1. Background
Salmonella spp. is one of the most important foodborne pathogens and a major agent of diarrheal complications worldwide. As estimated, it annually accounts for around 1 million cases of salmonellosis and over 450 deaths in the United States (1). It is a zoonotic microorganism (2). Commonly, the main sources of Salmonella infections are contaminated food products, poultry, pigs, contaminated drinking water, ruminants, and direct contact with infected animals (3). Salmonell enteritidis is commonly associated with poultry and poultry derivatives (4). The most common typical symptoms of illness caused by S. enteritidis are fever, vomiting, diarrhea, and abdominal cramps after the consumption of contaminated food products and water (5).
In the US, amongst different species, 36% of Salmonella outbreaks resulted from S. enterica serovar enteritidis (6). Salmonell enteritidis pathogenicity is attributed to its capability to invade non-phagocytic cells and survive and grow inside macrophages and dendritic cells (7). The capability of Salmonella spp. to cause disease can be attributed to various virulence genes found in the either chromosome or large virulence-associated plasmids (8). Different virulence genes of Salmonella, such as viz, sef, pef, spv, and inv are recognized to be involved in adhesion to and invading the host cells. Other virulence genes such as mgtC are associated with the survival of the host cells, and some other genes such as viz, sop, stn, pip A, B, and D have a significant role in real demonstration of pathogenic routes (9, 10).
It seems that selection pressure resulted from the use of antimicrobial drugs promotes growth or as prophylaxis in food-producing animals likely acts as the best selective pressure to develop resistant S. enteritidis (11). Surely, animals carrying resistant microorganisms may act as reservoirs; consequently, resistant strains may be spread from animals to humans via the food chain (12). And due to the similarity of the antibiotic classes used to treat the animals and humans, there is a high risk for antibiotic resistance and forming the multi-drug resistant (MDR) isolates (13). Regarding drug resistance, the role of antimicrobial drugs misuse and overuse in the emergence of antimicrobial resistance cannot be ignored (14).
Biofilm formation is one of the characteristics of these microorganism that intensifies the antibiotic resistance dilemma (15). Nowadays, the biofilm formation on equipment and tools is considered as a source of pathogenic bacteria that increases the risk of food product contamination in food processing systems (16). Biofilms protect bacteria from being damaged by environmental agents such as disinfectants and antibiotics; therefore, the bacteria can develop antibiotic resistance that is difficult to eliminate (17). Due to the high prevalence of resistant isolates recovered from poultry and domestic animals, and the possibility of their transmission through the food chain to humans, it seems to be necessary to evaluate the antibiotic resistance and determine the relationship between biofilm formation and antimicrobial resistance.
2. Objectives
The current study aimed at evaluating molecular detection, virulence genes, biofilm formation, and antibiotic resistance of S. enterica serotype enteritidis recovered from clinical and poultry strains.
3. Methods
3.1. Ethics Statement
The Ethics Committee of Pasteur Institute of Iran approved the current study protocol (IR.pII.REC.1394.83).
3.2. Isolates
The current study was conducted on 282 isolates including 165 isolates recovered from chicken meat, 30 isolates from live poultry feces, 31 isolates from eggs, and 36 isolates from human feces. The isolates were collected from five provinces including Tehran (n = 261), Mazandaran (n = 2), East Azerbaijan (n = 15), Qom (n = 1), and Sistan and Baluchistan (n = 3). The isolates were cultured on the TSI (triple sugar iron) agar (Himedia, India) and transferred to the Central Veterinary Laboratory of Iran Veterinary Organization (CVL-IVO) under standard conditions. Initially, several colonies were picked up from TSI, inoculated into a nutrient broth, and placed in an incubator at 37°C overnight. Afterward, the bacterial isolates were identified through culturing on xylose lysine deoxycholate (XLD) agar and Hektoen enteric (HE) agar (Himedia, India); the procedure was conducted according to ISO 6579 and Italian National Reference Centers (IZSVE) protocols. Then, Salmonella genus was determined using antiserum. Agglutination in polyvalent antisera and biochemical tests were used for final genus/species-level identification of Salmonella spp. (18). Finally, Luria-Bertani (LB) broth (Himedia, India) was used to culture the confirmed isolates.
