1. Background
Escherichia coli is a Gram-negative, rod-shaped bacterium that typically resides in the lower intestinal tract of humans. It is also found in hospital environments and can cause nosocomial infections (1). Escherichia coli is one of the most frequent causes of urinary tract infection (2, 3) and is among the most important pathogens causing bloodstream infections (4), otitis media, wound infections, neonatal meningitis, and nosocomial pneumonia (5, 6). Escherichia coli is a major cause of waterborne and foodborne human diarrhea worldwide, especially in developing countries, causing several deaths, particularly in children under five-years-old (7).
Globally, antimicrobial resistance is a serious public health concern, particularly in developing countries where infectious diseases, malnutrition, and poverty are endemic (8-10). It is one of the main causes of failure in the treatment of infectious diseases, including infections caused by E. coli, resulting in increased morbidity, mortality, and cost of healthcare services (11). The emergence of multidrug-resistant E. coli is a growing problem around the world (12). The prevalence and susceptibility profile of clinical isolates of E. coli show substantial variations in geographical locations, as well as significant differences in various populations, clinical samples, and environments (13).
The periodic epidemiology of E. coli to determine antibiotic resistance patterns in patients with urinary tract infections has been studied thoroughly in the region (3, 14). However, investigating antimicrobial susceptibility patterns of E. coli isolated from various clinical samples in our region is spares. The routine monitoring of antibiotic resistance patterns of E. coli from different specimens could help develop E. coli empirical treatment guidelines in the region.
2. Objectives
The present study was conducted to determine the sensitivity pattern of E. coli isolated from various clinical sources in Duhok city, Iraq.
3. Methods
3.1. Study Design and Specimen Collection
The study was conducted from January 2017 to February 2020 in Duhok city, Iraq. A total of 454 samples (418 urine, 18 wounds, 7 cervical, 4 blood, 3 semen, 2 ascitic fluid, and 2 cerebral spinal fluid samples) were collected from both genders (133 males and 321 females). The age of the participants ranged from 10 to 60 years. All clinical specimens were collected from participants referring to private health clinical centers in Duhok city, Iraq. All participants were chosen as clinically positive for E. coli and processed according to standard operating procedures.
Clean-catch midstream urine specimens were obtained using sterile disposable glass containers (5 mL) from patients to avoid contamination. Blood samples were taken from patients aseptically in blood culture tubes. Additionally, wound and cervical samples were collected using sterile cotton swabs. Semen, ascetic, and cerebral spinal fluid samples were collected in sterile wide-mouth containers.
3.2. Bacterial Identification and Antimicrobial Sensitivity
Escherichia coli isolates were initially identified by their morphological characteristics on MacConkey agar based on standard microbiological culture as per the Clinical and Laboratory Standards Institute (CLSI) guidelines (15). Identified E. coli was then confirmed using the Vitek-2 system (bioMerieux, US) following the manufacturer’s instructions. Antimicrobial susceptibility testing was performed on Mueller-Hinton agar (Oxoid Limited, Hampshire, England) using the disk diffusion (Kirby Bauer’s) technique following the CLSI guidelines. The inclusion criteria included patients from both genders, older than 10-years-old, with positive microbiological evidence of E. coli isolated from various samples, and agreement to participate in the study. Patients who did not agree to participate in the study were excluded.
3.3. Ethics
The study was approved by the Ethics Committee of the College of Medicine, University of Zakho, Kurdistan Region, Iraq. Written informed consent was obtained from all participants.
4. Results
Of the 454 clinical samples, the majority of the E. coli isolates were obtained from urine samples (418; 92.2%), followed by wound (18; 3.9%), cervical (7; 1.5%), blood (4; 0.9%), semen (3; 0.7%), ascitic (2; 0.4%), and cerebral spinal fluid (2; 0.4%) samples (Table 1). Escherichia coli isolates were more prevalent in females (70.7%) than in males (29.3%). In urine samples, the isolation of E. coli was higher in females (73.9%) than in males (26.1%) (Table 1). In wound samples, however, the incidence of E. coli was higher in males (55.6%) than in females (44.4%). The distribution of the isolates from other clinical samples in both genders is summarized in Table 1.
