1. Background
Obesity is a widespread health condition that has increased in recent decades. Body weight gain is associated with the etiology of a wide range of human cancers (1). Numerous studies have shown an association between obesity and cancer progression in various tissues, especially ovarian (2), pancreatic (3), esophageal, colon, prostate, and breast tissues (4).
Adipose tissues secrete various hormones, cytokines, and metabolites called Adipokines. As the body gains weight, some changes occur in the number and size of cells, adipokines secretion, adipocyte death, local hypoxia, and fatty acid motility (5). Adipose tissue dysfunction is known as the main cause of obesity and related disorders (6). Adipokines are produced and secreted by various adipose tissues, including subcutaneous, visceral, and breast adipose tissues. In the breast tissue, Adipokines are in paracrine and autocrine pathways (7). Adipose tissue itself (8) following the ovaries (9) is a source of estradiol production through the aromatization of androgens (8). A direct relationship has been reported between body weight and aromatase activity (8). With increased physical activity, the serum level of estradiol decreases; it indicated the effect of body weight on estradiol concentration (9).
Leptin is the most important adipokine that regulates energy balance and nutrient uptake in the hypothalamus. At the cellular level, it acts as a mitogen, a metabolic regulator, and an angiogenic agent. The role of this hormone in tumorigenesis, especially in the development of breast, colorectal, and prostate cancers, has been discussed (10). Leptin can directly affect obesity-caused breast cancer by the continuous proliferation of benign and malignant cells in the breast epithelium and reduced cell proliferation modulators (11).
Adiponectin, a peptide hormone released from fat tissue, is inversely related to body fat (12). This multifunctional protein, which affects various tissues and organs in the body, has been implicated in the development of obesity-related disorders, such as metabolic syndrome, diabetes, cardiovascular disease, and malignancies. Most studies have reported an association between low concentrations of this hormone and an increased risk of breast cancer (4). Generally, the level of this hormone is independent of adipose mass, and differences may be attributed to estradiol or bloodstream androgens (13).
2. Objectives
In this study, the effects of body mass index (BMI) on serum concentrations of estradiol, leptin, and adiponectin were evaluated. Also, the association between the risk of breast cancer and polymorphisms in leptin gene promoter (LEPG2548A rs7799039), polymorphisms in leptin receptors (Q223R rs1137101, K109 rs1137100, and K656N rs8179183), and adiponectin polymorphisms (T45G Rs2241766, G276T rs1501299, C11377G rs266729, and G11391A rs17300539) were studied concerning BMI in two groups of women.
3. Methods
3.1. Study Participants (Includes/Excludes Criteria)
The protocol of the study is following the ethical guidelines of the 1975 Helsinki Declaration, which was reflected in the prior approval of the University Medical Research Committee. After approval of the study protocol, blood samples were collected from 350 women, who had been referred to hospitals in Tehran. Totally, 175 women with breast cancer, whose disease was diagnosed by mammography, ultrasonography, and biochemical tests, needed surgical interventions and were selected by a specialist. Limitations of the study included not taking medications and not starting treatments such as chemotherapy, radiotherapy, etc. The healthy controls (175 women) also did not have specific diseases such as cardiovascular or diabetes. The case and control groups were matched for age, area of residency, and absence of cardiovascular disease or diabetes. In both groups, postmenopausal and non-menopausal women were selected (menopause if not for using estrogen therapy). They were in the same terms of not smoking, not alcohol consumption, and not having thyroid or fatty liver diseases, which resulted in the exclusion of large numbers of individuals.
According to the limitations of the study, individuals who had taken medications for migraine, thyroid problems, fatty liver, or any other drug were excluded from the study. After obtaining informed consent from all participants, some information, such as age, height, weight, and family history of breast cancer were collected. Pathological information was collected from their medical records after surgery.
3.2. DNA and Serum Isolation
For DNA extraction and serum preparation, blood samples (5 - 7 mL) were collected between 8 a.m. and 10 a.m. after 8 to 10 hours of fasting. Some of the blood samples were poured into a tube containing 200 μL of 0.5 mM EDTA and stored at -20°C to extract DNA based on the salting-out method. The remaining blood sample was transferred to a separate tube via centrifugation for serum separation and stored at -80°C. Hormone levels were measured after serum separation, using commercial 17-beta estradiol (DiaMetra Company, Italy), leptin (LDN, Nordhorn), and adiponectin (AviBion, Finland) kits. The assays were performed according to the protocol of each kit.
