1. Background
Congenital heart disease is the most common type of genetic disorder in children (1-3). The prevalence of congenital heart disease at birth has been reported to be between 6 and 13 subjects per 1000 live births (4-6), and more than half of these patients need surgery during the first year of life. These children are predisposed to more infections and more severe ones. In addition, some types of congenital heart disease are associated with genetic disorders that cause levels of immunodeficiency and increased susceptibility to infection (7); thus, these surgeries, in turn, increase the risk of nosocomial infections due to ulcers (8). Although post-surgical infections are well-known complications, different causative germs depend on the type and site of cardiac surgery. Post-cardiac surgery infections are responsible for the high rate of morbidity, prolonged ICU stay, and even mortality; therefore, having a comprehensive epidemiological knowledge of responsible germs makes antibiotic prophylaxis more accurate and effective in these kinds of surgeries. In this study, we tried to find a microbiologic pattern on germs responsible for post-cardiac surgery infections.
Several studies have examined postoperative infections in children undergoing heart surgery as the most common postoperative infections in surgical sites, with a prevalence of 1.2 to 48% (9-12). The difference in the prevalence rate may happen because of the difference in factors affecting the infectious events, such as body mass index (BMI), cardiopulmonary bypass time, the surgery duration, the age of less than one month, blood transfusion, intravenous feeding, central venous catheters indwelling time, severe bleeding 24 hours after surgery, and misuse scheduling of antibiotics before surgery (7, 9, 13, 14). Common microbial agents that are found at the site of surgery-related infections are Staphylococcus aureus (15), Coagulase-negative staphylococci (16), Escherichia coli (17), Enterococcus faecalis (18), and Pseudomonas spp. (19), respectively. In the last decade, the number of Gram-negative bacilli has decreased, and the proportion of infections associated with S. aureus has increased (20).
In spite of the importance of congenital heart disease and the higher risk of complications and mortality rate, few studies have evaluated the prevalence and factors affecting the disease and its treatment methods and complications (21, 22). Although few studies on surgical infections in children with congenital heart disease in developed and developing countries, these studies have shown the high prevalence of infection in these surgical procedures (7). However, these studies have some limitations, like a small number of samples and errors in retrospective data collection (21). Therefore, identifying the most common causes of infections can help prevent nosocomial infectious diseases. Accordingly, the costs and the length of hospitalization can be decreased, and more importantly, the indiscriminate use of broad-spectrum antibiotics and, consequently, antibiotic resistance can be reduced. Furthermore, monitoring the disinfection of surfaces and equipment and checking the principles of infection control cause a significant reduction in the frequency of such infections.
2. Objectives
The higher susceptibility to severe infection causes the increased risk of nosocomial infection and consequent surgery complications (7). Therefore, additional studies are needed to identify pathogens and the risk factors affecting the disease to improve preventive procedures. In this regard, the present study was conducted to evaluate the type of microbial agents inducing post-cardiac surgery infection and also the possible differences between these infections in terms of age, sex, disease types, and hospitalization length.
3. Methods
3.1. Study Population
In this cross-sectional study, the records of all patients who underwent cardiac surgery for congenital and structural heart diseases during four years (2014 - 2017) in Imam Reza training hospital affiliated with the Mashhad University of Medical Sciences were reviewed. Demographic information included age (between less than one month and 36 months), sex, and weight. Medical data, including hospitalization time, underlying congenital heart disease, history of nosocomial infection, location of nosocomial infection, and the type of microbial infection in the laboratory samples (sputum, blood, urine, and surgical wound) were collected using medical documents.
The nosocomial infections included ventilator-related infections, pneumonia, sepsis, urinary tract infections, infections at the surgical wound site, and cardiovascular, skin, and soft tissue infections.
All the patients with congenital, structural, and valvular heart diseases who underwent palliative or total reconstructive surgical correction in eighter of the pediatric cardiac surgery intensive care units in our center were included in this study.
