1. Background
Some species of Enterobacteriaceae, especially Klebsiella pneumoniae and Escherichia coli can produce carbapenemases; such pathogens are known as K. pneumoniae carbapenemase (KPC), New Delhi metallo-β-lactamase (NDM), and carbapenem-hydrolyzing oxacillinase (OXA-48). Carbapenemase-producing Enterobacteriaceae are a worldwide problem. In the study by Glasner et al. (1), carbapenemase-producing Enterobacteriaceae was identified in 26 of 31 (83.87%) analyzed European countries; only three years later, carbapenemase-producing Enterobacteriaceae was observed in 36 of 39 (92.31%) countries. From 01 November 2013 to 30 April 2014, in 2703 clinical samples from 455 hospitals in 36 countries, there were 85% K. pneumoniae and 15% E. coli isolates. Among them, 37% of K. pneumoniae and 19% of E. coli were KPC, NDM, OXA-48, and VIM (verona integron-encoded metallo-β-lactamase). The prevalence of these pathogens differed greatly, with the highest incidence in Mediterranean and Balkan countries (2). In 2013, only six countries in Europe reported the interregional spread of an endemic situation for carbapenemase-producing Enterobacteriaceae, whereas in 2015 this number increased to 13 countries (2).
Increased hospital mortality (27%) is reported in patients with bacteremia and pneumonia caused by carbapenem-resistant K. pneumoniae (CRKP) when compared with the 12% rate observed in individuals with urinary colonization by CRKP (3). The hospitalization time of patients with CRKP-related infections was also longer (4). Amongst all carbapenemase-producing Enterobacteriaceae, NDM pathogens represent the most important epidemiological and clinical problems (2, 5). The first case of E. coli NDM-1 was isolated in Poland in 2011 (6) from a patient recently returned from Congo. In 2013, the first K. pneumoniae NDM case was isolated in Warsaw, Poland. Over the next three years, this pathogen caused several dozens of outbreaks in Warsaw and the Warsaw region. Until the end of 2017, such infections were confirmed in more than 3,000 patients.
Prior use of carbapenems and previous hospitalization in countries with a high prevalence of carbapenemase-producing Enterobacteriaceae are reported as independent risk factors to acquire KPC and other carbapenemase-producing Enterobacteriaceae pathogens (7, 8). The health risks for a population associated with carbapenemase-producing Enterobacteriaceae infections result from three properties of these pathogens: (i) Resistance to almost all antimicrobials, except colistin and occasionally gentamycin and co-trimoxazole; (ii) the ability to transfer genes to other bacterial species due to the localization of such genes on plasmids and transposons; and (iii) the potential capacity to spread. Therefore, reducing the spread of bacteria requires restrictive compliance with hygiene rules, prompt methods of bacterial identification, and carrier isolation. The precise determination of bacterial susceptibility and minimal inhibitory concentration (MIC) values are necessary. It is also important not to administer antimicrobial therapy to asymptomatic carriers. In patients with urinary tract infections, monotherapy is adequate. For blood and pulmonary infections, combined therapy is indicated, with administration of colistin (9, 10).
Carbapenemase-producing Enterobacteriaceae infections are associated with high mortality, primarily due to delays in the administration of effective treatment and also the limited availability of effective treatment options. No new antimicrobials capable of replacing the carbapenems are available in the near future (5). Therefore, knowledge of the epidemiological situation and also the bacterial susceptibility and any resistance to antimicrobials is vital to make clinical decisions.
2. Objectives
The current study aimed at conducting a microbiological investigation of the antibiotic resistance of carbapenemase-producing Enterobacteriaceae pathogens isolated from the biological materials withdrawn from patients hospitalized at our center and analyzing the infections caused by the species.
3. Methods
3.1. Ethics Statement
The current study was approved by the Ethics Committee of the Military Institute of Medicine (No.105/WIM/2018). All patients’ personal information remained confidential.
