1. Introduction
Non-tuberculosis mycobacteria (NTM) are ubiquitous environmental organisms, which were introduced as human pathogenic in 1990’s. This group of bacteria can result in various infection including pulmonary, cutaneous, brain, eye, blood stream, and catheter-related infection in immune-compromised patients and even healthy individuals (1, 2).
According to the literature, several reports of drug-resistant Mycobacterium tuberculosis (DR-TB) from developing countries are wrong. In fact, Mycobacterium spp. was not identified at species level in these laboratories, and anti-tuberculosis drugs were routinely recommend for these patients. However, given that non-tuberculosis mycobacteria (NTM) are resistant to anti-tuberculosis drugs; the patients were not cured and were misdiagnosed with DR-TB (3). According to the American thoracic society (ATS) guideline, identification and differentiation of clinical isolates of Mycobacterium spp. is essential for final diagnosis, appropriate treatment, patient management, clinical significance, and epidemiological studies (1, 4).
Mycobacterium abscessus was originally classified as subspecies of Mycobacterium chelonae by Moore and Frerichs in 1953; 23 later, this bacterium was reclassified as novel species by Kubica et al. Today, M. abscessus is considered as a group of rapidly growing mycobacteria (RGM), which are emerged as an important pathogen in humans and recognized as the cause of approximately 65% - 80% of lung disease induced by rapidly growing mycobacteria. M. abscessus is resistant to a wide range of antibiotics and its treatment is very difficult (5, 6). Here, we describe a case of M. abscessus pulmonary infection mimicking pulmonary tuberculosis in a healthy patient with a history of previous pulmonary TB.
2. Case Presentation
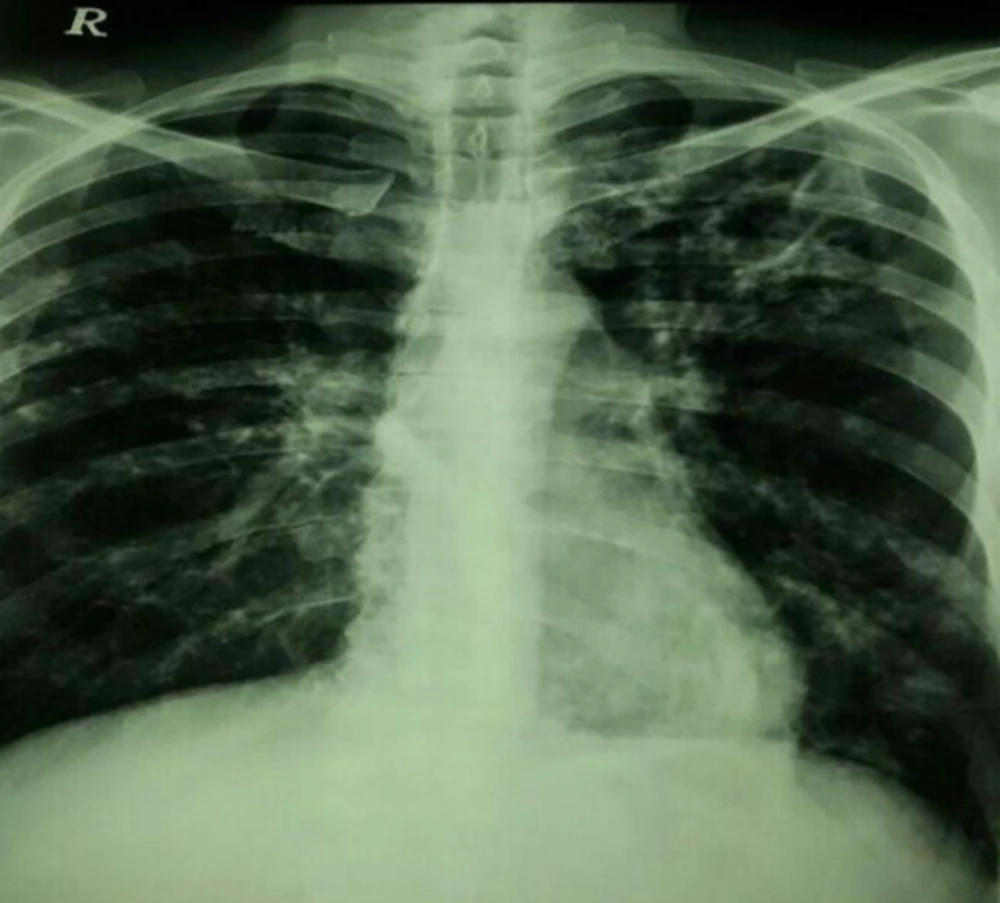
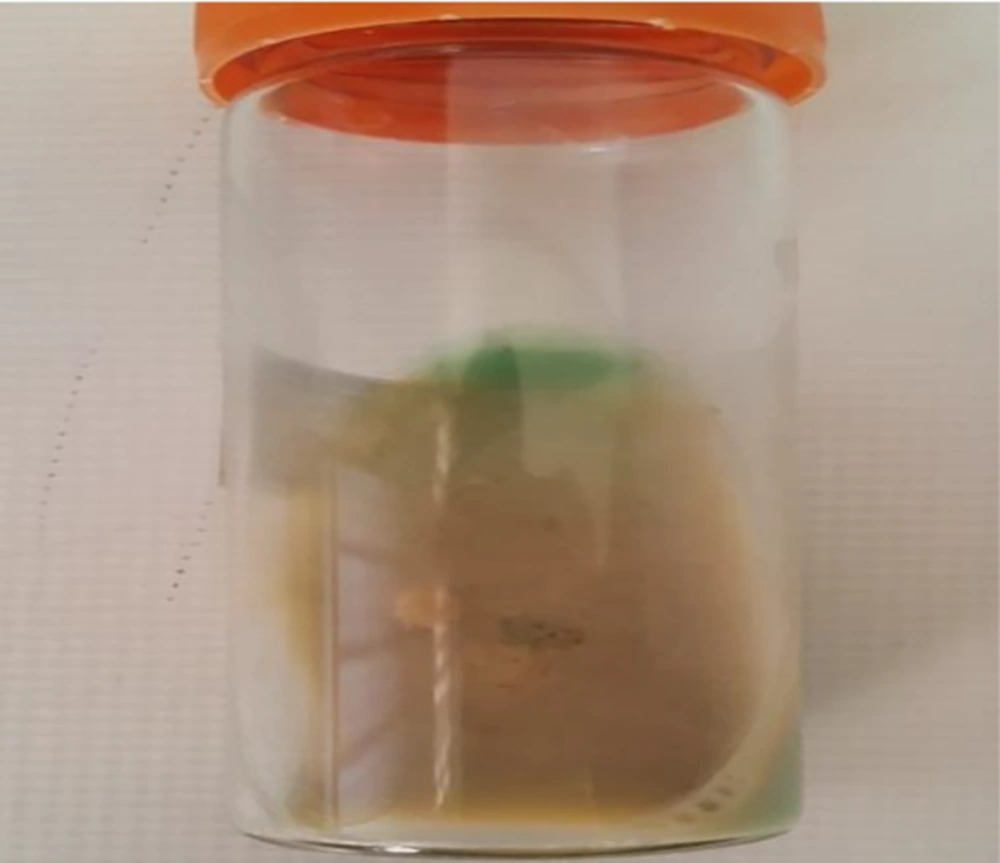