The study was approved by the Mashhad University of Medical Sciences Ethics Committee with the ethical approval number: IR.MUMS.MEDICAL.REC.1397.356. All study procedures followed Helsinki ethics, and a coding system was used to keep the patients' information confidential.
3.2. Statistical Analysis
Data were analyzed using SPSS V16. A P-value less than 0.05 was considered significant. Descriptive indices of mean, standard deviation, frequency, and percentage were used to describe the variables. The chi-square test and t-test (or its non-parametric equivalent) were used to compare qualitative and quantitative data.
4. Results
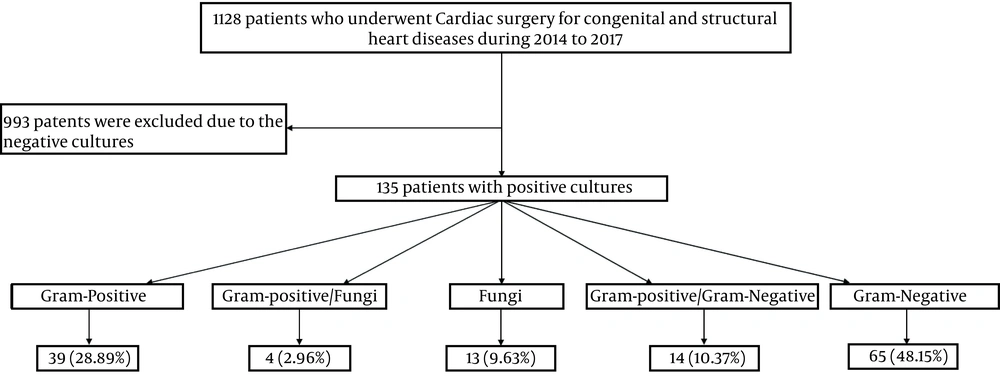
A total of 135 patients were enrolled in the study (Figure 1), of whom 80 patients (60.1%) were male, and 55 patients (39.9%) were female. The mean age of patients was 8.06 ± 3.86 months. Most patients (90/135, 66.7%) were under one month old. Most patients had ventricular septal defect (VSD) (45/135, 32.9%), atrial septal defect (ASD) (32/135, 21.9%), and atrioventricular septal defect (AVSD) (22/135, 10.5%). Ventilator-related infections were among the most reported nosocomial infections (36%), followed by catheter-related urinary tract infections (29%), infections at the site of the surgical wound (16%), sepsis (8%), and skin and soft tissue infections (11%).
Regarding the isolated microorganisms’ distribution, the most common Gram-negative types were Acinetobacter (14/135, 14.1%), followed by Pseudomonas spp. (13/135, 9.6%) and Enterobacter (13/135, 9.6%). Also, Corynebacterium diphtheria (10/135, 7.4%), Staphylococcus epidermidis (10/135, 7.4%), and S. aureus (10/135, 7.4%) were the most frequent Gram-positive types. Moreover, Candida albicans (14/135, 10.4%) was identified as the most common fungi. The type and frequency of all identified bacterial strains are reported in Table 1.