3.3. Detection of Salmonella spp. Using Triplex Polymerase Chain Reaction and Differentiation of S. enterica Serotype enteritidis from Other Salmonella spp.
3.3.1. Extraction of DNA
High pure PCR template preparation kit (Roche Company, REF: 11796828001) was used to extract DNA. Two hundred microliters of broth containing the cultured isolates were used, as recommended by the manufacturer’s manual.
3.3.2. Triplex PCR Performance
In order to perform PCR assay, three primer pairs were used. The invA primer (199 bp) with sequences of F-invA: AAA CGT TGA AAA ACT GAG GA, and R-invA: TCG TCA TTC CAT TAC CTA CC (19) was used to detect Salmonella genus. A tyv primer (614 bp) with sequences of F-tyv: GAGGAAGGGAAATGAAGCTTTT, and R- tyv: TAGCAAACTGTCTCCCACCATAC (20) was used to detect serogroup D. Finally, both of the two primers were used to confirm the findings in each culture and determine the serogroup with antiserum (Mast). Furthermore, sdf primer (299 bp) with sequences of F- sdf: AAA TGT GTT TTA TCT GAT GCA AGA GG and R- sdf: GTT CGT TCT TCT GGT ACT TAC GAT GAC was used to detect the serovar (20). The strains positive for all the three genes were considered as S. enterica serotype enteritidis.
Each PCR reaction contained 2.5 μL of 10X PCR buffer (Qiagen, Germany), 3 μL of DNA template, 1.5 μL of MgCl2 (25 mM) (Qiagen, Germany), 1.2 μL of dNTP (10 Mm) (Qiagen, Germany), 1.5 μL of each of the forward and reverse primers of invA (10 μM) (MWG Company, Germany), 0.75 μL of each of the forward and reverse primers of Sdf (10 μM) (MWG Company, Germany), 0.75 μL of each of the forward and reverse primers of tyv (10 mM) (MWG Company, Germany), 0.2 μL of hot start Taq DNA polymerase [(5 U/μL) (Qiagen, Germany)], and 10.5 μL of distilled water in a total volume of 25 μL. The following steps were taken to carry out PCR reaction: initial denaturation (95°C for 10 minutes), denaturation (94°C for 60 seconds, 30 cycles), annealing step (60°C for 90 seconds), extension step (72°C for 90 seconds), and final extension (72°C for 10 minutes, one cycle). The PCR products were analyzed on a 1.5% (w/v) agarose gel electrophoresis comprising SYBR® Safe (Qiagen, Germany) using the standard protocol. DNA ladder 100 bp (Qiagen, Germany) was used as a marker.
3.4. Antibiotic Resistance Testing
All S. enterica serotype enteritidis isolates were studied via Kirby-Bauer disk diffusion method to evaluate their resistance to antibiotic disks. The criteria proposed by the National Committee for Clinical Laboratory Standards was used to determine susceptibility rates (21). Antibiotic disks used in the current study were purchased from MAST, UK. The results of the tests were categorized into three classes of resistant, intermediate, and sensitive. Escherichia coli ATCC 35218 and ATCC 25922 were used as reference strains for quality control. The antibiotics used in the current study were as follows: nalidixic acid (30 µg), amoxicillin (30 µg), amoxicillin + clavulanic acid (20/10 µg), cefoxitin (30 µg), colistin (10 µg), chloramphenicol (30 µg), ciprofloxacin (5 µg), cefalotin (30 µg), enrofloxacin (5 µg), ceftazidime (30 μg), ceftriaxone (30 µg), cefepime (10 µg), kanamycin (30 µg), ampicillin (10 µg), neomycin (10 µg), ertapenem (10 µg), streptomycin (10µg), chloramphenicol (30 µg), trimethoprim + sulfamethoxazole (23.75/1.75 µg), tetracycline (30 μg), imipenem (10 µg), and meropenem (10 µg).