3.3. Selection of Single-Nucleotide Polymorphisms and Genotyping
In this study, single-nucleotide polymorphisms (SNPs), which were associated with the risk of multiple malignancies and diseases according to many previous studies, were selected (14-20). To investigate genetic variations in leptin, G2548A polymorphism at the gene promoter site, 3 polymorphisms, including Q223R, K109R, and K656N at the leptin receptor site were also selected. Also, genetic variations of adiponectin were investigated in G276T, T45G, C11377G, and G11391A polymorphisms.
After testing the quality and quantity of the extracted DNA, the samples were amplified with the designed primers (Table 1). After enzymatic digestion based on the size of formed bands on 2% gel, genotypes were determined in the samples. To confirm the results, some samples were randomly re-tested, and positive and negative controls were used to confirm enzymatic digestion.
| Polymorphism and Sequence 5’ → 3’ | TM | PCR Product | Restriction Enzyme | Digested Fragment |
|---|---|---|---|---|
| Leptin, G2548A, rs7799039 | 58.8°C | 229 bp | HhaI | GG: 181 bp, 48 bp; AA: 229 bp; GA: 229, 181 and 48 bp |
| F: TCCCGTGAGAACTATTCTTCTTTTG | ||||
| R: AAAGCAAAGACAGGCATAAAA | ||||
| Leptin R, Q223R, rs1137101 | 59.9°C | 366 bp | Mae III | AA: 243 bp and 123 bp; GG: 366 bp; AG: 366, 243 and 123 bp |
| F: TCCTGCTTTAAAAGCCTATCCAGTATTT | ||||
| R: AGCTAGCAAATATTTTTGTAAGCAAT | ||||
| Leptin R, K109R, rs1137100 | 59°C | 104 bp | BsgI | AA: 72 bp and 32 bp; GG: 104 bp; AG: 104, 72 and 32 bp |
| F: TTTTTTCCACTGTTGCTTTCGGA | ||||
| R: AAACTAAAGAATTTACTGTTGAAACAAATGTC | ||||
| Leptin R, K656N, rs8179183 | 61.1°C | 258 bp | BstUI | GG: 231 bp and 27 bp; CC: 258 bp; GC: 258, 231 and 27 bp |
| F: GCTAGATGGACTGGGATATTGGAGTAAT | ||||
| R: CTTCCAAAGTAAAGTGACATTTTTCTC | ||||
| Adiponectin, T45G, rs2241766 | 60.5°C | 372 bp | BsphI | TT: 216 bp and 156 bp; GG: 372 bp; TG: 372, 216 and 156 bp |
| F: GAAGTAGACTCTGCTGAGATGG | ||||
| R: TATCAGTGTAGGAGGTCTGTGATG | ||||
| Adiponectin, G276T, rs1501299 | 65.4°C | 468 bp | Mva1269I | GG: 320 bp and 148 bp; TT: 468 bp; GT: 468, 320 and 148 bp |
| F: TCTCTCCATGGCTGACAGTG | ||||
| R: AGATGCAGCAAAGCCAAAGT | ||||
| Adiponectin, C11377G, rs266729 | 58.8°C | 163 bp | BspCNI | CC: 120 bp and 43 bp; GG: 163 bp; CG: 163, 120 and 43 bp |
| F: CATCAGAATGTGTGGCTTGC | ||||
| R: AGAAGCAGCCTGGAGAACTG | ||||
| Adiponectin, G11391A, rs17300539 | 58.8°C | 163 bp | MspI | GG: 137 bp and 26 bp; AA: 163 bp; GA: 163, 137 and 26 bp |
| F: CATCAGAATGTGTGGCTTGC | ||||
| R: AGAAGCAGCCTGGAGAACTG |
Primer Sequences, TM Primers, PCR Products, Restriction Enzyme, and Digested Fragments
3.4. Statistical Analysis
The distribution of genotypes and alleles was determined for each polymorphism in the case and control groups based on the Hardy-Weinberg equilibrium, using unconditional univariate and multivariate logistic regression analyses for measuring the crude and adjusted odds ratios (ORs) of polymorphisms. The participants in the two groups were divided into subgroups based on BMI (BMI ≥ 25 and BMI < 25) to determine the relationship between SNPs and BMI, using logistic regression. In addition, the participants were subdivided into two age subgroups (< 50 and ≥ 50 years) to study the relationship between polymorphism genotypes and age via logistic regression analysis. The mean serum levels of estradiol, leptin, and adiponectin in the case and control groups, BMI subgroups, and different genotypes of polymorphisms were calculated, using an unpaired t test. In all analyses, serum concentrations of hormones were reported as mean ± SD and the significance level was considered ≤ 0.05.
