1. Background
Salmonella is an important pathogen transmitted through food, water or direct contact with animals. Most Salmonella surveillance programs rely on polymerase chain reaction (PCR)-based assays for rapid and accurate detection (1, 2). Among these molecular tools, the invA-PCR assay has been accepted as the conventional method for Salmonella detection (2-4). This PCR protocol amplifies a fragment of the invA gene, a Salmonella-specific locus (5, 6) proposed as an international standard tool for the accurate detection of this pathogen (7). Nearly, 7,000 scientific reports have used invA PCR assays for Salmonella detection and ~ 450 of them were published in 2018 (as of December) (8). Nevertheless, some reports have described the occurrence of false-positive results (5, 7, 9, 10).
2. Objectives
The main goal of the present study was to evaluate the performance of previously published invA PCR assays using a collection of isolates obtained from a Salmonella surveillance program.
3. Methods
3.1. Ethics Statement
School of Chemistry Biosafety Committee revised and approved the experiments under project #FQ-2017-01.
3.2. Bacterial Isolates
Salmonella enterica type strains (ATCC 140028, ATCC 700720, ATCC 23595, ATCC 14028 and ATCC 13076), Citrobacter spp., Escherichia coli, Enterobacter sp., Serratia sp., Hafnia sp. and Aeromonas sp. isolates were retrieved from our frozen-glycerol stock collection. This collection comprises more than 400 bacterial isolates obtained during a Salmonella surveillance program for poultry meat. All the selected isolates were grown overnight in Tryptic soy broth at 35°C to reach a concentration of ~ 4 × 109 CFU/mL. After incubation, one milliliter of the culture was used for genomic DNA extraction (Quick-DNA Miniprep Plus Kit, Irvine, CA).
3.3. PCR Assay Targeting the invA Gene
Genomic DNA from the selected isolates was subjected to invA PCR amplification using seven previously published primer sets (5, 11-14) and targeting the invA gene (STM2896; Table 1). When the published protocols generated non-specific amplicons, gradient PCR (temperature range: 41 - 64°C) was performed to identify optimum annealing temperatures. Reactions were carried out using maxima hot start Taq DNA polymerase (Thermo Fisher Scientific, Waltham, MA) and 5.0 ng/µL of purified DNA. The optimized PCR protocols consisted of an initial denaturation at 94°C for 3 minutes, 35 cycles of: desaturation at 94°C for 30 secods, annealing at 53 - 69.3°C for 30 seconds (Table 1), extension at 72°C for 30 seconds and a final extension at 72°C for 3 minutes. Specificity of the PCR method was analyzed on 1.5% agarose gel.
| Primer | Sequence (5’ - 3’) | Amplicon Size, bp | Locus | Annealing Temperature, °C | References |
|---|---|---|---|---|---|
| invA1 | CTGTTGAACAACCCATTTGT | 437 | invA | 57.4 | (11) |
| invA2 | CGGATCTCATTAATCAACAAT | ||||
| invAnest1 | AACCAGCAAAGGCGAGCAG | 199 | invA | 65.0 | (11) |
| invAnest2 | GCGCACGCCATAATCAATAAA | ||||
| invA3F | AACGTGTTTCCGTGCGTAAT | 262 | invA | 65.0 | (12) |
| invA3R | TCCATCAAATTAGCGGAGGC | ||||
| SA01 | TATCGTACTGGCGATATTGGTGTTTA | 540 | invA | 65.0 | (13) |
| SA02 | GGACAAATCCATACCATGGCGAGTCA | ||||
| SA03 | GAAATTATCGCCACGTTCGGG | 281 | invA | 65.0 | (13) |
| SA04 | TCATCGCACCGTCAAAGGAAC | ||||
| invA-139 | GTGAAATTATCGCCACGTTCGGGCAA | 284 | invA | 64.0 | (5) |
| invA-141 | TCATCGCACCGTCAAAGGAACC | ||||
| Salm 3 | GCTGCGCGCGAACGGCGAAG | 389 | invA | 65.0 | (14) |
| Salm 4 | TCCCGGCAGAGTTCCCATT | ||||
| 16SF1 | TGTTGTGGTTAATAACCGCA | 574 | 16s rRNA | 57.4 | (15) |
| 16SIII | CACAAATCCATCTCTGGA | ||||
| MINf | ACGGTAACAGGAAGMAG | 402 | 16s rRNA | 53.0 | (16) |
| MINr | TATTAACCACAACACCT | ||||
| STM3098-f2 | TTTGGCGGCGCAGGCGATTC | 423 | STM3098 | 69.3 | (17) |
| STM3098-r2 | GCCTCCGCCTCATCAATCCG | ||||
| ttr-6 | CTCACCAGGAGATTACAACATGG | 86 | ttrA/C | 65.0 | (18) |
| ttr-4 | AGCTCAGACCAAAAGTGACCATC |
PCR Primers Pairs Used in the Present Study, Its Amplicon Size, Targets and Annealing Temperature
3.4. PCR Assay Targeting the 16s rRNA, STM3098, and ttrA/C Genes
To improve the discriminatory power of PCR protocols, alternative Salmonella-specific PCR assays were performed. Four additional primer sets (16SF1 + 16SIII, MINf + MINr, STM3098-f2 + STM3098-r2, and ttr-6 + ttr-4) were evaluated using protocols published elsewhere (15-18) (Table 1). Similarly, gradient PCR was carried out to identify optimum annealing temperatures. PCR protocols were carried out as described above with annealing temperatures described in Table 1. Specificity of the PCR methods was analyzed on 1.5% agarose gel.