3.5. Primer Design, PCR Amplification Conditions, and Detection of Virulence Genes
As mentioned earlier, sdiA, spvC, ratA, lpfC, sitC, sifA, sopB, spiA, sipA, invA, and hilA virulence genes primer sequences were expected to form the desired products by mPCR technique on the DNA extracted from S. enterica serotype enteritidis isolates (22). The specificity of the primers was approved by the Blast program of GenBank database. The primers were designed using Gene Runner software and synthesized by MWG Company, Germany. The sequences of virulence genes primers are presented in Table 1.
| Target Gene, Primer Name | Sequences (5' to 3') | Annealing Temperature (ºC) | PCR Product Size (bp) | Reference |
|---|---|---|---|---|
| sdiA | 53 | 274 | (23) | |
| sdiA-F | AATATCGCTTCGTACCAC | |||
| sdiA-R | GTAGGTAAACGAGGAGCAG | |||
| spvC | 53 | 571 | (23) | |
| spvC-F | ACTCCTTGCACAACCAAATGCGGA | |||
| spvC-R | TGTCTCTGCATTTCGCCACCATCA | |||
| ratA | 56 | 243 | (24) | |
| ratA-F | GACGTCGCTGCCGTCGTACC | |||
| ratA-R | TACAGCGAACATGCGGGCGG | |||
| lpfC | 64 | 641 | (3) | |
| lpfC-F | GCC CCG CCT GAA GCC TGT GTT GC | |||
| lpfC-R | AGG TCG CCG CTG TTT GAG GTT GGA TA | |||
| sitC | 60 | 768 | (3) | |
| sitC-F | CAG TAT GCT CAA CGC GAT GTC TCC | |||
| sitC-R | CGG GGC GAA AAT AAA GGC TGT GAT GAA C | |||
| sifA | 60 | 449 | (3) | |
| sifA-F | TTT GCC GAA CGC GCC CCC CAC AGC | |||
| sifA-R | GTT GCC TTT TCT TGC GCT TTC CAC CCA TCT | |||
| sopB | 60 | 220 | (3) | |
| sopB-F | CGG ACC GGC CAG CAA CAA AAC AAG AAG | |||
| sopB-R | TAG TGA TGC CCG TTA TGC GTG AGT GTA TT | |||
| spiA | 60 | 550 | (3) | |
| spiA-F | CCAGGGGTCGTTAGTGTATTGCGTGAGATG | |||
| spiA-R | CGCGTAACAAAGAACCCGTAGTGATGGATT | |||
| sipA | 60 | 875 | (3) | |
| sip A-F | GGACGCCGCCCGGGAAAAACTCTC | |||
| sip A-R | ACACTCCCGTCGCCGCCTTCACAA | |||
| hilA | 52 | 497 | (25) | |
| hilA2-up | CTG CCG CAG TGT TAA GGA TA | |||
| hilA2-down | CTG TCG CCT TAA TCG CAT GT | |||
| invA | 58 | 796 | (26) | |
| invA-F | CGG TGG TTT TAA GCG TAC TCT T | |||
| invA-R | CGA ATA TGC TCC ACA AGG TTA | |||
| invA | 60 | 199 | (26) | |
| AAA CGT TGA AAA ACT GAG GA | ||||
| TCG TCA TTC CAT TAC CTA CC |
PCR assay was carried out in a DNA Thermocycler (Model CP2-003, Corbett, Australia) as follows: one cycle of 95°C for 5 minutes, followed by 35 cycles of denaturation (95°C for one minute), annealing step [(sdiA and spvC (53°C), ratA (56°C), lpfC (64°C), sitC, sifA, sopB, and spiA (60°C), hilA (52°C), and invA (58°C)] (incubation time for all genes in the annealing step were 40 seconds), and elongation step at 72°C for 30 seconds and 10 minutes of final elongation at 72°C. The total PCR reaction (25 μL) included 2.5 μL of 10X Buffer, 14.7 μL of distilled water, 1 μL of dNTP (10 mM), 1.5 μL of MgCl2 (25 mM), 0.3 of hot start Taq DNA polymerase (5 U/μL), 1 μL (10 μM) of each of the forward and revers primers, and 3 μL of the template DNA. Electrophoresis was performed on 1.5% agarose gel subjected to 140 V for one hour to visualize PCR product bands later analyzed using GeneTools and SnapGene software.