4. Results
The participants were in the age range of 35 to 72 years. Demographic and biochemical characteristics of patients with breast cancer and healthy controls are summarized in Table 2.
| Characteristics | Patients n = 175 | Controls n = 175 | P Value |
|---|---|---|---|
| Age (Range of years) | 0.470 | ||
| < 50 | 58 (33.14) | 56 (32) | |
| ≥ 50 | 117 (66.86) | 119 (68) | |
| Mean ± SD | 56.06 ± 8.09 | 55.39 ± 9.32 | |
| Body mass index (kg/m2) | 5.9 × 10 -5 | ||
| ≤ 25 | 74 (42.29) | 130 (74.29) | |
| > 25 | 101 (57.71) | 45 (25.71) | |
| Mean ± SD | 24.89 ± 2.06 | 24.02 ± 1.95 | |
| Menopausal statue | 0.052 | ||
| Premenopausal | |||
| Body mass index ≤ 25 | 34 (19.43) | 59 (33.71) | |
| Body mass index > 25 | 45 (25.71) | 21 (12) | |
| Mean ± SD | 24.67 ± 1.99 | 24.04 ± 2.06 | |
| Postmenopausal | 2.6 × 10-4 | ||
| Body mass index ≤ 25 | 40 (22.86) | 71 (40.57) | |
| Body mass index > 25 | 56 (32) | 24 (13.72) | |
| Mean ± SD | 25.08 ± 2.11 | 24.00 ± 1.87 | |
| Estrogen (ng/mL) | |||
| Body mass index ≤ 25 | 44.50 ± 13.98 | 37.45 ± 14.76 | 0.001 |
| Body mass index > 25 | 54.33 ± 15.97 | 37.61 ± 16.45 | 2.32 × 10-8 |
| Total Mean ± SD | 50.35 ± 15.92 | 37.49 ± 15.17 | 1.14 × 10-13 |
| Leptin (ng/mL) | |||
| Body mass index ≤ 25 | 17.64 ± 8.31 | 13.62 ± 4.31 | 1 × 10-5 |
| Body mass index > 25 | 17.69 ± 6.59 | 14.17 ± 5.11 | 0.002 |
| Total Mean ± SD | 17.67 ± 7.34 | 13.76 ± 4.53 | 5.3 × 10-9 |
| Adiponectin (ng/mL) | |||
| Body mass index ≤ 25 | 9.94 ± 3.44 | 11.67 ± 6.36 | 0.031 |
| Body mass index > 25 | 8.58 ± 3.68 | 11.40 ± 6.21 | 0.001 |
| Total Mean ± SD | 9.15 ± 3.63 | 11.60 ± 6.30 | 1.2 × 10-5 |
| Family history | 0.005 | ||
| Positive | 52 (29.71) | 30 (17.14) | |
| Negative | 123 (70.29) | 145 (82.86) | |
| Histological report | |||
| Estrogen receptor | |||
| Positive | 144 (82.29) | ||
| Negative | 31 (17.71) | ||
| Progesterone receptor | |||
| Positive | 106 (60.57) | ||
| Negative | 69 (39.43) | ||
| Type of cancer | |||
| Invasive lobular carcinoma (ILC) | 60 (34.29) | ||
| Invasive ductal carcinoma (IDC) | 70 (40) | ||
| Ductal carcinoma in situ (DCIS) | 31 (17.71) | ||
| Lobular carcinoma in situ (LCIS) | 14 (8) | ||
| Stage of cancer | |||
| I | 34 (19.43) | ||
| II | 26 (14.86) | ||
| II/III | 41 (23.43) | ||
| III | 47 (26.85) | ||
| Metastasis | 27 (15.43) |
Frequency Distribution of Demographic and Biochemistry Characteristics Among Cancer Patients and Controls a
4.1. Polymorphisms and Breast Cancer
Moreover, the relationship between SNPs and breast cancer was evaluated, using logistic regression analysis as shown in Table 3.
