1. Background
Klebsiella pneumoniae, as an opportunistic pathogen, can be the cause of a range of nosocomial and community - acquired infections (1-3). Pathogenic K. pneumoniae strains are responsible for urinary tract, respiratory, and blood infections and are associated with mortality and morbidity in patients (4-6). Pathogenicity of K. pneumoniae is the result of production of many virulence factors that help these bacteria overcome the immune system and cause various diseases. Different virulence factors, such as lipopolysaccharide (o - antigen), capsular polysaccharides (K antigen), fimbriae and siderophores contribute to the pathogenicity of Klebsiella (7, 8). Among these factors, the capsule is one of the most important virulence determinants, which helps achieve biofilm formation and leads to protection against phagocytosis, antimicrobial peptides, and serum bactericidal activity. There are more than 77 types of capsular antigens in K. pneumoniae strains, of which K1 and K2 serotypes are well known (9, 10).
Klebsiella pneumoniae also carries virulence - associated genes, which may encode capsules (magA, K2A, WcaG) and a capsular regulator gene (regulator of mucoid phenotype (rmpA). Mucoviscosity - associated gene A (magA) and WcGA genes located within the gene clusters are responsible for encoding capsular polymerase and capsular biosynthesis, respectively (11-13). Capsular biogenesis by magA is achieved through conversion of manose to fucose, which may enhance the ability of the bacteria to evade phagocytosis by macrophage. The regulator of mucoid phenotype A (rmpA) gene located on the plasmid increases capsular polysaccharide production and is found in the hypermucoviscous phenotype. Moreover, two genes, including the magA and rmpA, were initially associated with invasive infections (14, 15).
Resistance of pathogenic bacteria to different antibiotics has become a serious worldwide problem because of the fatal outcome of defective treatment and the difficulty to find treatment options (16). Klebsiella pneumoniae has been revealed to have the ability to acquire resistance to many antibiotics, especially third generation cephalosporins. Multidrug resistance of K. pneumoniae against the antibiotics used for therapy intensifies the virulence potential of Klebsiella. Beta - lactam antibiotics are one of the most commonly used antibiotics in the treatment of bacterial infections and the production of B - lactamase enzymes are the most common bacterial resistance mechanisms (17, 18). In the recent years, Extended Spectrum Beta Lactamase (ESBL) producer K. pneumoniae have increased over the world. The ESBLs are divided to several groups; the main groups are TEM, CTX, and SHV derivatives (19, 20).
2. Objectives
The present study aimed at determining the prevalence of magA, wcaG, rmpA, Capsular type K1 and Capsular type K2 virulence genes along with SHV and TEM beta lactam resistance genes in K. pneumoniae isolates.
3. Methods
3.1. Ethics Statement
The ethics committee of the Semnan University of Medical Sciences, Semnan, Iran, approved this study (IR.SEMUMS.REC 1394.29).
3.2. Sampling
A cross sectional study was performed. A total of 173 non - duplicate K. pneumoniae isolates were collected during year 2015 at two different hospitals of Semnan, Iran, from urine specimens.
3.3. Bacterial Identification
The specimens were cultured on blood agar and Eosin Methylene Blue (EMB) agar (Merck, Darmstadt, Germany), and incubated at 37°C for 24 hours. Klebsiella pneumoniae was identified by conventional bacteriological tests. Gram stain was performed for suspected colonies. Biochemical tests, including lactose fermentation, gas production from glucose, FeS production, motility, indole production, sodium citrate utilization, and urea utilization were used for K. pneumoniae identification (21).
3.4. Antibiotic Susceptibility Testing
Disks diffusion test was performed according to the guidelines of the Clinical and Laboratory Standards Institute (22). Used disks (Rosco, Taastrup, Denmark) were amikacin (30 μg), ciprofloxacin (5 μg), amoxicillin - clavulanic acid (20/10 μg), ceftazidime (30 μg), imipenem (10 μg), cefepime (30 μg), aztreonam (30 μg), nitrofurantoin (300 μg), and trimethoprim - sulfamethoxazole (1.25/23.75 μg). Escherichia coli (ATCC 25922) strain was used for quality control.
3.5. DNA Extraction
DNA of each K. pneumoniae isolate was extracted from 1 mL of overnight bacterial culture. Extraction was performed as previously described by Kuske et al. (23). The supernatant was used as the template DNA in PCR reactions.