3.6. Biofilm Formation
For quantification of biofilm formation 96-well microtiter plates were used; biofilm formation was measured in line with a previous study (27). Firstly, 230 μL of the nutrient broth medium (Himedia, India) was poured into wells of a sterile 96-well flat-bottomed polystyrene microplate. Three wells were allocated to each sample and the negative control (wells contained broth only). About 20 μL of bacterial culture (grown overnight) was poured into each well of the microplates. Afterward, incubation of the plates was performed overnight at 35ºC. The content of the microplates was discarded; the wells were washed three times using 300 μL of PBS. Then, 250 μL of methanol was added to each well to fix the microorganisms attached to the surface of microplates. After 15 minutes, the content of microplates was taken out and air dried at 25ºC.
The microplates were stained using Gram staining with 250 μL of Crystal violet per well for 5 minutes. To wash the extra stain, microplates were put under running tap water. Finally, microplates were air dried, and about 250 μL of 33% (v/v) glacial acetic acid was poured into each well to wash the bound dye. An automated Multiskan ELX808 reader (Labsystems, USA) at 570 nm and software (Gene5) were used to measure the optical density (OD) of wells. Taking into account the optical density (OD) of each well, isolates were categorized into four groups: these groups were isolates with a weak, moderate, or without the potential to produce the biofilm. The other group included isolates with a potential of producing a strong biofilm. In brief, optical density cut-off value (ODc) was equal to the average OD of negative control plus 3 x standard deviation (SD) of the negative control. The tested isolates were categorized as follows: OD ≤ ODc: non-adherent; ODc < OD ≤ (2 × ODc): weak biofilm producers; 2 × ODc < O.D ≤ 4 × ODc: moderate biofilm producers; and 4 × ODc < OD: strong biofilm producers. The obtained results are presented as means and standard deviation (SD).
3.7. Statistical Analysis of Data
The data were then analyzed with SPSS version 16.0 (Chicago, Illinois, USA). Consensus tables and Chi-square test were used to evaluate the correlation between biofilm formation potential and antibiotic resistance. P values < 0.05 were considered statistically significant.
| Sample (%) | Antibiotic | |||||||||||||||||||||
|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|
| Fox | AM | Aug | C | IMI | MEM | ATM | CIP | CAZ | CAZ+ Clav | CRO | CPM | ETP | P | AM | FM | LVX | GM | K | CO | T | NA | |
| Human | 47.1 | 20.6 | 14.7 | 35.3 | 14.7 | 2.9 | 2.9 | 23.5 | 2.9 | - | - | - | - | 94.1 | 14.7 | 50 | - | 11.8 | 47.1 | 91.2 | 91.2 | 88.2 |
| Chicken | 7.1 | - | - | 42.9 | - | - | - | 64.3 | 7.1 | - | 7.1 | 7.1 | - | 100 | - | 92.9 | 7.1 | 42.9 | 50 | 50 | 14.3 | 78.6 |
| Poultry | - | - | - | 5.9 | - | - | - | 11.8 | 5.9 | - | - | - | - | 70.6 | AM | 17.6 | - | - | - | - | - | 94.1 |
Abbreviations: Fox, cefoxitine; A, amoxicillin; ATM, aztreonam; Aug, augmentin; C, colistin; CAZ, ceftazidime; Clav, clavelanic acid; CO, chloramphenicol; CPM, cefepime; IM, imipenem; MEM, meropenem; CIP, ciprofloxacin; CRO, ceftriaxone; EP, ertapenem; K, kanamycin; NA, nalidixic Acid; Ne, neomycin; P, penicillin; TMP/SMX, trimethoprim/sulfamethoxazole; T/S, trimethoprim-sulfamethoxazole.