| Variables | Cases (n = 158) | Controls (n = 158) | Crude Odds Ratio (95% CI) | P Value | Adjusted Odds Ratio (95% CI) | P Value |
|---|---|---|---|---|---|---|
| Leptin G2548A rs7799039 | ||||||
| GG | 83 (47.43) | 112 (64) | Referent | |||
| AA | 60 (34.29) | 41 (23.43) | 0.506 (0.311 - 0.825) | 0.006 | 0.565 (0.325 - 0.980) | 0.042 |
| GA | 32 (18.28) | 22 (12.57) | 0.509 (0.276 - 0.940) | 0.031 | 0.598 (0.304 - 1.175) | 0.136 |
| Alleles | ||||||
| G | 198 (56.57) | 246 (70.29) | Referent | |||
| A | 152 (43.43) | 104 (29.71) | 0.579 (0.424 - 0.791) | 0.001 | ||
| Leptin R Q223R rs1137101 | ||||||
| AA | 116 (66.29) | 140 (80) | Referent | |||
| GG | 50 (28.57) | 25 (14.29) | 0.414 (0.242 - 0.711) | 0.001 | 0.375 (0.208 - 0.678) | 0.001 |
| AG | 9 (5.14) | 10 (5.71) | 0.921 (0.362 - 2.342) | 0.862 | 0.958 (0.347 - 2.639) | 0.933 |
| Alleles | ||||||
| A | 241 (68.86) | 290 (82.86) | Referent | |||
| G | 109 (31.14) | 60 (17.14) | 0.464 (0.324 - 0.664) | 2.7 × 10- 5 | ||
| Leptin R K109R rs1137100 | ||||||
| AA | 94 (53.71) | 118 (67.43) | Referent | |||
| GG | 67 (38.29) | 50 (28.57) | 0.594 (0.377 - 0.938) | 0.025 | 0.644 (0.389 - 1.066) | 0.087 |
| AG | 14 (8) | 7 (4) | 0.398 (0.155 - 1.027) | 0.057 | 0.529 (0.186 - 1.503) | 0.232 |
| Alleles | ||||||
| A | 202 (57.71) | 243 (69.43) | Referent | |||
| G | 148 (42.29) | 107 (30.57) | 0.624 (0.457 - 0.851) | 0.003 | ||
| Leptin R K656N rs8179183 | ||||||
| GG | 81 (46.29) | 118 (67.43) | Referent | |||
| CC | 55 (31.43) | 25 (14.29) | 0.312 (0.180 - 0.541) | 3.4 × 10-5 | 0.373 (0.206 - 0.676) | 0.001 |
| GC | 39 (22.28) | 32 (18.28) | 0.563 (0.326 - 0.973) | 0.039 | 0.605 (0.329 - 1.110) | 0.104 |
| Alleles | ||||||
| G | 201 (57.43) | 268 (76.57) | Referent | |||
| C | 149 (42.57) | 82 (23.43) | 1.412 (1.048 - 1.902) | 0.023 | ||
| Adiponectin T45G rs2241766 | ||||||
| TT | 111 (63.43) | 138 (78.86) | Referent | |||
| GG | 39 (22.29) | 25 (14.28) | 0.516 (0.294 - 0.903) | 0.021 | 0.496 (0.265 - 0.927) | 0.028 |
| TG | 25 (14.28) | 12 (6.86) | 0.386 (0.186 - 0.803) | 0.011 | 0.425 (0.192 - 0.942) | 0.035 |
| Alleles | ||||||
| T | 247 (70.57) | 288 (82.29) | Referent | |||
| G | 103 (29.43) | 62 (17.71) | 0.523 (0.366 - 0.749) | 4 × 10-4 | ||
| Adiponectin G276T rs1501299 | ||||||
| GG | 96 (54.86) | 119 (68) | Referent | |||
| TT | 38 (21.71) | 23 (13.14) | 0.488 (0.272 - 0.875) | 0.016 | 0.416 (0.222 - 0.781) | 0.006 |
| GT | 41 (23.43) | 33 (18.86) | 0.649 (0.382 - 1.105) | 0.111 | 0.648 (0.353 - 1.191) | 0.162 |
| Alleles | ||||||
| G | 233 (66.57) | 271 (77.43) | Referent | |||
| T | 117 (33.43) | 79 (22.57) | 0.598 (0.428 - 0.835) | 0.003 | ||
| Adiponectin C11377G rs266729 | ||||||
| CC | 120 (68.57) | 122 (69.71) | Referent | |||
| GG | 35 (20) | 29 (16.57) | 0.815 (0.469 - 1.417) | 0.468 | 0.891 (0.480 - 1.652) | 0.714 |
| CG | 20 (11.43) | 24 (13.72) | 1.180 (0.619 - 2.249) | 0.614 | 1.649 (0.777 - 3.497) | 0.192 |
| Alleles | ||||||
| C | 260 (74.29) | 268 (76.57) | Referent | |||
