1. Background
In the patients hospitalized in the ICU, hospitalized pneumonia is the most prevalent hospital infection which can be accompanied by mortality in 20 to 70% of the cases [1]. Rate of appearance of the pneumonia in the ICU has been reported in a range from 10% to 20% in the various studies [2]. Mechanical ventilation increases risk of occurrence and progression of pneumonia from 7 to 21 times [3]. VAP is a subset of the hospitalized pneumonia and referred to as infection of the pulmonary tissue which has been established within 48 hours or higher after intubation and connecting to mechanical ventilation system [4].
In the patients who are under mechanical ventilation, several factors can be led to the appearance of this infection. Accumulation of the pharyngeal secretions in the back of the splint pipe’s Kaf causing unclear aspirations, destruction of coughing reflex, dysfunction in the performance of cilia of aerial ways, lack of mobility and placement in the lying down situation for longtime, necessity for usage of suction for discharging of Secretions and existence of the resistant to-antibiotics organisms transferred to the patients through the personnel’s hands are the most significant factors in occurrence of this infection.
Many Gram-negative species can be replaced in the pharynx within 48 hours following the hospitalization [5]. Results of the studies show that, in majority of patients, VAP is created by the organisms such as S. aureus, H. influenza, S. pneumonia, P. aeroginosa and Entrobacter [6].
2. Objectives
In fact, they are the highest factors causing appearance of VAP. With dim of determination of frequency of the bacterial factors causing VAP in the patients hospitalized in the ICU of the Tonekabon Shahid Rajaei hospital and identification of the factors causing it’s appearance the current study was conducted.
3. Methods
3.1. Sampling Method
In this study, 35 individuals of the patients hospitalized in the hospital’s ICU with VAP were studied. Using Nedathol, a little quantity of the pulmonary secretions was sucked; then, 3 cm of the Nedathol’s end was cut under sterile conditions by sterile scissors and placed in the pipes containing Brain Heart Infusion Broth culture medium enriched by blood, numbering for each patient. After finishing the sampling, the pipes were transferred to laboratory and incubated in 37°C for 48 hours.
Following the incubation time, the pipes became vortex firstly; then, on the sterile conditions, a few loops of the microbic suspension were cultured in the spread form on the plate containing Brain Heart infusion agar. Later, the plates were incubated again in 37°C for 48 hours.
3.2. DNA Extraction and PCR
After passage of incubation time and observance of the bacteria’s colony, DNA extraction was carried out by use of the phenol- chloroform method [7]. For the purpose of amplification of universal 16s rRNA gene in the bacteria, primers 8f and 1492R” extracted from the scientific article were used. The sequence of forward and reverse primers were 5’-AGAGTTTGATCCTGGCTCAG-3’ and 5’-GGTTACCTTGTTACGACTT -3’, respectively [8].
Each reaction was performed in a total volume of 25 µL, which contained 14.5 µL of molecular biology-grade water (Sigma Aldrich Company LTD., USA), 2.5 µL of 10 × PCR buffer (Promega, USA), 1 µL of 10 pmol of forward and reverse PCR primers, 1 µL of 10 mM dNTPs (Promega, USA), 0.5 µLof smart taq DNA polymerase (Promega, USA), 1 µL of 50 mM MgCl2 (Promega, USA) and 5 µLof DNA template. The negative control tube contained the same PCR reagents as above but had 5 µL of water substituted for the DNA template.
PCR amplification conditions on thermocycler (eppendrof-Germany) were as follows: 95°C for 5 min, followed by 35 cycles of 95°C for 75 s, 56°C for 45 seconds, and 72°C for 45 seconds, with a final extension at 72°C for 5 minutes and storage at 4°C.
An aliquot of all PCR products was run on a 1.5% (w/v) agarose gels with a 100 bp DNA ladder (Fermntas-Russia) and electrophoresed at 75 V for 40 minutes. The bands were visualized using ethidium bromide staining and photographed after UV treatment by a transilluminator (UV doc, England).
3.3. 16S rRNA Gene Sequencing
16s rRNA PCR products were sent to macrogen in South Korea (http://www.macrogen.com/) for DNA sequencing.
3.4. Bioinformatics Applications
All sequences data was subjected to BLAST analysis (ncbi.nlm.nih.gov/BLAST/) to definitively identify each respective 16S rRNA gene amplicon.