| Characteristic | No. (%) |
|---|---|
| Age | |
| < 1 month | 90 (66.7) |
| 2 - 5 | 26 (19.3) |
| 6 - 10 | 7 (5.2) |
| 11 - 15 | 3 (2.2) |
| 16 - 20 | 0 (0) |
| 21 - 25 | 3 (2.2) |
| 26 - 30 | 4 (3.0) |
| ≥ 31 | 2 (1.5) |
| Sex | |
| Male | 80 (60.1) |
| Female | 55 (39.9) |
| Hospitalization unit | |
| ICUA | 3 (1.9) |
| ICUB | 62 (48.6) |
| ICUC | 2 (1.0) |
| ICUD | 2 (1.0) |
| NICU | 9 (6.7) |
| open-heart surgery ICU | 44 (31.4) |
| Children ward | 2 (1.0) |
| open-heart surgery ward | 11 (8.6) |
| Final diagnosis | |
| VSD | 45 (32.9) |
| ASD | 32 (21.9) |
| AVSD | 22 (10.5) |
| PDA | 8 (7.7) |
| TOF | 6 (5.6) |
| Cardiomyopathy | 6 (5.6) |
| PS | 4 (4.0) |
| PH | 4 (4.0) |
| TGA | 1 (1.0) |
| CAD | 1 (1.0) |
| COA | 1 (1.0) |
| TAPVC | 1 (1.0) |
| Lupus+ cardiac tamponade | 1 (1.0) |
| Heart failure | 3 (2.8) |
| Bacteria type | |
| Acinetobacter | 19 (14.1) |
| Candida albicans | 14 (10.4) |
| Pseudomonas spp. | 13 (9.6) |
| Enterobacter | 13 (9.6) |
| Corynebacterium diphtheriae | 10 (7.4) |
| Staphylococcus epidermidis | 10 (7.4) |
| Methicillin-resistant Staphylococcus epidermidis | 8 (5.9) |
| Klebsiella | 7 (5.2) |
| Hemolytic Staphylococcus | 7 (5.2) |
| Enterococcus | 6 (4.4) |
| Escherichia coli | 4 (3.0) |
| Multidrug resistant Acinetobacter | 3 (2.2) |
| Gram-negative bacilli | 2 (1.5) |
| Stenotrophomonas maltophilia | 2 (1.5) |
| Vancomycin-resistant Enterococcus | 1 (0.7) |
| Methicillin-resistant Staphylococcus aureus | 1 (0.7) |
| Micrococcus | 1 (0.7) |
| Coagulase-negative staphylococci | 1 (0.7) |
| Gram-positive bacilli | 1 (0.7) |
Abbreviations: VSD, ventricular septal defect; ASD, atrial septal defect; AVSD, atrioventricular septal defect.
Comparing group differences based on age revealed that the most positive cultured subjects were under ten months of age, specifically one-month-old. Among 19 positive cultures of Acinetobacter, 15 cases were in subjects under one month of age, and four cases were in the second group (1 to 5 months). Moreover, the number of C. albicans was higher (10 out of 14) in the first group (one-month-old) than in others. However, there was no significant difference in infectious bacteria between different age groups (P = 0.571). The frequency of isolated infectious bacteria in age groups is reported in Table 2.
| Bacteria Type | Age; No. (%) | |||||||
|---|---|---|---|---|---|---|---|---|
| < 1 Month | 2 - 5 | 6 - 10 | 11 - 15 | 21 - 25 | 26 - 30 | ≥ 31 | Total | |
| Corynebacterium diphtheriae | 7 (7.8) | 1 (3.8) | 2 (28.6) | 0 | 0 | 0 | 0 | 10 (7.4) |
| Staphylococcus epidermidis | 7 (7.8) | 1 (3.8) | 2 (28.6) | 0 | 0 | 0 | 0 | 10 (7.4) |
| Methicillin-resistant Staphylococcus epidermidis | 5 (5.6) | 1 (3.8) | 1 (14.3) | 0 | 0 | 0 | 1 (50.0) | 8 (5.9) |
| Klebsiella | 4 (4.4) | 2 (7.7) | 0 | 0 | 0 | 1 (25.0) | 0 | 7 (5.2) |
| Acinetobacter | 15 (16.7) | 4 (15.4) | 0 | 0 | 0 | 0 | 0 | 19 (14.1) |
| Enterococcus | 5 (5.6) | 0 | 0 | 0 | 0 | 0 | 1 (50.0) | 6 (4.4) |