| G | 90 (25.71) | 82 (23.43) | 0.912 (0.647 - 1.286) | 0.6 | ||
| Adiponectin G11391A rs17300539 | ||||||
| GG | 105 (60) | 126 (72) | Referent | |||
| AA | 57 (32.57) | 41 (23.43) | 0.599 (0.372 - 0.966) | 0.036 | 0.803 (0.467 - 1.382) | 0.429 |
| GA | 13 (7.43) | 8 (4.57) | 0.513 (0.195 - 1.284) | 0.154 | 0.548 (0.201 - 1.493) | 0.240 |
| Alleles | ||||||
| G | 223 (63.71) | 260 (74.29) | Referent | |||
| A | 127 (36.29) | 90 (25.71) | 0.615 (0.445 - 0.851) | 0.003 |
Association Between Single Nucleotide Polymorphisms in LEP, LEPR, ADIPO, and Odds Ratio of Breast Cancer by Logistic Regression a
As shown in Table 4, the association between polymorphisms and breast cancer was adjusted for age, BMI, and family history. A significant association was observed in rs7799039, rs1137101, rs8179183, and rs1501299.
| Genotypes | Case (%) | Controls (%) | P-Value | Odds Ratio (CI 95%) |
|---|---|---|---|---|
| Leptin G2548A rs7799039 | ||||
| GG | 47.43 | 64 | 1 (Referent) | |
| AA | 34.29 | 23.43 | 0.024 | 2.001 (1.098-3.647) |
| GA | 18.28 | 12.57 | 0.069 | 1.967 (0.948-4.080) |
| Leptin R Q223R rs1137101 | ||||
| AA | 66.29 | 80 | 1 (Referent) | |
| GG | 28.57 | 14.29 | 0.004 | 2.537 (1.356-4.747) |
| AG | 5.14 | 5.71 | 0.705 | 1.249 (0.395-3.946) |
| Leptin R K109R rs1137100 | ||||
| AA | 53.71 | 67.43 | 1 (Referent) | |
| GG | 38.29 | 28.57 | 0.053 | 1.713(0.993-2.953) |
| AG | 8 | 4 | 0.203 | 2.036(0.681-6.091) |
| Leptin R K656N rs8179183 | ||||
| GG | 46.29 | 67.43 | 1 (Referent) | |
| CC | 31.43 | 14.29 | 0.001 | 2.842 (1.510-5.351) |
| GC | 22.28 | 18.28 | 0.120 | 1.676 (0.873-3.218) |
| Adiponectin T45G rs2241766 | ||||
| TT | 63.43 | 78.86 | 1 (Referent) | |
| GG | 22.29 | 14.28 | 0.055 | 1.892 (0.986-3.630) |
| TG | 14.28 | 6.86 | 0.121 | 1.951 (0.837-4.545) |
| Adiponectin G276T rs1501299 | ||||
| GG | 54.86 | 68 | 1 (Referent) | |
| TT | 21.71 | 13.14 | 0.005 | 2.654 (1.352-5.208) |
| GT | 23.43 | 18.86 | 0.462 | 1.278 (0.665-2.456) |
| Adiponectin C11377G rs266729 | ||||
| CC | 68.57 | 69.71 | 1 (Referent) | |
| GG | 20 | 16.57 | 0.518 | 1.239 (0.648-2.369) |
| CG | 11.43 | 13.72 | 0.195 | 0.583 (0.257-1.319) |
| Adiponectin G11391A rs17300539 | ||||
| GG | 60 | 72 | 1 (Referent) | |
| AA | 32.57 | 23.43 | 0.807 | 1.076 (0.597-1.940) |
| GA | 7.43 | 4.57 | 0.248 | 1.839 (0.654-5.171) |
The Association Between Single Nucleotide Polymorphisms in Leptin and Adiponectin Gene and Risk of Breast Cancer After Adjustment for Age, Body Mass Index, and Family History
In another analysis, the relationship between BMI and SNPs was analyzed via logistic regression. Both homozygous mutant and heterozygous LEPG2548A genotypes had a significant relationship with BMI and increased the risk of breast cancer by 2.200 and 2.279 folds, respectively. Moreover, with an increase in BMI, the risk of breast cancer increased by 2.426, 2.899, and 2.796 folds in carriers of LEPRQ223R, LEPRK656N, and AdipoG276T genotypes, respectively, compared to individuals with other genotypes.