| Source of Isolation | No. (%) | Male (No.%) | Female (No.%) |
|---|---|---|---|
| Urine | 418 (92.2) | 109 (26.1) | 309 (73.9) |
| Wound swab | 18 (3.9) | 10 (55.6) | 8 (44.4) |
| Cervical swab | 7 (1.5) | 7 (100) | 0 (0) |
| Blood | 4 (0.9) | 1 (25) | 3 75) |
| Semen | 3 (0.7) | 3 (100) | 0 (0) |
| Ascitic fluid | 2 (0.4) | 1 (50) | 1 (50) |
| Cerebral Spinal Fluid | 2(0.4) | 2 (100) | 0 (0) |
| Total | 454 (100) | 133 (29.3) | 321 (70.7) |
The Occurrence of Escherichia coli Isolated from Various Clinical Specimens Based on Sex
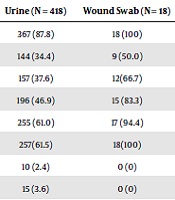
The antimicrobial sensitivity pattern in urine samples showed that E. coli isolates were highly sensitive to ertapenem (97.6%) and imipenem (96.4%) while resistant to ampicillin (87.8%), cefepime (61.5%), and ceftriaxone (61.0%) (Table 2). Escherichia coli isolates from wound samples were highly resistant to ampicillin (100%), cefepime (100%), and ceftriaxone (94.4%), whereas they showed 100% sensitivity to ertapenem and imipenem. Escherichia coli isolates from cervical samples were sensitive to ertapenem (100%) and imipenem (100%) (Table 2). In addition, 100% of E. coli isolates from blood samples were resistant to ampicillin, cefoxitin, and ceftriaxone and 75% of them were sensitive to ertapenem, ciprofloxacin, and levofloxacin (Table 2). The results of the antibiotic susceptibility pattern of isolates from other clinical samples are shown in Table 2.
| Antibiotic | Number of Isolates Recovered from Clinical Sources (Percent (%) of Resistance) | |||||
|---|---|---|---|---|---|---|
| Urine (N = 418) | Wound Swab (N = 18) | Cervical Swab (N = 7) | Blood (N = 4) | Semen (N = 3) | Other Samplesa (N = 4) | |
| Ampicillin | 367 (87.8) | 18 (100) | 5 (71.4) | 4 (100) | 2 (66.7) | 4 (100) |
| Amoxicillin/clavulanic acid | 144 (34.4) | 9 (50.0) | 3 (42.9) | 3 (75) | 2 (66.7) | 3 (75) |
| Cefazolin | 157 (37.6) | 12(66.7) | 3 (42.9) | 3 (75) | 0 (0) | 0 (0) |
| Cefoxin | 196 (46.9) | 15 (83.3) | 3 (42.9) | 4 (100) | 2 (66.7) | 4 (100) |
| Ceftriaxone | 255 (61.0) | 17 (94.4) | 4(57.1) | 4 (100) | 2 (66.7) | 3 (75) |
| Cefeprine | 257(61.5) | 18(100) | 4(57.1) | 3 (75) | 2 (66.7) | 3 (75) |
| Ertapenem | 10 (2.4) | 0 (0) | 0 (0) | 1 (25) | 0 (0) | 0 (0) |
| Imipenem | 15 (3.6) | 0 (0) | 0 (0) | 3 (75) | 0 (0) | 0 (0) |
| Gentamicin | 116 (27.8) | 8 (44.4) | 2 (28.6) | 3 (75) | 2 (66.7) | 3 (75) |
| Tobramycin | 156 (37.3) | 12 (66.7) | 3 (42.9) | 2 (50) | 2 (66.7) | 2 (50) |
| Ciprofloxacin | 186 (44.5) | 12 (66.7) | 4(57.1) | 1 (25) | 2 (66.7) | 2 (50) |
| Levofloxacin | 98 (23.5) | 5 (27.8) | 2 (28.6) | 1 (25) | 0 (0) | 0 (0) |
| Nitrofurantoin | 43 (10.3) | 2 (11.1) | 1 (14.3) | 4 (100) | 0 (0) | 0 (0) |
| Trimethoprim/ Sulfamethoxazole | 203 (48.6) | 8 (44.4) | 2 (28.6) | 4 (100) | 2 (66.7) | 1 (25) |
Antibiotic Resistance Profiles of Multiresistant Escherichia coli Isolated from Various Clinical Sources
The overall susceptibility patterns of E. coli isolates from various clinical sources are presented in Table 3. It was found that 88.3%, 63.3%, and 63.9% of the isolated E. coli strains were resistant to ampicillin, ceftriaxone, and cefepime, respectively. On the other hand, E. coli showed to be sensitive to ertapenem (97.6%), imipenem (96.1%), and nitrofurantoin (88.9%) (Table 3).
| Antibiotic | Susceptibility Patterns (N = 454) | |
|---|---|---|
| Resistant No. (%) | Sensitive No. % | |
| Ampicillin | 400 (88.3) | 54 (11.7) |
| Amoxicillin/clavulanic acid | 164 (36.2) | 290 (63.8) |
| Cefazolin | 175 (38.6) | 279 (61.4) |
| Cefoxin | 224 (49.4) | 230 (50.6) |
| Ceftriaxone | 285 (63.9) | 169 (37.1) |
| Cefepime | 287 (63.3) | 167 (36.7) |
| Ertapenem | 11 (2.4) | 443 (97.6) |
| Imipenem | 18 (3.9) | 436 (96.1) |
| Gentamicin | 218 (48.1) | 236 (51.9) |
| Tobramycin | 177 (38.9) | 277 (61.1) |
| Ciprofloxacin | 206 (45.4) | 248 (54.6) |
| Levofloxacin | 106 (23.4) | 348 (76.6) |
| Nitrofurantoin | 50 (11.1) | 404 (88.9) |
| Trimethoprim/Sulfamethoxazole | 220 (48.5)) | 234 (51.5) |
Overall Antimicrobial Resistance Patterns of Escherichia coli Isolated from Various Clinical Specimens
5. Discussion
Escherichia coli is one of the most common causative agents of bacterial infections (16). Antimicrobial resistance patterns of E. coli continue to pose a great threat to public health worldwide and lead to serious health problems such as prolonged hospitalization and treatment failure (8, 10). Therefore, this study aimed to detect the antibiotic susceptibility profile of E. coli isolates from various clinical sources in Duhok city, Iraq.
