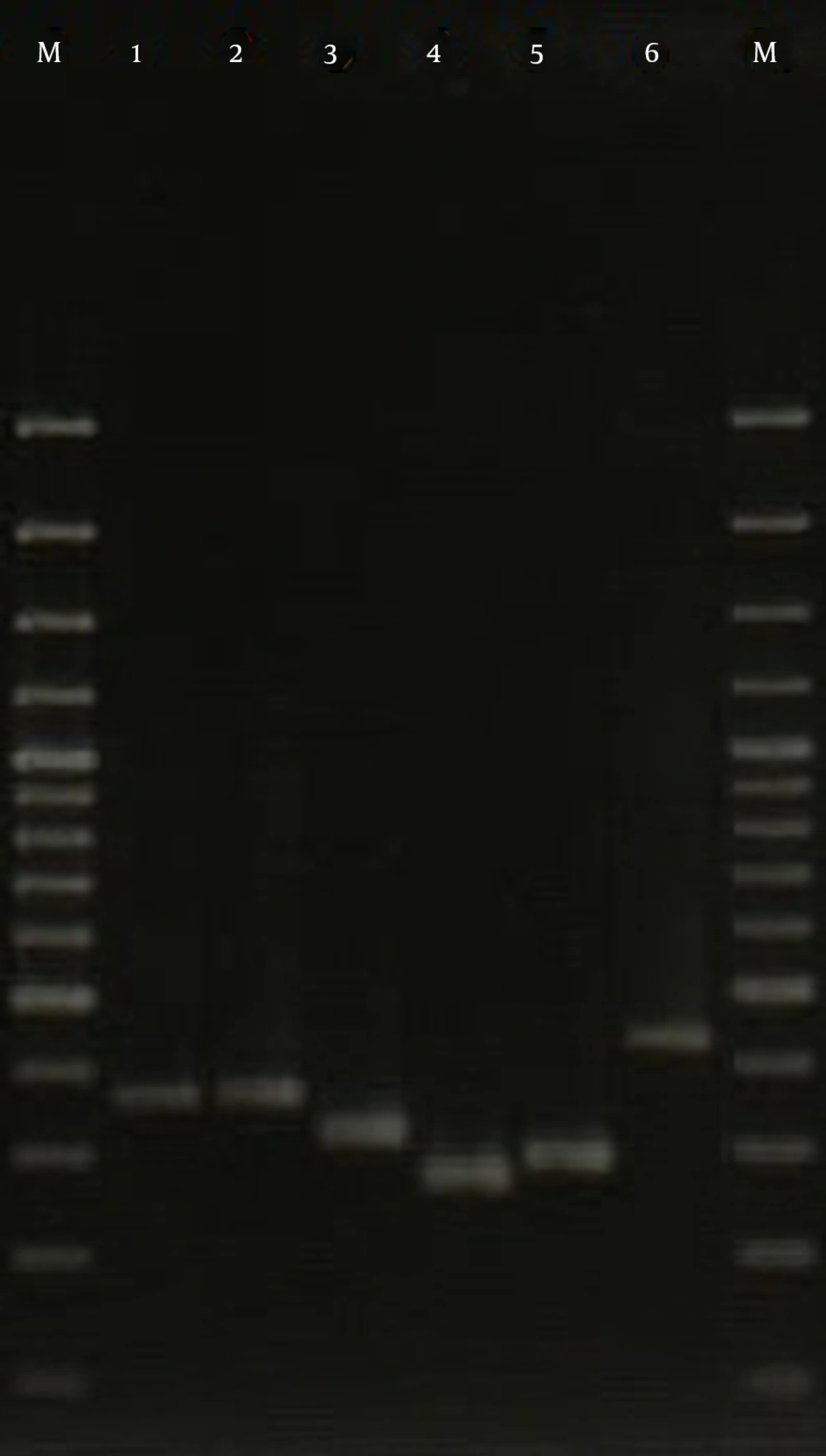
The isolates investigated in the current study were assigned to 10 different spa types, with the most common being spa type t037, accounting for 23.3% of isolates followed by t924 (15.6%), t383 (15.6%), t426 (12.2%), t044 (12.2%), t790 (6.7%), t084 (4.4%), t021 (4.4%), t7580 (4.4%), and t064 (1.2%) (Figure 2). Also, 25 PVL-positive isolates were distributed among 5 different spa types, including t426 (8 isolates), t044 (6 isolates), t021 (4 isolates), t790 (4 isolates), and t084 (3 isolates).