Concerning the variable of age in the two subgroups (< 50 and ≥ 50 years), ORs of 2.662, 2.970, 1.990, and 2.237 were reported in breast cancer patients with advancing age, carrying homozygous mutant genotypes of LEPRQ223R, LEPRK656N, AdipoT45G, and AdipoG276T, respectively.
In addition to the above studies, the relationship between polymorphism genotypes and history of disease in first-degree relatives was studied among patients. The results showed that just the TT genotype of AdipoG276T with an OR of 3.023 (P = 0.043, OR: 3.023, CI95%: 0.992 - 9.214) and GG genotype of AdipoC11377G polymorphism with an OR of 1.374 (P = 0.049, OR: 1.374, CI95%: 0.545 - 3.462) had significant relationships with the risk of breast cancer.
4.2. Influence of Risk Allele on Serum Concentration of Hormones in Pre- and Post-menopause Samples
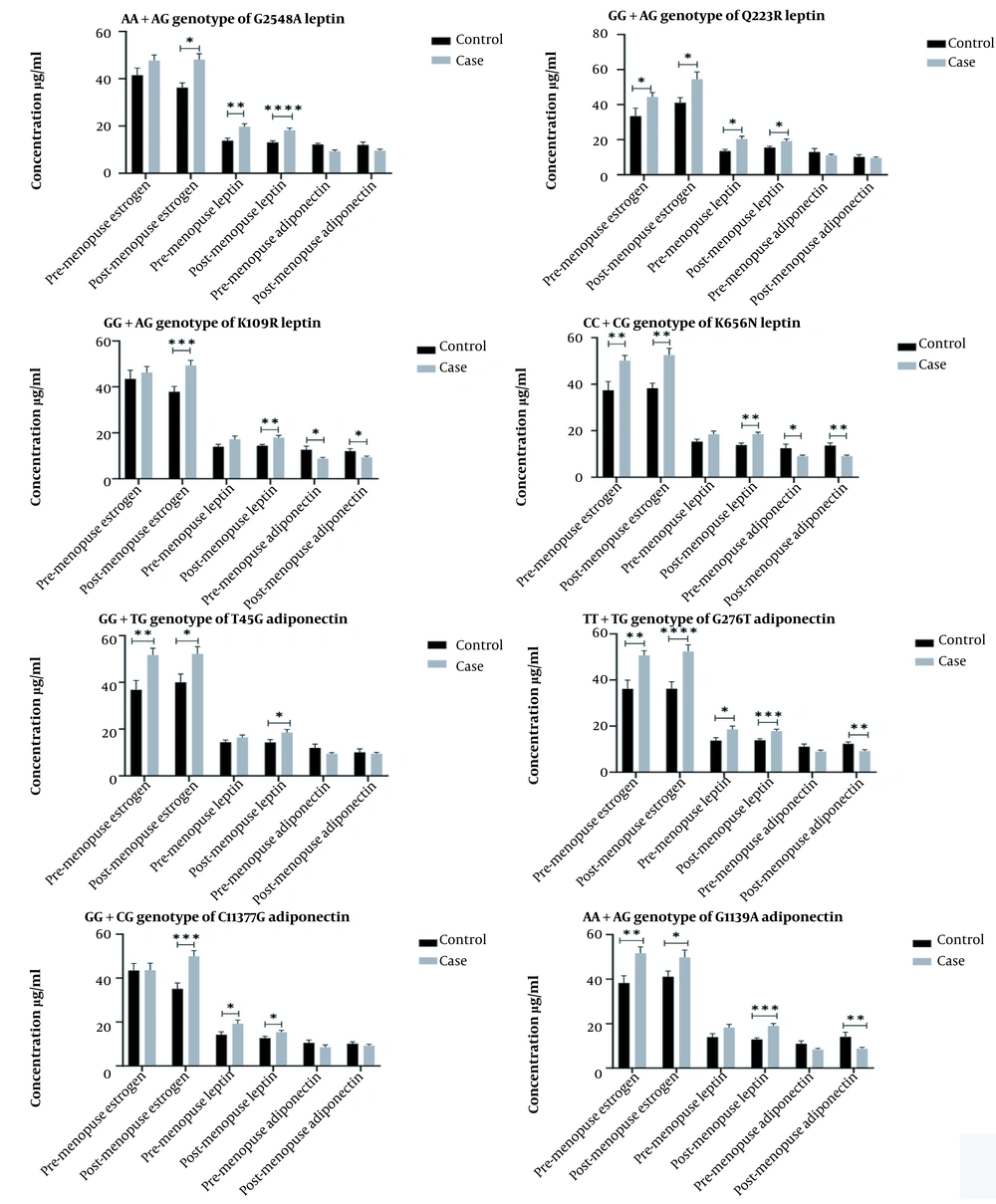
In another study, the effect of the risk alleles of studied polymorphisms on the serum concentration of the estrogen, leptin, and adiponectin hormones in premenopausal and postmenopausal women regarding the incidence of breast cancer was investigated. The effect of risk allele genotypes on serum concentration of hormones indicated changes in serum concentration of estrogen and leptin in all studied polymorphisms associated with breast cancer in postmenopausal women. Adiponectin level was affected only by polymorphisms K109R, K656N, and G276 and G11391A in postmenopausal women with breast cancer. Mean serum concentrations of hormones and their association with breast cancer are shown in Figure 1.
Influence of mutant allele of polymorphisms on serum concentration of estradiol, leptin, and adiponectin hormones in pre-menopause and post-menopause of two groups. Postmenopausal women with gene mutations are more likely to develop breast cancer. However, some mutations can affect the risk of cancer by affecting the concentration of adiponectin. * P ≤ 0.01, ** P ≤ 0.001, *** P ≤ 0.0001, **** P ≤ 0.00001
5. Discussion