In the present study, a total of 454 E. coli isolates were collected from various clinical specimens. The frequency of clinical isolates of E. coli in urine samples was higher in females than in males. This result is consistent with other studies reporting a higher prevalence of E. coli in UTI in females (14, 17, 18). The reason for the high prevalence of this microorganism in females is that the urethra of females are short and this shortens the distance to be moved by bacteria to the bladder and sexual activities, which increases the inoculation of bacteria into the bladder (19). These predisposing factors of UTI are accelerated by poor hygiene and low socioeconomic status (17). Alteration in the vaginal microflora may play a major role in encouraging the colonization of the vagina with coliforms and this can be associated with UTI (20). In terms of wound swabs, the occurrence of clinical isolates of E. coli was higher in males than in females. In contrast, in a study conducted in Pakistan, the incidence of E. coli isolates from wound swabs was the same in both males and females (5). It is difficult to explain this variation, and further studies with larger sample sizes are needed to explore the reason.
In the current study, E. coli isolated from various clinical specimens, showed differences in antibiotic sensitivity patterns. In the case of urine samples, the antibiotic sensitivity profile showed that E. coli isolates were extremely sensitive to ertapenem and imipenem and highly resistant to ampicillin. This result is in agreement with previous studies showing that E. coli isolates from urine samples were highly resistant to ampicillin (5) and highly sensitive to imipenem (21). In another study conducted in Iraq, the majority of clinical E. coli isolates from urine samples were resistant to amoxicillin/clavulanic acid (3). The same study found that E. coli isolates were sensitive to amikacin and imipenem (3). Other studies found that 100% of E. coli isolates were sensitive to gentamycin, amikacin, imipenem, meropenem, piperacillin-tazobactam, and tobramycin (22). Previous studies conducted in India and Kenya also showed high sensitivity to gentamycin (23, 24). Our findings are alarming and call for urgent measures to control the threatening development of antibacterial resistance, particularly to ampicillin, in the region.
In this study, ertapenem and imipenem were found to be the most effective antimicrobials against E. coli isolates from the wound and cervical swabs, whereas the microorganism showed high resistance to ampicillin and cefepime. Additionally, it was observed that the E. coli isolates had relatively high resistance to ciprofloxacin. The blood isolates were found to be 100% resistant to ampicillin, cefoxitin, ceftriaxone, and nitrofurantoin and about 75% of them were sensitive to levofloxacin, ertapenem, and ciprofloxacin. Our results are in agreement with a study conducted in Ethiopia (25) that found that E. coli isolates were resistant to ampicillin (100%). In all other clinical specimens, including semen, ascetic, and cerebral spinal fluid specimens, E. coli showed high sensitivity to cefazolin, ertapenem, imipenem, levofloxacin, and nitrofurantoin. These results are in agreement with other research (26).
Antimicrobial resistance rates obtained in this study were higher than the resistance patterns reported by previous studies (27, 28). This may be attributed to the prevailing use and misuse of antibiotics in the area under study. On the other hand, E. coli isolates from various sources were sensitive to ertapenem, imipenem, and nitrofurantoin. This is in contrast to previous studies performed in different countries that found gentamicin, nitrofurantoin, ciprofloxacin, norfloxacin, and chloramphenicol as the most effective antimicrobial agents against E. coli (29, 30). Other studies reported that the isolates showed a high level of resistance to ampicillin, cefuroxime, amoxicillin-clavulanic acid, ceftriaxone, ciprofloxacin, and cefepime (31). The same study found that the isolates were highly sensitive to imipenem, nitrofurantoin, amikacin, chloramphenicol, piperacillin-tazobactum, gentamicin, azetreonam, and norfloxacin (31). These differences in the sensitivity pattern of antibiotics could be attributed to the time difference between the two studies, population variations, and significant differences in the sample sizes and types,
5.1. Conclusion
The high prevalence and spread of infection in females can be reduced by proper hygienic and medical care. In this study, ertapenem, imipenem, and nitrofurantoin were found to be the most effective antibiotics against E. coli isolates from various clinical sources. However, E. coli isolates exhibited high resistance to ampicillin, ceftriaxone, and cefepime. Therefore, it is proposed to perform antibiotic susceptibility testing to ensure effective prescriptions.