4. Results and Discussion
4.1. PCR Assay Targeting the invA Gene
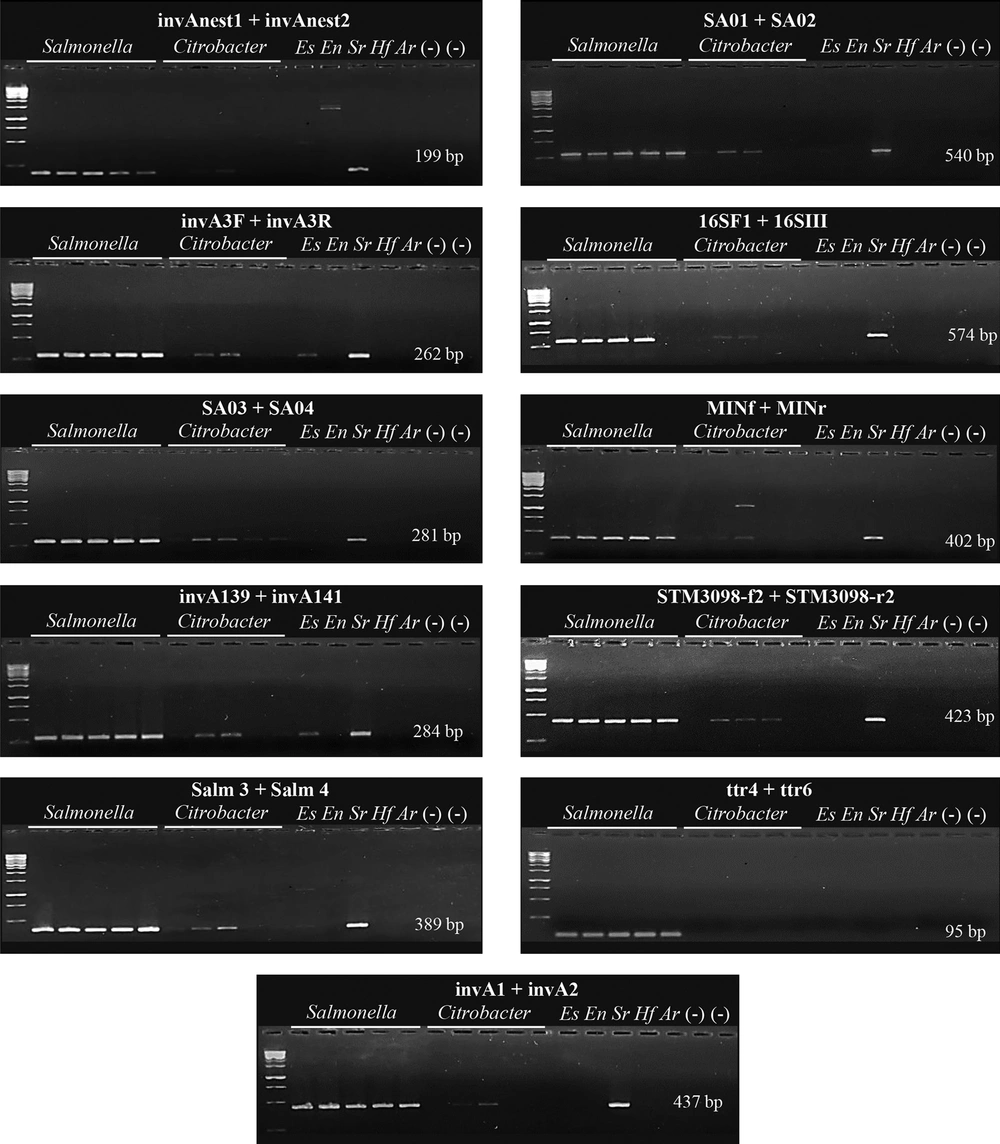
During a Salmonella surveillance program, it was observed that some bacterial isolates generated false-positive signals using the conventional invA PCR assay (5); thus, it was decided to evaluate the performance of other published primers targeting the invA gene. Using a collection of Citrobacter spp., E. coli and Serratia sp. recovered from poultry meat, it was revealed that all the selected invA gene primers generated non-specific signals (Figure 1); comparable results have been reported in reactions containing genomic DNA from non-Salmonella isolates (5, 7, 9, 10). Recent studies have reported a high specificity for invA PCR assays (19, 20); however, these analyses were carried out using DNA obtained from type strain collections. The advantage of the present study was that field isolates known to generate conflictive results were used to evaluate the specificity of the assays. Overall, the results of the present study indicate that PCR assays based on invA gene amplification are not reliable for Salmonella detection.
4.2. Evaluation of Other Salmonella-Specific PCR Assays
Taking advantage of this collection of isolates, the performance of the other four additional primer sets was evaluated. Primers pairs 16SF1 + 16SIII and MINf + MINr targeting the 16S rRNA gene generated non-specific signals in reactions containing Citrobacter spp. and Serratia sp. DNA, even after gradient PCR was performed (Figure 1). This lack of specificity has been reported in other studies (15, 21) and could be caused because primer sets 16SF1 + 16SIII and MINf + MINr target the V3 region of the 16S rRNA, a segment with a high level of homology between members of the Salmonella, Citrobacter and Enterobacter genera (21, 22). Also, the primer set STM3098-f2 + STM3098-r2 targeting locus STM3098, a genomic region coding for a putative transcriptional regulator (17), generated non-specific signals in reactions