3.2. Study Population
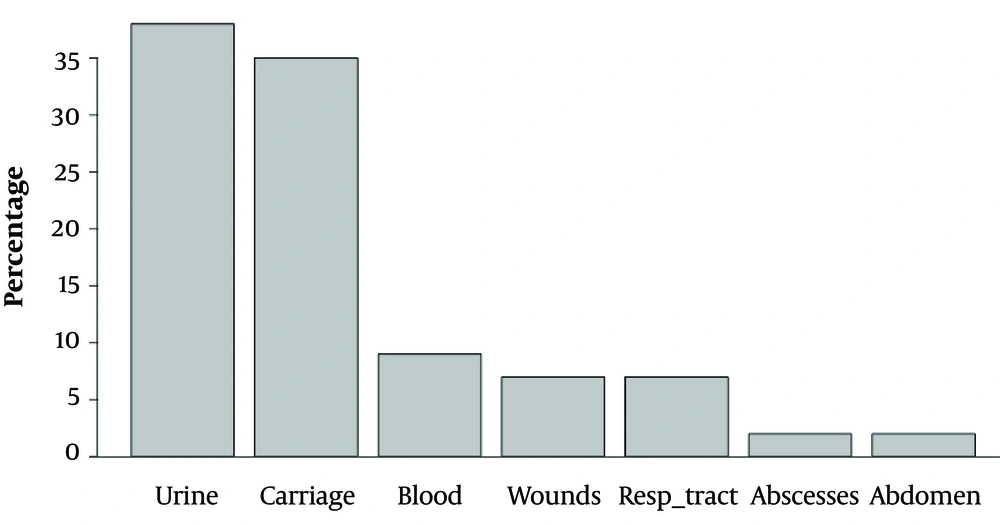
The current study analyzed 122 carbapenemase-producing Enterobacteriaceae isolates obtained from 122 patients (78 males and 44 females) hospitalized at the Military Institute of Medicine in Warsaw, Poland. The data were collected from 01 January 2009 to 03 December 2016 from the following sample types: Urine (n = 46), cloacal swabs for fecal carriage (n = 43), blood (n = 11), wounds (n = 9), bronchial tree aspirates collected via an endotracheal tube (n = 8), abscesses (n = 3), and fluid collected from the abdominal cavity (n = 2). Only one isolate per patient was included in the analysis. In case of multiple isolates, the first isolate was included.
3.3. Microbiological Examination
The samples were processed according to generally accepted microbiological procedures. They were inoculated on standard culture media (e g, Columbia agar, MacConkey agar, and chromID® Carba Smart, a selective chromogenic medium to screen carbapenemase-producing Enterobacteriaceae to detect carbapenemase-producing Enterobacteriaceae carriers; bioMérieux, France). The cultures were incubated at 37°C in air containing a 5% carbon dioxide atmosphere for 24 - 48 hours. When the growth of mixed bacterial cultures was observed, separating procedures were performed to obtain pure pathogen isolates.
3.4. Antibiotic Susceptibility
Detailed microbial identification was performed with the automatic VITEK® 2 testing system (bioMérieux, France). A microbroth dilution method with VITEK® 2 AST cards was used to test the antibiotic susceptibility of the isolated pathogens. The microbiological analyses were conducted from 2009 to 2010 according to the Clinical and Laboratory Standard Institute (CLSI) recommendations. The analyses performed from 2011 to date were according to the regulations of the european committee on antimicrobial susceptibility testing (EUCAST) and the national reference center for susceptibility testing (NRCST, Warsaw, Poland).
Control susceptibility tests included reference isolates of Klebsiella pneumoniae ATCC BAA-1705, ATCC BAA-1706, and ATCC 25955.The carbapenemase production was detected in the analyzed isolates of Enterobacteriaceae via phenotypic, biochemical, and molecular methods. The phenotypic tests were performed and interpreted according to the EUCAST and NRCST criteria to identify class A (KPC) carbapenemase, class B (MBL), and class D (OXA-48). To detect class A carbapenemase, the combined disk test with phenylboronic acid on Mueller-Hinton agar was used. The test was considered positive when the diameter of the growth-inhibitory zone around a β-lactam disk with phenylboronic acid was ≥4 mm larger than that of a disk containing the β-lactam substrate alone.