| Vancomycin-resistant Enterococcus | 1 (1.1) | 0 | 0 | 0 | 0 | 0 | 0 | 1 (0.7) |
| Pseudomonas spp. | 6 (6.7) | 5 (19.2) | 1 (14.3) | 0 | 0 | 1 (25.0) | 0 | 13 (9.6) |
| Enterobacter | 9 (10.0) | 2 (7.7) | 0 | 2 (66.7) | 0 | 0 | 0 | 13 (9.6) |
| Hemolytic Staphylococcus | 4 (4.4) | 1 (3.8) | 0 | 1 (33.3) | 1 (33.3) | 0 | 0 | 7 (5.2) |
| Candida albicans | 10 (11.1) | 2 (7.7) | 1 (14.3) | 0 | 0 | 0 | 1 (50.0) | 14 (10.4) |
| Escherichia coli | 4 (4.4) | 0 | 0 | 0 | 0 | 0 | 0 | 4 (3.0) |
| Methicillin-resistant Staphylococcus aureus | 1 (1.1) | 0 | 0 | 0 | 0 | 0 | 0 | 1 (0.7) |
| Gram-negative bacilli | 1 (1.1) | 1 (3.8) | 0 | 0 | 0 | 0 | 0 | 2 (1.5) |
| Multidrug resistant Acinetobacter | 1 (1.1) | 0 | 0 | 0 | 1 (33.3) | 0 | 0 | 3 (2.2) |
| Stenotrophomonas maltophilia | 2 (2.2) | 0 | 0 | 0 | 0 | 0 | 0 | 2 (1.5) |
| Micrococcus | 1 (1.1) | 0 | 0 | 0 | 0 | 0 | 0 | 1 (0.7) |
| Coagulase- negative staphylococci | 0 | 1 (3.8) | 0 | 0 | 0 | 0 | 0 | 1 (0.7) |
| Gram-positive bacilli | 0 | 0 | 1 (14.3) | 0 | 0 | 0 | 0 | 1 (0.7) |
Analysis of the distribution of bacterial type between sex groups indicated that the prevalence of Klebsiella, vancomycin-resistant enterococci, E. coli, methicillin-resistant S. aureus, methicillin-resistant S. aureus, Coagulase-negative staphylococci, and Gram-positive bacilli was higher in males. Also, Enterococcus, C. albicans, Gram-negative bacilli, and Strophomonas maltophilia had equal prevalence in both groups. Although the frequency of other bacteria was higher in females, there was no significant difference between different sex groups regarding the bacteria causing the infection (P = 0.621) (Table 3).
| Bacteria Type | Sex; No. (%) | |
|---|---|---|
| Male | Female | |
| Corynebacterium diphtheriae | 4 (7.3) | 6 (7.5) |
| Staphylococcus epidermidis | 4 (7.3) | 6 (7.5) |
| Methicillin-resistant Staphylococcus epidermidis | 1 (1.8) | 7 (8.8) |
| Klebsiella | 4 (7.3) | 3 (3.8) |
| Acinetobacter | 3 (5.5) | 16 (20.0) |
| Enterococcus | 3 (5.5) | 3 (3.8) |
| Vancomycin-resistant Enterococcus | 1 (1.8) | 0 |
| Pseudomonas spp. | 6 (10.9) | 7 (8.8) |
| Enterobacter | 5 (9.1) | 8 (10.0) |
| Hemolytic Staphylococcus | 1 (1.8) | 6 (7.5) |
| Candida albicans | 7 (12.7) | 7 (8.8) |
| Escherichia coli | 3 (5.5) | 1 (1.3) |
| Methicillin-resistant Staphylococcus aureus | 1 (1.8) | 0 |
| Gram-negative bacilli | 1 (1.8) | 1 (1.3) |
| Multidrug resistant Acinetobacter | 1 (1.8) | 2 (2.5) |
| Stenotrophomonas maltophilia | 1 (1.8) | 1 (1.3) |
| Micrococcus | 0 | 1 (1.3) |
| Coagulase-negative staphylococci | 1 (1.8) | 0 |
| Gram-positive bacilli | 1 (1.8) | 0 |
Regarding the isolated bacteria in different diseases, Candida was the most frequently isolated bacteria in ASD subjects. Moreover, Acinetobacter was the most abundant bacteria in VSD and AVSD patients. However, there was no significant difference in terms of the bacteria causing infection between different congenital diseases (P = 0.831) (Table 4).