4. Results
4.1. Identification of Salmonella enterica Serotype enteritidis
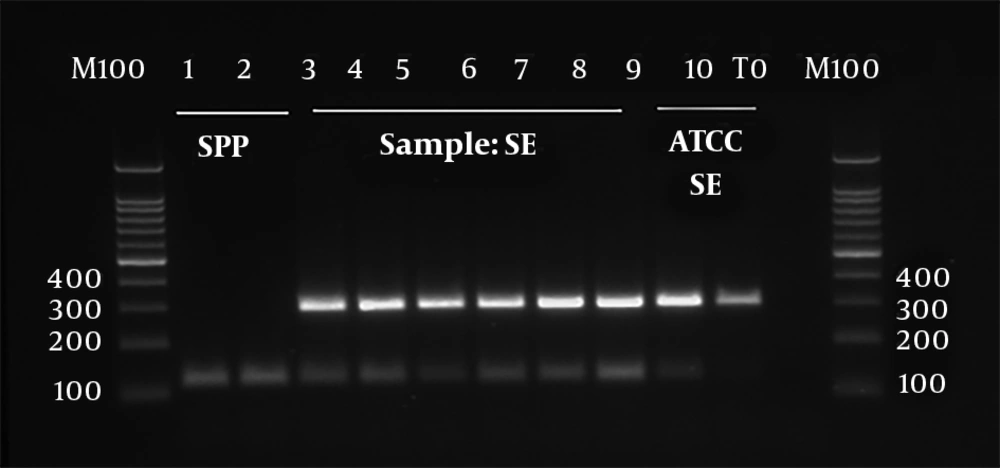
Genus, serogroup, and serovar were determined using triplex-PCR. The invA, tyv, and sdf genes were used to detect Salmonella genus, serogroup D, and serovar, respectively. All the tested isolates were positive for all the three genes and were identified as S. enterica serotype enteritidis (Figure 1).
4.2. Results of the Analysis of Virulence Genes in Salmonella enterica Serotype enteritidis
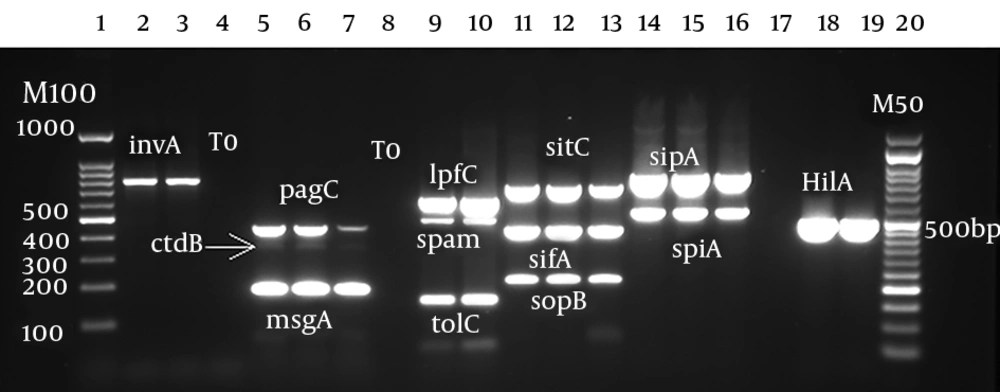
Analysis of virulence genes showed that the prevalence of invA, sdiA, hilA, and ratA was 100% (n = 282). The prevalence of other genes was as follows: sopB (n = 281, 99.6%), sitC and sifA (n = 275, 97.9%), and spiA and sipB (n = 280, 99.3%). The lowest prevalence was observed in spvC (n = 106, 37.6%) (Figure 2).
Virulence genes detection by multiplex PCR. Lane 1: marker 100 bp; lanes 2 and 3: invA (780 bp); lanes 4, 8, and 17: negative control; lanes 5, 6, and 7: pag (454 bp), cdtB (268 bp), and msgA (189 bp); lanes 9 and 10: ipfC (641 bp), span (504 bp), and tolC (161 bp); lanes 11, 12, and 13: sitC (768 bp), sifA (449 bp), and sopB (220 bp); lanes 14, 15, and 16: sipA (875 bp) and spiA (550 bp); lanes 18 and 19: hilA (497 bp), lane 20: marker 50 bp.