Class B carbapenemases (metallo-β-lactamase: NDM, IMP, and VIM types) were isolated using a double disc synergy test (DDST) with ethylenediamine tetraacetic acid (EDTA), ceftazidime (30 µg), and imipenem (10 µg) disks placed on Mueller-Hinton agar (bioMérieux, France). The tests were considered positive when a growth-inhibition zone was present around the disk with ceftazidime and/or carbapenem from the side of the disk with EDTA, an MBL inhibitor. To detect class D carbapenemase (OXA types like OXA-48), a disk containing temocyclin (30 µg) was placed on Mueller-Hinton agar (bioMérieux) (11, 12). The test result was considered positive and OXA-48 production was suspected when the growth-inhibition zone around the disk was 10 mm or less. The Rapidec® Carba NP (bioMérieux) biochemical test, which is based on the direct detection of carbapenem hydrolysis by carbapenemase-producing bacteria, was introduced into diagnostics in 2015.
3.5. Detection of Carbapenemase Genes
All isolates identified as carbapenemase-producing pathogens were sent to the NRCST in Warsaw, Poland, for reference diagnostics and surveillance purposes. The NRCST confirmed carbapenem resistance using the Carba NP test (13, 14) and polymerase chain reaction (PCR) for blaVIM-, blaIMP-, blaNDM-, blaKPC-, and blaOXA-48-like genes, as described previously (15-17).
3.6. Statistical Analysis
The data were analyzed using the R statistical software (R Development Core Team (2016). R: A language and environment for statistical computing. R Foundation for Statistical Computing, Vienna, Austria). Descriptive statistics were used in the statistical analysis.
4. Results
From 2009 to 2016, a total of 122 total carbapenemase-producing Enterobacteriaceae isolates were identified. There were 73 isolates of KPC pathogens (Klebsiella pneumoniae 71/122, Escherichia coli 1/122, Citrobacter freundii 1/122), mostly in 2009. During the next four years, the number of KPC isolates remained at a constant level (eight or nine per year) and then decreased to three isolates from 2014 to 2015 and one isolate in 2016. The first NDM pathogens in our center were observed in 2015, and the number increased almost four-fold during the next year. Klebsiella pneumoniae (46/122) and K. oxytoca (2/122) were amongst the NMD isolates.
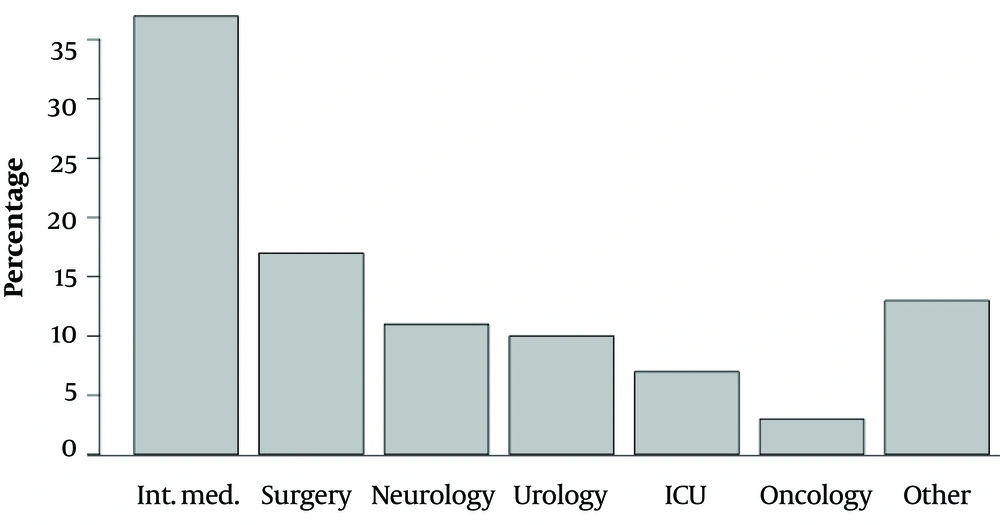
Only one OXA-48 pathogen (E. coli) was isolated in 2015. All isolated carbapenemase-producing Enterobacteriaceae were susceptible to colistin and resistant to ciprofloxacin, cefepime, imipenem, ertapenem, meropenem, piperacillin/tazobactam, and amoxicillin/clavulanate. Escherichia coli OXA-48 was also susceptible to tigecycline and resistant to amikacin, gentamycin, and trimethoprim/sulfamethoxazole. The susceptibility and resistance of KPC, NDM, and OXA-48 to other antimicrobials varied. The number of identified pathogens is presented in Table 1, and the origin of the biological samples is shown in Figure 1. Figure 2 contains the incidence of carbapenemase-producing Enterobacteriaceae species in different departments of the Military Institute of Medicine. The specific mechanisms of resistance to antimicrobials are shown in Table 2. Furthermore, the susceptibility and resistance of the isolated carbapenemase-producing Enterobacteriaceae pathogens to different antimicrobial agents are presented in Table 3.