| Bacteria type | Disease Type; No. (%) | |||||||||||||
|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|
| ASD | AVSD | CAD | COA | COM | HF | L+CT | PDA | PH | PS | TAPVC | TGA | TOF | VSD | |
| Corynebacterium diphtheriae | 2 (6.3) | 1 (4.5) | 0 | 0 | 1 (16.7) | 0 | 0 | 0 | 1 (25.0) | 0 | 0 | 0 | 1 (16.7) | 3 (6.7) |
| Staphylococcus epidermidis | 3 (9.4) | 1 (4.5) | 0 | 0 | 1 (16.7) | 0 | 0 | 0 | 2 (50.0) | 0 | 0 | 0 | 0 | 0 |
| Staphylococcus epidermidis+ Corynebacterium diphtheriae | 0 | 0 | 0 | 0 | 0 | 0 | 0 | 0 | 0 | 0 | 0 | 0 | 0 | 1 (2.2) |
| Staphylococcus epidermidis+ Candida albicans | 1 (3.1) | 0 | 0 | 0 | 0 | 0 | 0 | 0 | 0 | 0 | 0 | 0 | 0 | 0 |
| Methicillin-resistant Staphylococcus epidermidis | 1 (3.1) | 1 (4.5) | 0 | 0 | 1 (16.7) | 0 | 0 | 0 | 0 | 0 | 0 | 0 | 0 | 3 (6.7) |
| Methicillin-resistant Staphylococcus epidermidis + Enterobacter | 0 | 0 | 0 | 0 | 0 | 0 | 0 | 0 | 0 | 0 | 0 | 0 | 0 | 1 (2.2) |
| Methicillin-resistant Staphylococcus epidermidis+ Pseudomonas spp. | 0 | 1 (4.5) | 0 | 0 | 0 | 0 | 0 | 0 | 0 | 0 | 0 | 0 | 0 | 0 |
| Klebsiella | 2 (6.3) | 0 | 0 | 0 | 1 (16.7) | 0 | 0 | 0 | 0 | 1 (25.0) | 0 | 0 | 0 | 2 (4.4) |
| Klebsiella+ Enterobacter | 0 | 0 | 0 | 0 | 0 | 0 | 0 | 0 | 0 | 1 (25.0) | 0 | 0 | 0 | 0 |
| Klebsiella+ Pseudomonas spp. | 0 | 1 (4.5) | 0 | 0 | 0 | 0 | 0 | 0 | 0 | 0 | 0 | 0 | 0 | 0 |
| Acinetobacter | 3 (9.4) | 5 (22.7) | 0 | 0 | 0 | 0 | 0 | 0 | 1 (25.0) | 2 (50.0) | 0 | 0 | 1 (16.7) | 6 (13.3) |
| Acinetobacter+ Pseudomonas spp.+ Enterobacter | 0 | 0 | 0 | 0 | 0 | 0 | 0 | 1 (12.5) | 0 | 0 | 0 | 0 | 0 | 0 |
| Enterococcus | 0 | 2 (9.1) | 0 | 1 (100.0) | 0 | 1 (33.3) | 0 | 0 | 0 | 0 | 0 | 0 | 0 | 1 (2.2) |
| Enterococcus + Candida albicans | 0 | 0 | 0 | 0 | 0 | 0 | 0 | 0 | 0 | 0 | 0 | 0 | 0 | 1 (2.2) |
| Vancomycin-resistant Enterococcus | 0 | 0 | 0 | 0 | 0 | 0 | 0 | 0 | 0 | 0 | 0 | 0 | 1 (16.7) | 0 |
| Pseudomonas spp. | 2 (6.3) | 3 (13.6) | 0 | 0 | 0 | 0 | 0 | 3 (37.5) | 0 | 0 | 0 | 0 | 0 | 3 (6.7) |
| Enterobacter | 2 (6.3) | 1 (4.5) | 1 (100.0) | 0 | 0 | 0 | 0 | 1 (12.5) | 0 | 0 | 0 | 0 | 0 | 4 (8.9) |