4. Results
After one- Year study in the ICU, 35 patients have been studied totally out of which 16 patients (45.7%) are women and 19 patients (54.3%) are men, and age range of the patients in from 15 to 88 years which 10 individuals (28.6%) were placed in under 50 years age group, 15 individuals (42.8%) were place in 50 year to 70 year age group and 10 individuals (28.6%) were placed in higher than 70 years age group. After confirmation of existence of VAP, the patients were entered into the study, and sampling from splint pipe was accomplished out of which 7 individuals (20%) were died for this reason. Dispersion of causes of hospitalization in the studied samples, respectively, includes: 4 individuals (11.4%) Decrease of alertness level, 2 individuals (5.7%) pneumonia, 5 individuals (14.2%) Accident, 2 individuals (5.7%) Respiratory failure, 3 individuals (8.5%) cardiac arrest, 3 individuals (8.5%) pulmonary infection, 2 individuals (5.7%) Septicemia and enteric infection, prostatectomy, stroke, vesical infection, internal infection, diabetes, hepato cellular carcinoma, cerebral hemorrhage, gastric ulcer, Poisoning and distress each of which is 1 undivided (2.8%) in number. Totally, of 35 under study patients, 53 strains of bacterium were isolated out of which 28 bacteria (52.8%) were gram-positive and 25 bacteria (47.2%) were Gram negative. Among verities after Gram positive bacteria 11 strains (20.7%) were Staphylococcus aureus, 4 strains (7.5%) were Corynebacterium, 3 strains (5.6%) were Enterococcus faecalis, 2 strains (3.7%) were Enterococcus faecium, 2 strains (3.7%) were Staphylococcus haemolyticus, 2 strains (3.7%) were Staphylococcus hominis, 2 strains (3.7%) were Staphylococcus lugdunensis, 1 strain (1.8%) was Staphylococcus epidermis and 1 strain (1.8%) was Staphylococcus intermedius.
Among the Gram- negative bacilli, 7 strains (13.2%) were Klebsiella pneumonia, 6 strains (11.3%) were Pseudomonas aeruginosa, 5 strains (9.4%) were Actinobacter baumannii, 2 strains (3.7%) were Alcaligenes faecalis, 2 strains (3.7%) were Enterobactercloacae, and Enterobacter aeruginosa, Escherichia coli and Morganella morganii each of which is one strain.
| Name of Isolated Bacteria | Number of Isolated |
|---|---|
| Staphylococcus aureus | 11 (20.7) |
| Klebsiella pneumonia | 7 (13.2) |
| Pseudomonas aeruginosa | 6 (11.3) |
| Actinobacter baumannii | 5 (9.4) |
| Corynebacterium | 4 (7.5) |
| Enterococcus faecalis | 3 (5.6) |
| Enterococcus faecium | 2 (3.7) |
| Staphylococcus haemolyticus | 2 (3.7) |
| Staphylococcus hominis | 2 (3.7) |
| Staphylococcus lugdunensis | 2 (3.7) |
| Alcaligenes faecalis | 2 (3.7) |
| Enterobacter cloacae | 2 (3.7) |
| Staphylococcus epidermis | 1 (1.8) |
| Staphylococcus intermedius | 1 (1.8) |
| Enterobacter aeruginosa | 1 (1.8) |
| Escherichia coli | 1 (1.8) |
| Morganella morganii | 1 (1.8) |
| Total | 53 (100) |
aValues are expressed as No. (%).
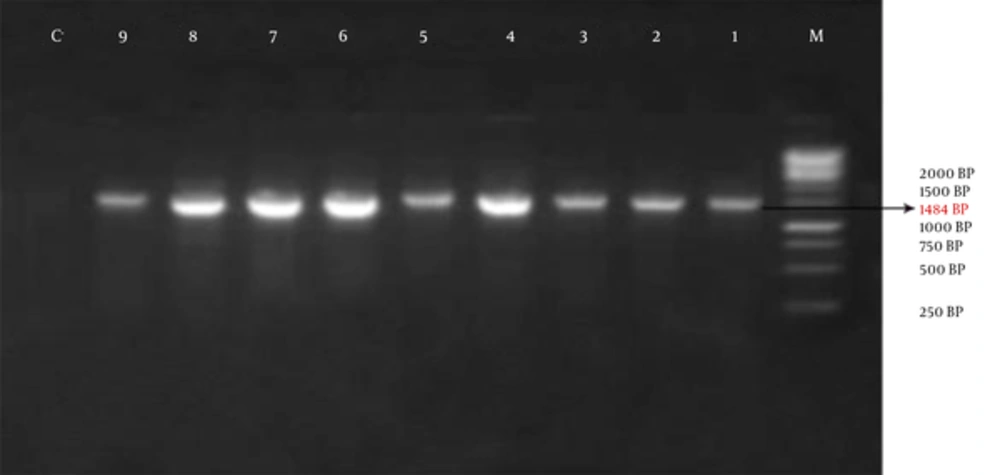
The obtained information showed that, out of 35 studied patients, only one bacterium was isolated from 14 patients (40%) and 2 types of the bacterium were isolated from the other 21 patients (60%). 1484 bp band on the Agarose gel shows amplification of the 16s rRNA gene (Figure 1).