According to the results of MRSA screening, all the spa types, t044, t790 and t084, analyzed in the study, were found to be MRSA while all the spa types t924 and t7580 were MSSA. The tst gene was identified in 7 spa types (t037, t924, t383, t426, t084, t021, and t7580). None of the spa types t044, t790, and t064 were positive for tst encoding gene. The distribution of the spa types among MRSA and MSSA strains are presented in Table 3. Distribution of different spa types among MRSA and MSSA isolates with different MDR profiles was variable. The spa types t7580 and t924 were only isolated from MSSA strains. The most prevalent spa type within the MRSA and MSSA population were spa type t037 (17/55:30.9%) and t924 (12/35:34.3%), respectively. Table 4 shows the distribution of antimicrobial resistance profiles among MRSA and MSSA spa types.
| Characteristic | MRSA | MSSA | pvl | tst | Total |
|---|---|---|---|---|---|
| spa types | |||||
| 037 | 17 (81) | 4 (19) | 0 (0) | 16 (76.2) | 21 (23.3) |
| 924 | 0 (0) | 14 (100) | 0 (0) | 9 (64.3) | 14 (15.6) |
| 383 | 8 (57.1) | 6 (42.9) | 0 (0) | 5 (35.7) | 14 (15.6) |
| 426 | 6 (54.5) | 5 (45.5) | 8 (72.7) | 3 (27.3) | 11 (12.2) |
| 044 | 11 (100) | 0 (0) | 6 (54.5) | 0 (0) | 11 (12.2) |
| 790 | 6 (100) | 0 (0) | 4 (66.7) | 0 (0) | 6 (6.7) |
| 084 | 4 (100) | 0 (0) | 3 (75) | 1 (25) | 4 (4.4) |
| 021 | 2 (50) | 2 (50) | 4 (100) | 2 (50) | 4 (4.4) |
| 7580 | 0 (0) | 4 (100) | 0 (0) | 4 (100) | 4 (4.4) |
| 064 | 1 (100) | 0 (0) | 0 (0) | 0 (0) | 1 (1.2) |
aValues are expressed as No. (%).
| No of Antibiotics | Resistance Profile | MSSA spa Type (N = 35), (No; %) | MRSA spa Type (N = 55), (No; %) | Total, No. (%) |
|---|---|---|---|---|
| 6 | AP, CIP, GM, E, T, CD | t426 (2; 4.9), t021 (2; 4.9), t924 (5; 12.2) | t037 (7; 17.1), t426 (6; 14.6), t044 (6; 14.6), t790 (6; 14.6), t084 (3; 7.3), t021 (2; 4.9), t383 (2; 4.9) | 41 (45.5) |
| 5 | AP, CIP, E, TS, AK | t7580 (3; 13.1), t037 (2; 8.7), t924 (2; 8.7), t383 (3; 13.1) | t084 (1; 4.3), t037 (5; 21.7), t383 (2; 8.7) | 23 (25.6) |
| AP, CIP, AK, E, TS | t924 (2; 8.7) | t064 (1; 4.3), t037 (2; 8.7) | ||
| 4 | AP, GM, T, AK | t924 (5; 20), t383 (3; 12), t426 (3; 12) | t037 (3; 12), t383 (4; 16), t044 (3; 12) | 25 (27.8) |
| CIP, AK, GM, TS | t037 (2; 8) | t044 (2; 8) | ||
| 3 | E, AK, TS | t7580 (1; 100) | - | 1(1.1) |
Abbreviations: AK, Amikacin; AP, Ampicillin; CD, Clindamycin; CIP, Ciprofloxacin; E, Erythromycin; GM, Gentamicin; T, Tetracycline; TS, Trimethoprim-Sulfamethoxazole.
5. Discussion
The present study describes the antibiotic resistant pattern and molecular types of S. aureus strains isolated from patients with UTI using SCCmec and spa typing methods. According to the obtained data, it was revealed that S. aureus accounted for 10.4% of UTI. The current findings were relatively high in comparison with those of studies conducted in Great Britain (0.5%) (15) and France (1.3%) (16). Overall, it is well established that isolation rate of S. aureus in UTI is between 2% and 15% (5). In the present study, MRSA screening showed that 61.1% of the tested isolates were confirmed to be MRSA, which was lower than those reported in previous studies from Iran (8), and higher than those reported in Croatia (17). Moreover, different prevalence rates are reported for MRSA by different researchers. This variety in the prevalence rates could be due to the difference among these studies in terms of applying standard infection control programs in ICUs, study designs, antibiotic prescription, sample type, investigated population, and laboratory testing in order to determine methicillin resistance.
Despite the previous data about the emergence of MRSA resistance to vancomycin in Iran (18), the current data showed that all the MRSA and MSSA isolates were sensitive to vancomycin revealing the limited and appropriate use of vancomycin and prescription protocols in the Iranian health care systems. According to the literature, MRSA strains are resistant not only to beta-lactams but also to macrolides, lincosides, and aminoglycosides (6, 8, 18). In the present study, the majority of the MRSA isolates were resistant to ampicillin (100%), ciprofloxacin (94.5%), gentamicin (98.2%), erythromycin (90.9%), tetracyclin (87.3%), and clindamycin (98.2%), which is in line with the findings reported in Iran (8), Croatia (17), and Turkey (19). The rate of resistance to tested antibiotics, with the exception of amikacin among MRSA isolates, was higher than those among MSSA isolates, which is in accordance with the findings reported by Yu et al. (20). In contrast to Ajantha et al. (21), the lowest resistance rate (31.1%) was noted against trimethoprim-sulfamethoxazole, as an empirical drug of choice in the treatment of UTI.