| Hemolytic Staphylococcus | 0 | 2 (9.1) | 0 | 0 | 0 | 1 (33.3) | 0 | 1 (12.5) | 0 | 0 | 0 | 0 | 0 | 2 (4.4) |
| Hemolytic Staphylococcus + Enterobacter | 0 | 1 (4.5) | 0 | 0 | 0 | 0 | 0 | 0 | 0 | 0 | 0 | 0 | 0 | 0 |
| Candida albicans | 8 (25.0) | 0 | 0 | 0 | 1 (16.7) | 0 | 0 | 0 | 0 | 0 | 0 | 0 | 0 | 3 (6.7) |
| Escherichia coli | 0 | 1 (4.5) | 0 | 0 | 0 | 0 | 0 | 0 | 0 | 0 | 0 | 0 | 0 | 3 (6.7) |
| Methicillin-resistant Staphylococcus aureus | 0 | 0 | 0 | 0 | 0 | 0 | 0 | 1 (12.5) | 0 | 0 | 0 | 0 | 0 | 0 |
| Gram-negative bacilli | 0 | 0 | 0 | 0 | 0 | 0 | 0 | 1 (12.5) | 0 | 0 | 0 | 0 | 0 | 0 |
| Gram-negative bacilli+ Staphylococcus epidermidis | 1 (3.1) | 0 | 0 | 0 | 0 | 0 | 0 | 0 | 0 | 0 | 0 | 0 | 0 | 0 |
| Multidrug resistant Acinetobacter | 2 (6.3) | 1 (4.5) | 0 | 0 | 0 | 0 | 0 | 0 | 0 | 0 | 0 | 0 | 0 | 0 |
| Stenotrophomonas maltophilia | 1 (3.1) | 0 | 0 | 0 | 1 (16.7) | 0 | 0 | 0 | 0 | 0 | 0 | 0 | 0 | 0 |
| Micrococcus | 0 | 0 | 0 | 0 | 0 | 0 | 0 | 0 | 0 | 0 | 0 | 0 | 1 (16.7) | 0 |
| Coagulase-negative staphylococci | 0 | 0 | 0 | 0 | 0 | 0 | 0 | 0 | 0 | 0 | 0 | 0 | 1 (16.7) | 0 |
| Gram-positive bacilli | 0 | 0 | 0 | 0 | 0 | 0 | 0 | 0 | 0 | 0 | 0 | 0 | 0 | 1 (2.2) |
| Negative culture | 4 (12.5) | 1 (4.5) | 0 | 0 | 0 | 1 (33.3) | 1 (100.0) | 0 | 0 | 1 (25.0) | 1 (100.0) | 1 (100.0) | 1 (16.7) | 11 (24.4) |
Abbreviations: VSD, ventricular septal defect; ASD, atrial septal defect; AVSD, atrioventricular septal defect.
Weight-based categorization indicated that 13 subjects with weight less than 5 kg, 45 patients with a weight of 5 to 10 kg, and 77 cases with more than 10 kg had positive cultures. The most abundant bacterium in the less than 5 kg group was S. epidermidis. The three most common bacteria in the 5 to 10 kg group were Acinetobacter, S. aureus (alone and with Enterobacter), and Enterococcus, respectively. The most common bacteria in the third group (> 10 kg) were Acinetobacter, Candida, Pseudomonas, and Corynebacterium diphtheria. There was no significant difference in the type of bacteria between the three weight groups (P = 0.786) (Table 5).