4.3. Results of Susceptibility Testing
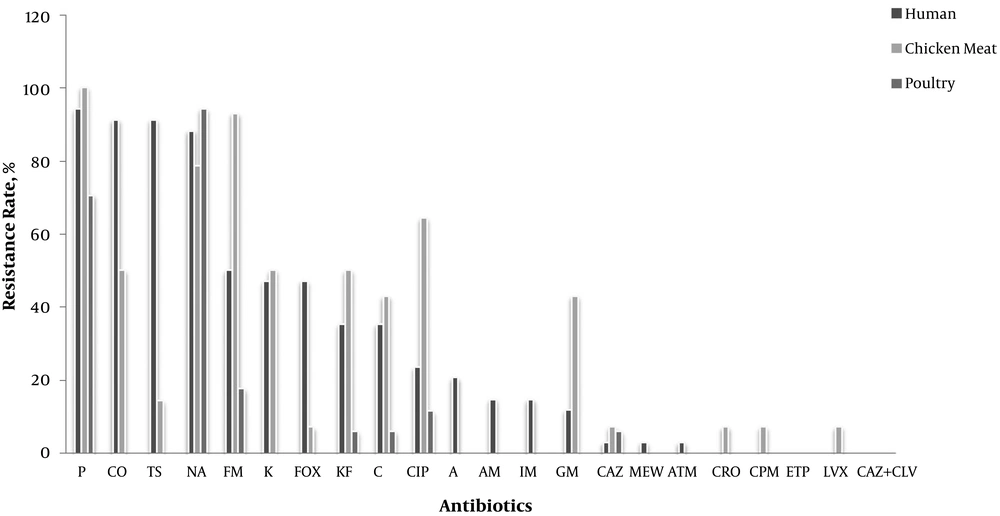
Of all strains, 67% were MDR, of which 51.7% were recovered from humans and the remaining 15.3% from chicken meat. No species was isolated from live poultry. Only five isolates were resistant to one antibiotic; in addition, one strain was resistant to two antibiotics, and the other isolates were resistant to three or more antibiotics. The isolates recovered from human samples were mainly resistant to penicillin, followed by colistin, and trimethoprim-sulfamethoxazole with a prevalence of 94%, 91.2%, and 91.2%, respectively. Furthermore, there was no resistance to ciprofloxacin, levofloxacin, ertapenem, and cefepime. The isolates recovered from live poultry were mainly resistant to nalidixic acid ad penicillin, with a prevalence of 94.1% and 70.6%, respectively; however, there was no significant resistance to other antibiotics. Finally, the isolates recovered from chicken meat were mainly resistant to nitrofurantoin and penicillin, with a prevalence of 100%; however, they showed no resistance to ampicillin, amoxicillin, and meropenem (Figure 3).
The total results of antibiotic resistance in live poultry, human, and chicken meat. A, Amoxicillin; C, colistin; CAZ, ceftazidime; CIP, ciprofloxacin; CO, chloramphenicol; CPM, cefepime; CRO, ceftriaxone; EP, ertapenem; FM, framycetin; Fox, cefoxitine; GM, gentamicin; IM, imipenem; K, kanamycin; KF, cefalotine; LVX, levofloxacin; MEM, meropenem; NA, nalidixic acid; NIT, nitrofurantoin; P, penicillin; TMP/SMX, trimethoprim/sulfamethoxazole; T/S, trimethoprim-sulfamethoxazole.