In this study, the effects of BMI and genetic variations of leptin and adiponectin genes on the serum concentrations of estradiol, leptin, and adiponectin in postmenopausal and non-menopausal patients with breast cancer were evaluated. Because taking medication or exposure to radiation affects hormone concentrations, patients whose diseases had been diagnosed and had not started any treatment were selected.
Evaluation of genetic variations in SNPs revealed that G2548A polymorphism was associated with breast cancer risk. This finding was in line with the results reported by Cleveland et al. (21) and Tang et al. (22). On the other hand, in a study by Luan et al., no significant association (G2548A polymorphism) was found with breast cancer (23).
In a previous study of Luan, LEPR Q223R polymorphism was associated with a reduced risk of breast cancer in Asians (23). The present results also showed the firmest association between Q223R polymorphism and reduced risk of breast cancer, which is in line with the results reported by Wang et al. (24). Moreover, in a meta-analysis by Liu and Liu, LEPRQ223R was associated with the spread of breast cancer in East Asia (25). In addition, Okobia et al. attributed the increased relative risk of breast cancer before menopause to changes in leptin signaling capacity (16). Conversely, Rong et al. found no significant association between Q223R polymorphism and cancer risk (26). But, in this study, the allele frequency of LEPR K109R polymorphism was associated with a reduced risk of breast cancer. A similar finding was reported by Shi et al. (27). But Rodrigo et al. reported that LEPR K109R polymorphism increased the risk of breast cancer (OR = 4.125) (28).
In this study, LepR K656N polymorphism had the firmest allelic association with the reduced risk of breast cancer, which is in line with the results reported by Huerta et al. (29). Nevertheless, Liu et al. did not find any relationship between this SNP and cancer in a meta-analysis (30). Also, Wang et al. did not report any association between this polymorphism and breast cancer (31).