| Bacteria Type | Weight (kg); No. (%) | ||
|---|---|---|---|
| < 5 | 5 - 10 | > 10 | |
| Corynebacterium diphtheriae | 1 (7.7) | 1 (2.2) | 8 (10.4) |
| Staphylococcus epidermidis | 2 (15.4) | 1 (2.2) | 5 (6.5) |
| Staphylococcus epidermidis+ Corynebacterium diphtheriae | 0 | 0 | 1 (1.3) |
| Staphylococcus epidermidis+ Candida albicans | 0 | 0 | 1 (1.3) |
| Methicillin-resistant Staphylococcus epidermidis | 1 (7.7) | 0 | 5 (6.5) |
| Methicillin-resistant Staphylococcus epidermidis + Enterobacter | 0 | 1 (2.2) | 1 (2.2) |
| Methicillin-resistant Staphylococcus epidermidis+ Pseudomonas spp. | 0 | 1 (2.2) | 1 (1.3) |
| Klebsiella | 1 (7.7) | 0 | 5 (6.5) |
| Klebsiella+ Enterobacter | 0 | 0 | 1 (1.3) |
| Klebsiella+ Pseudomonas spp. | 0 | 0 | 1 (1.3) |
| Acinetobacter | 0 | 11 (24.4) | 10 (12.9) |
| Acinetobacter+ Pseudomonas spp. + Enterobacter | 1 (7.7) | 1 (2.2) | 1 (1.3) |
| Enterococcus | 1 (7.7) | 6 (13.3) | 0 |
| Enterococcus + Candida albicans | 1 (7.7) | 1 (2.2) | 1 (1.3) |
| Vancomycin-resistant Enterococcus | 0 | 1 (2.2) | 0 |
| Pseudomonas spp. | 1 (7.7) | 3 (6.6) | 8 (10.4) |
| Enterobacter | 1 (7.7) | 5 (11.1) | 3 (3.9) |
| Hemolytic Staphylococcus | 0 | 0 | 3 (3.9) |
| Hemolytic Staphylococcus + Enterobacter | 1 (7.7) | 5 (11.1) | 1 (1.3) |
| Candida albicans | 0 | 3 (6.7) | 10 (12.9) |
| Escherichia coli | 1 (7.7) | 2 (4.4) | 1 (1.3) |
| Methicillin-resistant Staphylococcus aureus | 0 | 0 | 1 (1.3) |
| Gram-negative bacilli | 0 | 0 | 1 (1.3) |
| Gram-negative bacilli+ Staphylococcus epidermidis | 1 (7.7) | 1 (2.2) | 1 (1.3) |
| Multidrug resistant Acinetobacter | 0 | 1 (2.2) | 4 (5.2) |
| Stenotrophomonas maltophilia | 0 | 1 (2.2) | 1 (1.3) |
| Micrococcus | 0 | 1 (2.2) | 0 |
| Coagulase-negative staphylococci | 0 | 0 | 1 (1.3) |
| Gram-positive bacilli | 0 | 0 | 1 (1.3) |
| Total | 13 (100) | 45 (100) | 77 (100) |
The duration of hospitalization (P = 0.002), intubation time (P = 0.004), bypass time (P = 0.006), and duration of urinary catheterization (P > 0.0001) were significantly longer in positive cultures than in negative ones (Table 6).
| Characteristics | Culture | P-Value | |
|---|---|---|---|
| Positive | Negative | ||
| Hospitalization time (day) | 6.34 ± 3.21 | 4.05 ± 2.44 | 0.002 |
| Intubation time (h) | 14.28 ± 5.14 | 10.43 ± 4.31 | 0.004 |
| Bypass time (min) | 130.3 ± 10.4 | 91.5 ± 2.8 | 0.006 |
| Urinary catheterization time (day) | 13.16 ± 1.8 | 3.1 ± 0.3 | < 0.0001 |
5. Discussion
This study was designed to identify microorganisms that caused post-cardiac surgery infection in these patients. In this regard, 135 patients who underwent heart surgery were enrolled the study. Most of the subjects were infants, and the most common abnormalities were VSD, followed by ASD and PDA. Acinetobacter, C. albicans, Pseudomonas spp., Enterobacter, C. diphtheria, and S. epidermidis were identified as the most common infectious bacteria. Our findings showed no significant difference between age, sex, weight, and disease types, and bacterial species. However, the duration of hospitalization, intubation, bypass time, and urinary catheterization was significantly different between positive and negative cultures.