3.6. Detection of Capsular type K1, Capsular type K2, rmpA, wcaG, TEM, and SHV
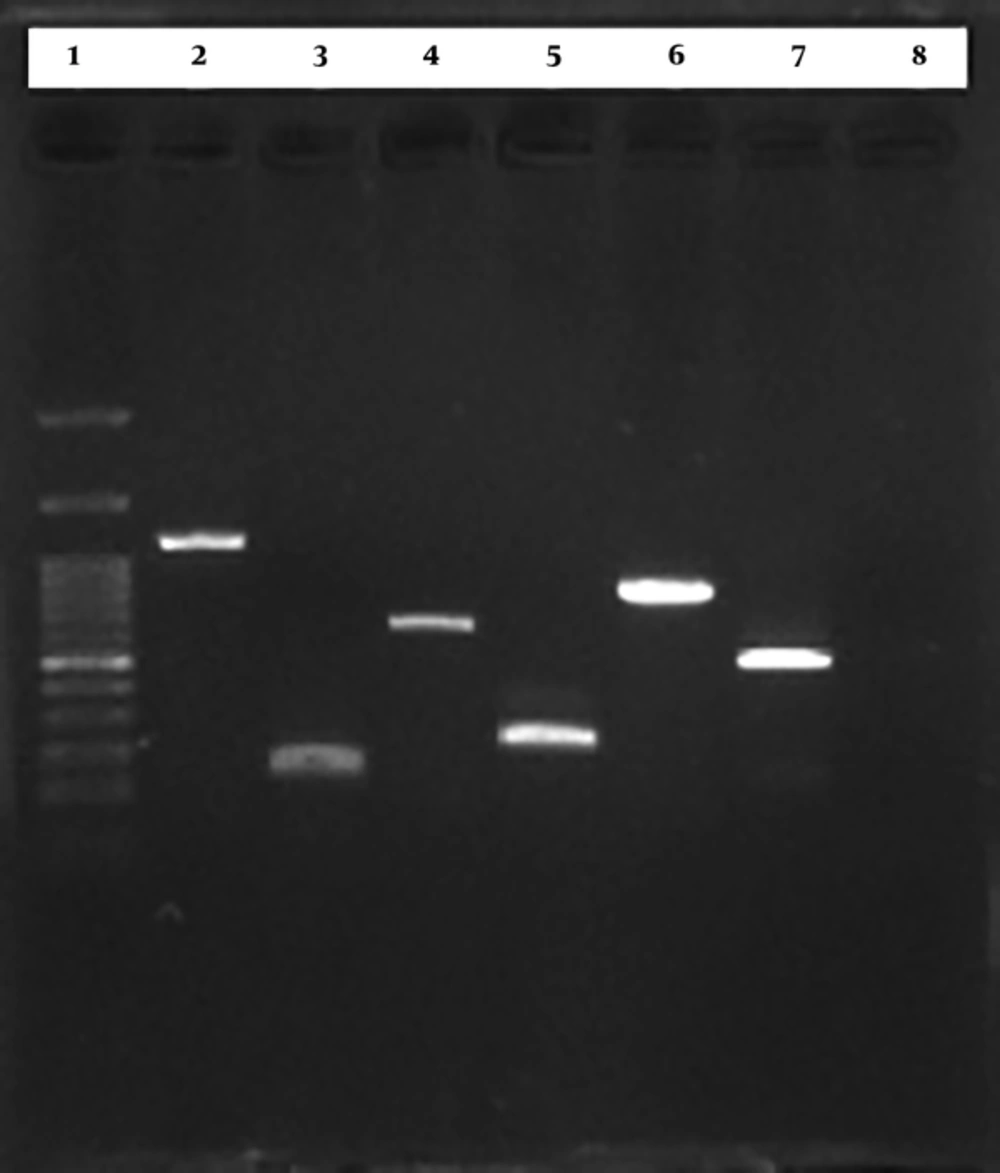
Specific primers for detection of Capsular type K1, Capsular type K2, rmpA, wcaG, TEM, and SHV are shown in Table 1. For capsular type K1, capsular type K2, rmpA, and wcaG amplification were performed as follow: initial denaturation at 95°C for five minutes, followed by 35 cycles at 94°C for 30 seconds, 58°C for 90 seconds and 72°C for 90 seconds and a final extension at 72°C for 10 minutes (24). Polymerase Chain Reaction (PCR) conditions for TEM amplification were as follow: initial denaturation at 94°C for three minutes, 35 cycles at 94°C for 30 seconds, 45°C for one minute, 72°C for one minute, final extension at 72°C for 10 minutes (25). The SHV was amplified under the following conditions: 94°C for three minutes, 35 cycles at 94°C for 30 minutes, 60°C for one minute, 72°C for one minute, and a final extension at 72°C for 10 minutes (25). The PCR products were analyzed by electrophoresis with 1% agarose gel in 1X Tris - Acetate - EDTA buffer. The gels were stained with ethidium bromide and the PCR products were visualized under UV light (Figure 1).