4.4. Results of Biofilm Formation
According to the results of biofilm formation test, 98 (34.5%) isolates were strong biofilm producers; in addition, 168 (59.6%) and 16 (5.7%) isolates were moderate and weak biofilm producers, respectively. As shown in Table 3, there was a statistically significant correlation between strong biofilm formation and resistance to colistin (P = 0.001), ceftazidime (P = 0.04), penicillin (P = 0.001), gentamicin (P = 0.04), chloramphenicol (P = 0.001), trimethoprim (P = 0.001), and trimethoprim-sulfamethoxazole (P = 0.001).
| Antibiotic | Sensitive (%) | Intermediate (%) | Resistant (%) | P Value |
|---|---|---|---|---|
| Fox | 64.6 ± 0.07 | 9.2 ± 1.14 | 26.2 ± 3.29 | 0.075 |
| A | 81.5 ± 2.1 | 7.6 ± 0.11 | 10.7 ± 1.2 | 0.174 |
| C | 44.1 ± 0.111 | 26.1 ± 0.06 | 29.2 ± 0.111 | 0.02 |
| IM | 81.5 ± 4.5 | 10.8 ± 0.08 | 7.7 ± 1.3 | 0.174 |
| MEM | 98.4 ± 5.7 | 0 ± 0.01 | 1.6 ± 0.67 | 0.674 |
| CIP | 0 ± 0.01 | 70.7 ± 4.2 | 29.3 ± 2.34 | 0.138 |
| CAZ | 90.7 ± 4.56 | 4.6 ± 0.06 | 4.6 ± 0.06 | 0.04 |
| CRO | 98.4 ± 3.23 | 0 ± 0.001 | 1.6 ± 0.95 | 0.24 |
| CPM | 96.9 ± 3.1 | 1.5 ± 0.08 | 1.5 ± 0.21 | 0.214 |
| P | 9.2 ± 0.08 | 1.1 ± 0.5 | 89.2 ± 3.1 | 0.001 |
| FM | 32.3 ± 2.3 | 16.9 ± 3.32 | 50.7 ± 3.76 | 0.537 |
| AM | 76.9 ± 0.07 | 4.6 ± 0.9 | 18.4 ± 0.07 | 0.966 |
| LVX | 98.4 ± 5.6 | 0 ± 0.001 | 1.6 ± 1.1 | 0.24 |
| GM | 60 ± 3.7 | 24.6 ± 2.1 | 15.4 ± 1.3 | 0.04 |
| NA | 1.5 ± 02.1 | 10.7 ± 1.32 | 87.6 ± 3.33 | 0.602 |
| TMP/SMX | 49.2 ± 0.95 | 0 ± 0.01 | 50.8 ± 2.13 | 0.001 |
| Co | 41.5 ± 0.06 | 0 ± 0.002 | 58.5 ± 3.89 | 0.001 |
| ATM | 98.4 ± 4.1 | 0 ± 0.01 | 1.6 ± 0.11 | 0.674 |
| EP | 98.4 ± 1.14 | 0 ± 0.02 | 1.60 ± 1.11 | 0.674 |
| T | 49.2 ± 0.06 | 0 ± 0.002 | 50 ± 5.1 | 0.001 |
Abbreviations: Fox, cefoxitine; A, amoxicillin; C, colistin; CAZ, ceftazidime; CIP, ciprofloxacin; CO, chloramphenicol; CPM, cefepime; CRO, ciprofloxacin; EP, ertapenem; FM, framycetin; GM, gentamicin; IM, imipenem; MEM, meropenem; NA, nalidixic acid; P, penicillin; TMP/SMX, trimethoprim/sulfamethoxazole; T/S, trimethoprim-sulfamethoxazole.
5. Discussion
The results of virulence genes analysis showed that all the isolates (100%) were positive for invA, sdiA, hilA, and ratA. Moreover, the lowest prevalence (37.6%) was observed in spvC. In the current study, S. enteritidis isolated from poultry, chicken meat, and eggs had the virulence genes similar to the ones present in isolates recovered from human feces. The results indicated that isolates recovered from poultry, chicken meat, and eggs were human pathogens. This finding was also reported by Akiyama et al., since they stated that the isolates recovered from the environmental specimens carried the genes similar to those harbored by clinical strains, which contributed to human pathogenesis (28). In line with the current study findings, a study conducted by Mezal et al., showed that all isolates were positive for most of the virulence genes found in S. enterica (3).