| Variables | 2009 | 2010 | 2011 | 2012 | 2013 | 2014 | 2015 | 2016 | Total |
|---|---|---|---|---|---|---|---|---|---|
| KPC | 31 | 9 | 9 | 8 | 9 | 3 | 3 | 1 | 73 |
| NDM | - | - | - | - | - | - | 10 | 38 | 48 |
| OXA-48 | - | - | - | - | - | - | 1 | - | 1 |
Abbreviations: KPC, Klebsiella pneumoniae carbapenemase; NDM, New Delhi metallo-β-lactamase; OXA-48, carbapenem-hydrolyzing oxacillinase.
| Pathogen | Number of Isolates | Percentge |
|---|---|---|
| Klebsiella pneumoniae KPC | 71/122 | 58.2 |
| K. pneumoniae NDM | 46/122 | 37.7 |
| K. oxytoca NDM | 2/122 | 1.6 |
| Citrobacter freundi KPC | 1/122 | 0.8 |
| Escherichia coli KPC | 1/122 | 0.8 |
| E. coli OXA-48 | 1/122 | 0.8 |
| Antimicrobial | KPC, n = 73 | NDM, n = 48 | OXA-48, n = 1 | |||
|---|---|---|---|---|---|---|
| Agent | Susceptible | Resistant | Susceptible | Resistant | Susceptible | Resistant |
| Amikacin | 36 (49.3) | 37 (50.7) | 5 (10.4) | 43 (89.6) | 0 (0) | 1 (100) |
| Gentamycin | 38 (52.1) | 35 (47.9) | 25 (52.1) | 23 (47.9) | 0 (0) | 1 (100) |
| Ciprofloxacin | 0 (0) | 73 (100) | 0 (0) | 48 (100) | 0 (0) | 1 (100) |
| Cefepime | 0 (0) | 73 (100) | 0 (0) | 48 (100) | 0 0) | 1 (100) |
| Imipenem | 0 (0) | 73 (100) | 0 (0) | 48 (100) | 0 (0) | 1 (100) |
| Ertapenem | 0 (0) | 73 (100) | 0 (0) | 48 (100) | 0 (0) | 1 (100) |
| Meropenem | 0 (0) | 73 (100) | 0 (0) | 48 (100) | 0 (0) | 1 (100) |
| PIP/TAZO | 0 (0) | 73 (100) | 0 (0) | 48 (100) | 0 (0) | 1 (100) |
| Colistin | 73 (100) | 0 (0) | 48 (100) | 0 (0) | 1 (100) | 0 (0) |
| Tigecycline | 68 (93.2) | 5 (6.8) | 9 (18.8) | 39 (81.2) | 1 (100) | 0 (0) |
| TMP/SMX | 4 (5.5) | 69 (94.5) | 23 (47.9) | 25 (52.1) | 0 (0) | 1 (100) |
| Amox/Clav | 0 (0) | 73 (100) | 0 (0) | 48 (100) | 0 (0) | 1 (100) |
Abbreviations: Amox/clav, amoxicillin/clavulanate; PIP/TAZO, piperacillin/tazobactam; TMP/SMX, trimethoprim/sulfmethoxazole.
aValues are expressed as No. (%).