| Gene | Primer | Sequence | Product Size (bp) | Reference |
|---|---|---|---|---|
| Capsular type K1 | MagA - F | GGTGCTCTTTACATCATTGC | 1283 | Turton et al. (24) |
| MagA - R | GCAATGGCCATTTGCGTTAG | |||
| Capsular type K2 | K2wzy - F | GACCCGATATTCATACTTGACAGAG | 641 | Turton et al. (24) |
| K2wzy - R | CCTGAAGTAAAATCGTAAATAGATGGC | |||
| rmpA | rmpA - F | ACTGGGCTACCTCTGCTTCA | 516 | Turton et al. (24) |
| rmpA - R | CTTGCATGAGCCATCTTTCA | |||
| wcaG | wcaG - F | GGTTGGKTCAGCAATCGTA | 169 | Turton et al. (24) |
| wcaG - R | ACTATTCCGCCAACTTTTGC | |||
| SHV | SHV - F | AAGATCCACTATCGCCCAGCAG | 200 | Shahcheraghi et al. (25) |
| SHV - R | ATTCAGTTCCGTTTCCCAGCGG | |||
| TEM | TEM - F | GAGTATTCAACATTTCCGTGTC | 800 | Shahcheraghi et al. (25) |
| TEM - R | TAATCAGTGAGGCACCTATCTC |
3.7. Statistical Analysis
Confidence interval test was used to assess the statistical significance with confidence level of 95% (α = 0.05).
4. Results
In this study, frequency of virulence factors was as follow: capsular type K2: 57 out of 173 (32.9% CI: 39.9%, 25.9%), rmpA: 35 out of 173 (20.2% CI: 26.2%, 14.2%), wcaG: 28 out of 173 (16.2% CI: 19.4%, 13%), capsular type K1: 12 out of 173 (6.9% CI: 10.6%, 3.2%). Also, the SHV was observed in 81 out of 173 (46.8% CI: 51.2%, 42.4%) and TEM was observed in 58 out of 173 (33.5% CI: 37.5%, 29.5%) cases. The resistant rates to antibiotics were as follow, imipenem: 13 out of 173 (7.5% CI: 9.8%, 5.2%), ciprofloxacin: 28 out of 173 (16.1% CI: 19.2%, 13%), amoxicillin - clavulanic acid: 52 out of 173 (30% CI: 33.9%, 26.1%), trimethoprim - sulfamethoxazole: 57 out of 173 (32.9% CI: 36.9%, 28.9%), cefepime: 59 out of 173 (34.1% CI: 38.2%, 30%), nitrfurantoin: 62 out of 173 (35.8% CI: 35.8%, 27.8%), amikacin: 63 out of 173 (36.4% CI: 40.5%, 32.3%), aztreonam: 68 out of 173 (39.3% CI: 43.5%, 35.1%).
5. Discussion
Klebsiella pneumoniae is the cause of nosocomial and community acquired infections, such as urinary tract, respiratory, and blood infections. It harbors many virulence-associated genes including magA, rmpA, WcaG, fimbriae, and siderophores, which help the bacteria overcome the immune system and cause infection. Drug resistant, especially ESBL - producing K. pneumoniae strains, contribute to treatment failure and increase morbidity and mortality in patients (26-28).
In this study, the frequency of SHV was significantly lower than 69.6% and 67.4% reported by Shahcheraghi et al. (25) and Feizabadi et al. (29), respectively. Frequency of TEM was significantly lower than 54%, reported by Feizabadi et al. (29), yet in accordance with 32.1% reported by Shahcheraghi et al. (25). These differences were the result of differences in infection control system and therapeutic regimens in different parts of Iran. Age of over 60, long hospitalization, diabetes and previous antibiotic treatment were other factors associated with the ESBL phenotype (30).