In the current study, 67% of all the strains were MDR, of which 51.7% were recovered from humans and the remaining 15.3% were isolated from chicken meat and eggs; furthermore, no isolate was recovered from live poultry. Moreover, five strains were resistant to only one antibiotic, one strain was resistant to two antibiotics, and the remaining were resistant to three or more antibiotics. Nowadays, it is believed that animals and their products are the main sources of MDR Salmonella species; the resistance first emerges in animals, before transmitting to humans via the food chain (29). Surprisingly, no resistant strains were retrieved from poultry. On the other hand, the resistant strains were mainly isolated from human and partly from chicken and the eggs. This finding might be attributed to the correct administration of antibiotics for chickens. Our findings were inconsistent with those demonstrating that the increase of antimicrobial resistance amongst food-borne bacteria in the recent decades might be attributed to the selection pressure produced by the unsuitable or unrestricted use of antibiotics in veterinary medicine (30).
Overall, there was no resistance to ertapenem and ceftazidime-clavulanic acid; on the other hand, the resistance to penicillin and nalidixic acid was the most common type. Resistance to nalidixic acid in S. enterica serovar enteritidis recovered from different sources was high, and the prevalence rates were as follows: human isolates (88.2%), poultry (94.1%), and chicken/eggs (78.6%). The resistance to nalidixic acid in the current study was higher than those reported by other studies in Brazil (21.5%) and Europe (26%) (31). To the best of the author's knowledge, fluoroquinolones are widely used in veterinary for therapeutic purposes; they are also selectively used for mutant Salmonella isolates resistant to nalidixic acid or for cases with decreased susceptibility to fluoroquinolones (32, 33). As mentioned in the results section, all the isolates showed a high susceptibility to ceftazidime-clavulanic acid (100%), ceftriaxone (92.1%), ceftazidime (94.7%), and cefepime (7.1%), which may indicate the limited and controlled administration of such antibiotics in poultry and animal food products in Iran. These results were inconsistent with those reported from Turkey (34); but in agreement with the current study findings, a study conducted in Brazil showed the resistance about 88.8% to nalidixic acid (35). Also, Ghasemmahdi et al. showed the resistance (97%) to nalidixic acid (36).
The current study aimed at investigating the correlation between biofilm formation potential and antibiotic resistance of isolates species, if any. Based on the results of biofilm formation assessments, 34.5% of the isolates were strong biofilm producers, while 59.6% and 5.7% were moderate and weak biofilm producers, respectively. In line with our findings, a study conducted by Sereno et al., reported that 72.7% of all the isolates could form biofilms, although 13.6%, 59.1%, and 34% were moderate, weak, and strong biofilm producers, respectively. Nonetheless, 27.3% did not form biofilms (15) and inconsistent with the current study, they found no strong biofilm producers. Furthermore, there was a statistically significant correlation between strong biofilm formation and resistance to colistin, ceftazidime, penicillin, gentamicin, chloramphenicol, trimethoprim, and trimethoprim-sulfamethoxazole. However, contrary to the current study findings, a research conducted by Reza et al., reported that biofilm formation potential could not change the antibiotic resistance profile in the microorganisms (37).
In general, owing to the administration of antibiotics to improve growth and prevent diseases in food animals, the augmentation of human salmonellosis started with the emergence of foodborne MDR Salmonella (23, 38). The information acquired from various studies could be utilized to develop better strategies to eliminate biofilms in the food chain, and accordingly, reduce Salmonella contamination in animal/food chain. Furthermore, to ensure the effectiveness of control programs, it is necessary to constantly monitor antibiogram results reported worldwide.
5.1. Conclusions
The presented results showed that more than half of all the assessed strains were MDR. Moreover, there was a significant correlation between biofilm formation potential and antibiotic resistance. Due to the isolation of MDR strains both from clinical and chicken/egg samples, and the likely transmission of resistant species through food chain to human, it seems that there was a significant correlation between biofilm formation and antibiotic resistance; thus, control programs are needed to prevent uncontrollable outbreaks.