In our study, there was a significant relationship between breast cancer and risk alleles of AdipoT45G and AdipoG276T polymorphisms. Similar results were reported by Reddy in an Indian population (32), whereas Nyante et al. found no significant relationship with breast cancer (33).
Two same results were obtained for C11377G polymorphism in the gene promoter site. In a previous study of 156 samples, Adipo C11377G polymorphism was not significantly associated with breast cancer (34). Also, in the current study, no changes occurred by increasing the sample size to 350. It seems that this polymorphism is not related to breast cancer. Conversely, in a study by Liu et al., this polymorphism was associated with non-alcoholic fatty liver disease (35). Polymorphism G11391A was associated with a reduced risk of breast cancer in terms of genotypic and allelic frequencies. Furthermore, in a study by Vasseur et al., C11377G and G11391A polymorphisms were associated with the risk of type II diabetes (36).
Our findings revealed that carriers of LEPRQ223R, LEPRK656N, AdipoT45G, and AdipoG276T polymorphisms, who were above 50 years, had higher risks of developing breast cancer by 2.662, 2.970, 1.990, and 2.237 folds, respectively. With increasing BMI, ORs of cancer development in individuals with AA and AG genotypes of LepG2548A polymorphism were 2.200 and 2.279, respectively. Also, ORs of cancer in individuals with GG genotype of LEPRQ223R, CC genotype of LEPRK656N, and TT genotype of AdipoG276T were 2.426, 2.899, and 2.796 folds, respectively. The relationship between polymorphism genotype and family history of patients showed ORs of developing breast cancer in individuals carrying GC genotype of LEPRK656N, homozygous TT mutant of AdipoG276T, and GG genotype of AdipoC11377G were 1.352, 3.023, and 1.374, respectively.
The results showed that elevated serum levels of estradiol and leptin, besides the decreased level of adiponectin, were associated with the risk of breast cancer; similar results were reported by Chen et al. (37). Furthermore, in a meta-analysis by Pan et al., leptin was identified as an important biomarker in women at risk of breast cancer, especially in overweight/obese or postmenopausal women (38). In addition, by a meta-analysis, Gu et al. found that serum leptin level might play an essential role in the pathogenesis and metastasis of breast cancer and could be used as a suitable therapeutic target for breast cancer treatment (39). On the contrary, Hu et al. (40) and Assiri and Kamel found an inverse association between serum leptin concentration and breast cancer risk in premenopausal women (12).
The study of the effect of risk alleles on serum concentration of estradiol and leptin hormones showed that postmenopausal individuals were associated with breast cancer for all studied polymorphisms. But, the serum level of adiponectin was only affected by K109R, K656N, and G276T and G11391A polymorphisms. During perimenopause, polymorphisms Q223R, K656N, T45G, and G276T and G11391A affected serum level of estradiol, polymorphisms G2548A, Q223R, and G276T and C11377G affected leptin level, and K109R, K656N affected adiponectin serum level.
The association of menopausal status with the risk of breast cancer and leptin level has been reported in several studies. Some research has indicated an inverse relationship between leptin level and menopausal status. Also, in a study by Pan et al., leptin level was associated with a higher risk of breast cancer in postmenopausal women (38), while no significant association was observed in premenopausal women. In addition, a study by Ando et al. revealed the role of postmenopausal estradiol (4).
In terms of BMI, there was a significant difference between premenopausal and postmenopausal women in the control and cancer groups, and a significant association was observed in the BMI > 25. In line with the present study, the finding of Renehan et al. indicated a specific association between breast cancer and BMI increase in premenopausal and postmenopausal women in Asia-Pacific (8). Some recent studies have suggested BMI as a risk factor for breast cancer in postmenopausal women (41-44) similar to this study. In contrast, Rinaldi et al. reported an inverse relationship between BMI and breast cancer risk (44). Van den Brandt et al. also found a strong association between BMI and breast cancer risk in postmenopausal women. Also, a negative relationship was reported in premenopausal women (45).
5.1. Conclusions
The current results indicated that weight gain and obesity, along with polymorphisms in leptin and adiponectin genes, are major contributing factors to the development of breast cancer. Also, findings suggest that the expression of estradiol, leptin and adiponectin should be reduced in premenopausal women by following a healthy body weight to prevent breast cancer development during menopause.