Capsular type K1 frequency was significantly lower than 39.5% and 64.3% reported by Liu et al. (31) and Lee et al. (32), yet significantly higher than 1.4% reported by Maatallah et al. (33). Capsular type K2 frequency was significantly higher than 18.4% and 20% reported by Liu et al. (30) and Lee et al. (32), yet significantly lower than 5% reported by Maatallah et al. (33). Frequency of rmpA was significantly higher than 5.5%, 3.6%, and 7.0% reported by Derakhshan et al. (27), Maatallah et al. (33), and Derakhshan et al., respectively (34). Frequency of wcaG was significantly higher than 2.7% and 8.6% reported by Derakhshan et al. (27) and Maatallah et al. (33), yet significantly lower than 23.5% reported by Derakhshan et al. (34).
The resistance rate to imipenem was significantly higher than 1.4% and 0.05% reported by Liu et al. (31), and Derakhshan et al. (34), and also reports of Shahcheraghi et al. (25), Peerayeh et al. (26), Derakhshan et al. (27), Peerayeh et al. (28), and Ranjbar et al. (35), which reported no resistance to imipenem, yet significantly lower than 20.8% reported by Amraie et al. (36). The resistance rate to ciprofloxacin was significantly lower than 44.4%, 37.1%, 42.5%, and 52.5% reported by Derakhshan et al. (27), Liu et al. (31), Derakhshan et al. (34), and Ranjbar et al. (35), yet in accordance with 18%, 18%, and 15.6% reported by Shahcheraghi et al. (25), Peerayeh et al. (26), and Amraie et al. (36). The resistance rate to amoxicillin - clavulanic acid was significantly lower than 31.7% and 28.6% reported by Liu et al. (31) and Pereira et al. (37), yet in accordance with 100% and 60.5% reported by Derakhshan et al. (27) and Derakhshan et al. (34), respectively.
The resistance rate to nitrofurantoin was significantly lower than 40% and 71% reported by Ranjbar et al. (35) and Amraie et al. (36). The resistance rate to trimethoprim - sulfamethoxazole was significantly lower than 39%, 55.5%, 39.3%, 50%, 51.5%, and 54% reported by Peerayeh et al. (26), Derakhshan et al. (27), Peerayeh et al. (26), Derakhshan et al. (34), Ranjbar et al. (35), and Amraie et al. (36), respectively, yet in accordance with 32.9% reported by Liu et al. (31). The resistance rate to cefepime was significantly higher than 12.9% reported by Liu et al. (31), yet in accordance with 37.5% reported by Peerayeh et al. (26), also significantly lower than 80.5% and 57% reported by Peerayeh et al. (26) and Derakhshan et al. (34).
The resistance rate to amikacin was significantly higher than 14%, 27%, 10%, 22%, and 4.8% reported by Shahcheraghi et al. (25), Peerayeh et al. (26), Liu et al. (31), Amraie et al. (36), and Pereira et al. (37), yet in accordance with 31.5% reported by Derakhshan et al. (34), also significantly lower than 50% reported by Derakhshan et al. (27). The resistant rate to aztreonam was significantly lower than 100%, 59%, and 92.5% reported by Derakhshan et al. (27), Derakhshan et al. (34), and Ranjbar et al. (35), yet in accordance with 39% and 38.1% reported by Peerayeh et al. (26) and Pereira et al. (37), also, significantly higher than 30% reported by Liu et al. (31). The resistance rate to ceftazidime was significantly higher than 31.3%, 51.1%, 35.8%, and 4.8% reported by Shahcheraghi et al. (25), Peerayeh et al. (28), Liu et al. (31), and Pereira et al. (37), yet significantly lower than 100%, 57%, 100%, and 49.7% reported by Derakhshan et al. (27), Derakhshan et al. (34), and Ranjbar et al. (35).
6. Conclusions
Frequency of some virulence factors including Capsular type K2, rmpA, wcaG, and also resistance rates to imipenem, amikacin, and ceftazidime were significantly higher than similar studies. Presence of virulence factors accompanied by drug resistance make bacteria an infectious agent and lead to treatment failure.