5. Discussion
Ventilator associated pneumonia (VAP) is a prevalent complexity in the patients hospitalized in the intensive care unit (ICU) [5]. This disease leads to increase of mortality in patients, duration of hospitalization and therapeutic costs [9, 10]. Rate of appearance of pneumonia caused by ventilator has been reported in a range from 7.5% to 45% in the various studies [11-15].
Cause of difference in outbreak of VAP in the various studies in due to different attribute of the hospitalized patients and accompaniment of susceptible - making factors and their various background diseases.
The most prevalent bacterial factors of pneumonia resulted from ventilator in the current study were genera of Enterobacteriaceae Family (35.4%), including strains of Klebsiella pneumonia, Enterococcus, Enterobacter, Escherichiacoli and Morganella. Then, they have been Staphylococcus aureus (20.7%), negative coagulase Staphylococci (14.7%), Pseudomonas aeruginosa (11.3%), Acinetobacter baumannii (9.4%) and Corynebacterium (7.5%).
In a study carried out in the Shariati hospital of Tehran [16], Frequency of the microorganisms causing VAP was 37.6% in Gram- negative bacterium, 31.7% in Gram- positive bacterium, 9.4% in Fungus and 21.3% without a special factor.
In case Frequency of the Gram- positive bacteria is 52.8% and that of the Gram- negative bacteria is 47.2% in this research which differs from the above research, this subject represents increase of invasive inclination of the Gram-positive bacteria towards establishment of pneumonia in these patients.
In the study conducted by Vincent et al., bacterial factors of VAP were reported to be 34.4% in Enterobacteriaceae, 30.1% in Staphylococcus aureus and 19.1% in the negative coagulase staphylococci. while in the research carried out by Jordi rello et al., Pseudomonas aeruginosa, Staphylococcusaureus, homphillusinfluenza, Acenitobacter, Klebsiellapneumonia and Enterobacteriaceae include 31.7%, 11.8%, 8.4%, 11.8%, 7.7% and 13.7% of the bacteria isolated from the patients, respectively [17, 18].
In the study of Diouf et al. [19], rate of outbreak of Pseudomonas aeruginosa and Acetinobacter has been reported to be 68% approximately, while, in the current study, outbreak of these two bacteria is 11.3% and 9.4%, respectively which doesn’t correspond with the findings of the present research.
In this study, rate of prevalence of species of the bacteria belonging to family of Enterobacteriaceae and Staphylococcus aureus among those with hospitalized pneumonia required mechanical ventilation was higher than the other micro organisms, suggesting more tendency of these microorganisms to establish pneumonia in the patients who are dependent on them mechanical ventilation.
As it was observed earlier, the greatest microorganisms isolated from several cultures for the patients with VAP are gram-negative bacteria, while, in the Shariati hospital of Tehran, the gram-negative bacteria have been causative factor of establishment of VAP in less than half of the cases. However, it that study, these microorganisms had acquired the first rank in terms of outbreak, suggesting invasive inclination and adaptability of these microorganisms in the ICU and easy transfer of these factors, through apparatuses and personnel, to the patients hospitalized in this ward.
5.1. Conclusion
VAP is a serious challenge in the patients who are in need of mechanical ventilation, and accurate enforcement of controlling the hospitalized infection to prevent from it in ICU is to be advised. Reduction of duration of intubation and number of microorganisms in mouth through repetition of oral suction and disinfection and keeping clean the suction after usage leads to decrease of the organisms reached the lung and causes reduction of the VAP.