When MDR S. aureus increases, appropriate therapeutic options decrease and thus morbidity and mortality in hospitalized patients increases. Similar to the studies carried out in Serbia (83.9%) (22) and Taiwan (75.8%) (23), a high prevalence rate of MDR was detected among tested isolates of the present study. The high MDR rate in the current study reflected the lack of stringent antibiotic policies in clinical practice. It is well-established that SCCmec types I, II, and III are related to HA-MRSA while SCCmec types IV and V are prominent types in CA-MRSA (8). The results of the current study showed that the most prevalent SCCmec type was type III (38.9%). This finding is in agreement with those reported from China (24) and Brazil (25), yet differs from that reported by Vazquez et al. in Spain (26). The high frequency of SCCmec type III in the current study emphasizes the nosocomial origin of these strains in patients with UTIs.
Regarding PVL-positive strains, these isolates were mostly found in MRSA strains. In contrast to the current finding, O’Malley et al. (27) indicated that pvl-encoding genes were identified in 24% of known S. aureus isolates, with all but one found in MSSA. All the PVL-positive isolates belonged to SCCmec type IV and II. This finding is different from that reported by Rodrigues et al., who observed that none of the PVL-positive MRSA strains harbored SCCmec type IV (25), yet the current findings were supported by other studies (8, 28). All the PVL positive isolates were resistance to six antimicrobial drugs. Our findings suggest that pvl-encoding genes existed in antibiotic-resistant organisms in Tehran, Iran. In addition, 40 tested isolates (44.4%) harbored tst, which was the most prevalent toxin gene observed in the present study. Isolates harboring tst were distributed in all SCCmec types with high frequency of SCCmec type III. Several different molecular types of strains harboring tst were previously described (4).
Distribution of spa types varied from one geographic region to another. In the present study, 10 different spa types were identified, among which spa type t037 was the most prevalent, accounting for 23.3% of the isolates. This spa type was reported in previous studies conducted in Iran (10), China (29), and Saudi Arabia (30). It is noteworthy to mention that the majority of isolates (81%) with spa type t037 were MRSA. This finding was previously reported by several researchers (31). It appears that spa type t037 is linked to resistance to methicillin. The second most common spa types identified in the current study were t924 and t383 (15.6%). In a study conducted by Ellington et al. in the UK, in order to determine the prevalence of the pvl genes among MSSA and MRSA bacteremia isolates from UK and Ireland, PVL-positive spa type t383 was reported (32). All the spa types 924 in the present study were found to be MSSA. Also, a search on the Ridom Spa server (http://spa.ridom.de/spa-t924.shtml/) revealed that spa type 924 was reported in Germany and Sweden.
Considering the literature, spa type t044 PVL-positive MRSA is widely disseminated in European countries (33). All the isolates with t044 (12.2%), as the third most common spa type identified in the current study, were MRSA and belonged to SCCmec IV and some of them (54.5%) were PVL-positive. In line with the results of the present study, spa type t044 PVL-positive MRSA was reported from Sweden (34), UK (32), and Lebanon (35) In accordance with the findings of the present study, in a study conducted by Meemken et al. (36) on S. aureus isolated from animals in Germany, spa type t426 was detected in both MRSA and MSSA isolates. These data suggest transmission of MRSA spa type t426 between different hosts (humans and animals). Also, similar to the results of previous studies from other countries, spa type t790 was found in both PVL-negative and positive MRSA strains (37). It was also found that most spa t790 isolates belonged to SCCmec IV.
The spa type t790 was reported in Goudarzi et al.’s study, which was carried out to study molecular characterization of MRSA clinical strains (8). Unlike many studies, which reported t790 as the predominant spa type, there are several investigations reporting lower frequencies of this strain in different geographic areas. Considering the high prevalence of t790, the current study demonstrated its transfer from the community to hospitals. In the present study, other infrequent spa types, such as t084 and t064, were obtained from MSSA strains. The spa type 084 was reported earlier by Goudarzi et al. from Iran (6). According to the literature, spa type t064 was reported in MRSA clinical isolates from hospitals and horses in other countries (38, 39).
Four (4.4%) t021 isolates were characterized in the present study, which were equally distributed between MRSA and MSSA isolates. This spa type has been detected in previous investigations from Lebanon, Ireland, Romania, and Portugal (40). The spa type t7580 was observed among 4 isolates (4.4%); all these 4 isolates were MSSA, harbored tst encoding gene, and belonged to SCCmec III. None of these spa types were PVL-positive. The results of the current study are in agreement with those of another investigation conducted in Iran, showing that the spa type t7580 was low (4.3%) and the pvl-encoding gene was not confirmed in any of these types (8). The current study had some limitations, which should be pointed out. First, the sample size was low. The second limitation was the restriction on the application of molecular techniques, such as PFGE and MLST.
6. Conclusions
To summarize, the current study revealed different spa and SCCmec types that were disproportionately distributed between MRSA and MSSA isolates. In the present study, spa types t426 and t021 were identified from Iran for the first time, which places emphasis on the fact that continuous and nationwide MRSA surveillance studies should be periodically performed to identify the major molecular type and also to characterize the emergence of new strains.