containing Citrobacter spp. and Serratia sp. DNA (Figure 1). To the best of our knowledge, only one study has evaluated the specificity of this primer set, showing a high specificity against 37 non-Salmonella isolates; however, these isolates belonged to type strain collections (23). These results highlight the importance of using field isolates during PCR protocol validations.
The present study also revealed that the primer set ttr-6 + ttr-4 targeting the ttrA/C genes (tetrathionate reductase subunit A/C) was able to discriminate between S. enterica and non-Salmonella isolates (Figure 1). Comparable results were reported using a set of 110 S. enterica strains, representing 38 different serovars and 87 non-Salmonella strains (18). Importantly, the primer set ttr-6 + ttr-4 has shown to be an excellent molecular target for quantitative assays (e.g., qPCR) due to its high specificity and amplicon size (~ 90 bp) (24, 25).
5. Conclusions
In sum, the results of the present manuscript indicate that PCR assays based on invA gene amplification are not reliable for Salmonella detection. False-positive results are commonly obtained from Citrobacter spp., E. coli and Serratia sp. isolates. Other loci, such as ttrA/C genes, should be used for the accurate and reliable detection of this pathogen.
Performance evaluation of PCR assays for Salmonella detection. Representative PCR reactions using Salmonella enterica type-strains and isolates recovered from poultry meat samples. Eleven primer sets targeting the invA (invA1 + invA2, invAnest1 + invAnest2, invA3F + invA3R, SA01 + SA02, SA03 + SA04, invA-139 + invA-141 and Salm 3 + Salm 4), 16S rRNA (16SF1 + 16SIII and MINf + MINr), STM3098 (STM3098-f2 + STM3098-r2) and ttrA/C (ttr-6 + ttr-4) genes were evaluated against Salmonella enterica type-strains (placed in this order: ATCC 140028, ATCC 700720, ATCC 23595, ATCC 14028 and ATCC 13076), Citrobacter spp., Escherichia coli (Es), Enterobacter sp. (En), Serratia sp. (Sr), Hafnia sp. (Hf), Aeromonas sp. (Ar).