5. Discussion
The microbiological and epidemiological monitoring of infections caused by carbapenemase-producing Enterobacteriaceae is a routine prophylactic practice. During the eight years of the current analysis, 122 strains of carbapenemase-producing Enterobacteriaceae were isolated from biological materials obtained from patients hospitalized at our center. Among all the isolates, 96% were identified as K. pneumoniae, 2% K. oxytoca, 2% E. coli, and 1% C. freundii. KPC-like resistance was observed in 60% of the analyzed pathogens, NMD-like resistance in 39%, and OXA-48-like resistance in 1%.
The results of the EuSCAPE study (5) showed that, among 2703 Enterobacteriaceae isolates, 85% were K. pneumoniae and 15% Escherichia coli isolates; 37% of K. pneumoniae and 19% of E. coli isolates could produce KPC, NDM, and OXA-48 carbapenemases. These data were slightly different from the current study results. In the current study, KPC-like carbapenemases were observed in 60% of samples, compared with 45% in the EuSCAPE study (5), while NDM was present in 40% of the current study samples, but in only 11% from the EuSCAPE data. These findings indicated that the microbiological situation in our center was different due to the incidence of isolated NDM pathogens, as well as the relatively low number of the available therapeutic options. New antimicrobials, such as cefazolin/tazobactam, ceftazidime/avibactam, ertapenem/meropenem, and sometimes gentamicin are not effective against these pathogens (18-20).
The first isolate of KPC was identified in Warsaw in May 2008 (21).From 2008 to 2016, a total of 907 cases of KPC were documented. The first isolate of E.coli NDM-1 was observed in August 2011 in a 58-year-old male returned to Poland from Congo. He was hospitalized in the intensive care unit of a Warsaw hospital. However, this patient died due to multiple organ failure (6). From 2011 to 2016, a total of 2605 isolates of NMD were identified. Regarding OXA-48 isolates, 53 isolates were observed from 2012 to 2016 and were mainly from abroad. Eighty percent of NDM isolations in Poland occurred during the last few years, and they were mostly obtained from biological samples collected from hospitals in the Warsaw region. This trend explains the significant increase in the detected number of NDM isolates in our center from 10 in 2015 to 38 in 2016.
The first two carbapenemase-producing Enterobacteriaceae pathogens with resistance mechanisms, NDM and OXA-48, were isolated in our center in 2005 from victims of the terrorist attack at the Bardo Museum in Tunis, Tunisia. These patients were initially hospitalized in Tunis (17). All carbapenemase-producing Enterobacteriaceae pathogens were susceptible to colistin. Although some information on pathogen resistance to colistin exists, the current study encountered no resistance. Colistin resistance is threatening because it results from the presence of the plasmid-mediated MCR-1 gene (22). Both a decrease in bacterial resistance to colistin and an increase in the clinical efficacy of this antibiotic may be achieved when colistin is administered in conjunction with tigecycline, fusidic acid, aminoglycosides, and rifampicin (23).
The current study observed that NDM-carbapenemase-producing species were significantly less susceptible to tigecycline compared with KPC and OXA-48-producing pathogens (19% vs. 93% and 100%, respectively; Table 3). The current study results were comparable to those of the SENTRY study (24). About 50% of the KPC isolates, 11% of the NDM pathogens, and none of the OXA-48 isolates were susceptible to aminoglycosides. OXA-48 carbapenemase had the lowest hydrolytic activity against cephalosporins and carbapenems. All carbapenemase-producing Enterobacteriaceae isolates with KPC, NDM, and OXA-48 phenotypes were resistant to carbapenems at breakpoint values > 8 µg/mL for imipenem and meropenem and > 1 µg/mL for ertapenem. At MIC values of ≤ 16 µg/mL and especially ≤ 8 µg/mL, the administration of large doses of carbapenems together with colistin and tigecycline (6 g daily by intravenous infusion followed by a bolus of 2 g) may improve the clinical outcome (25)
5.1. Conclusions
The current study results on carbapenemase-producing Enterobacteriaceae susceptibility to antimicrobial agents were comparable to those of other studies that analyzed the microbiological situation in Europe. The relatively high incidence of NDM pathogens is alarming and may be resulted from the unique epidemiological situation in our region. This finding indicated the importance of identifying pathogen carriers for epidemiological services and also wide microbiological analyses, especially in patients admitted from other hospitals and long-term care facilities or after foreign travel.