The prevalence of post-cardiac surgery infection in children was reported at 11.96% in our study. Several examinations have studied the distribution of nosocomial infection in children. However, a vast variation was observed in prevalence reported by different studies ranging from 8.7% to 17.7% (23-26). In a new survey on a total of 11,651 subjects (0 - 10 years old), the rate of nosocomial infection was reported to be 10.8% (9). However, another study reported an extremely high frequency of the infection, almost 36% in 155 cases under consideration (13).
On the other hand, a study in the UK on 3090 consecutive pediatric cardiac surgical admissions revealed the post-surgical infection in 0.9 to 2.9% of subjects, independently and along with other morbidities, respectively, that was a lower frequency than previous studies (27). These variations can be explained by individual characteristics in considered populations, like genetic background and probable higher susceptibility to infectious diseases. Also, environmental factors, like infection control protocols and antiseptic materials used in hospitals, may be different. Therefore, evaluation of health issues, attention to disinfection of equipment, and the infection control disciplines lead to a significant reduction in the prevalence of the infection.
In the present study, the most common infectious bacteria in infants were Acinetobacter, Candida albicans, Enterobacter, Pseudomonas spp., C. diphtheria, and S. epidermidis. Similarly, a previous study reported S. epidermidis, S. aureus, and Klebsiella as blood-borne infectious pathogens, and methicillin-resistant staphylococci, methicillin-susceptible staphylococci, C. diphtheria, E. faecalis, and Escherichia coli as infectious microorganisms at the surgical site (24). Other studies have also indicated the higher prevalence of S. epidermidis, Klebsiella, Pseudomonas, Acinetobacter, S. pneumonia, Beta hemolytic streptococcus, Candida, and H. influenza as main infectious post-cardiac surgery agents in children (23, 25, 28, 29). Although the identified infectious organisms were similar between studies, the rate and rank of pathogens were different.
Based on our results, bacterial type distribution was similar between different age, sex, and heart disease groups. Although the infection rate in patients under one month old was higher than in others, no difference was observed between age groups. Conversely, previous studies have reported age as an independent risk factor for surgical site infections after pediatric cardiovascular surgery (30, 31). Also, another study indicated that the younger age is related to more susceptibility to attract infections (32). However, sex was not a risk factor for infection (9). Furthermore, we found that prolonged hospitalization, intubation, bypass, and urinary catheterization time was more significant in patients with positive cultures. Consistent with these findings, it has previously been reported that a longer duration of surgery, prolonged ICU stay, catheter indwelling time, and longer previous hospital stay were associated with nosocomial post-cardiac surgery infections (13, 30, 31).
As one of the limitations, this study was limited to one center, which reduced the sample size. Furthermore, the prevalence of post-cardiac surgery infection has not been evaluated compared with other types of surgeries based on the age distribution. However, our study is one of the few studies in this area, especially to examine the age distribution and distribution of infectious masses among different types of congenital heart disease. The results of this study can be used in two parts. First, identifying the bacteria causing the infection in children who have had heart surgery can improve our knowledge about antibiotics that could be used for empirical treatment or as prophylaxis. Also, the distribution of infectious bacteria at different ages can help to prescribe these antibiotics more accurately. Second, this study can add more evidence to the literature about the distribution of pathogens in different age categories and various heart diseases, as previous researches have paid less attention to these features.
5.1. Conclusion
The prevalence of nosocomial infection in open-heart surgery in our patients was 11.96%. The duration of hospitalization, intubation, bypass, and urinary catheterization was significantly higher in positive cultures than negative ones. Therefore, decreasing the time of these situations may reduce nosocomial infection. Consequently, treatment costs, as well as resistance to antibiotics, can be reduced. However, more studies should be conducted with a larger sample size focusing on age and microbial type distribution among different heart diseases to confirm these findings.