An 85-year-old woman admitted to the hospital due to productive cough, dyspnea, fever, night sweats, weigh lost, hemoptysis, chest pain, anorexia, vomiting, and hematuria is presented here. She had a history of previous pulmonary tuberculosis, however, no evidence for human immunodeficiency virus (HIV) infection or immune-disorder was observed. Chest X-ray showed lung diffuse infiltrates lesions (Figure 1), laboratory examination results were as follows: WBC 8,300 μL, RBC 4.17 μL × 106, platelet 297,000 μL, Hemoglobin 10.9 g/dL and Hematocrit 33.1%; other laboratories indexes: BUN 24 mg/dL, creatinine 0.9 mg/dL, calcium 7.5 mg/dL, phosphorus 3.4 mg/dL, sodium 127 mEq/L, potassium 2.2 mEq/L, AST 26 U/L, ALT 17 U/L, CPK 55 U/L, CPK-MB 9 U/L, and troponin I < 25 ng/L. Urine culturing was negative and direct smear examination of sputum revealed the presence of acid-fast bacilli (AFB); based on the previous clinical history, reactivation of tuberculosis was assumed and anti-tuberculosis therapy with isoniazid, rifampin, and ethambutol was started. However, the patient did not improve and in the culture of sputum, mycobacterium spp. grew in Lowenstein-jensen and Sauton’s agar (Figure 2). The isolate was identified using phenotypic tests consisting growth rate of < 7 days, ability for growth on MacConkey agar without crystal violet, production of urease, positive for iron uptake, Aryl-sulfatase (3 days), heat stable catalase, hydrolysis of tween 80 and negative results for pigment, niacin production, and nitrate reduction. Following DNA extraction (simple boiling method), amplification and sequencing of the nearly full length of 16S rRNA gene and partial segment of hsp65 gene was carried out for this isolate as previously described (1). The hsp65 gene sequence of the isolate showed 100% similarities with the corresponding sequence of mycobacterium abscessus strain ATCC 19977. The GenBank accession number of this isolate is MG930482.
The antibiotic susceptibility testing was performed using micro-broth dilution as recommended by clinical and laboratory standards institute (CLSI) (7). The antibiotic sensitivity test showed that the isolate was sensitive to amikacin, cefoxitin, and linezolid. Moreover, this isolate exhibited resistance to clarithromycin and doxycycline. Finally, the patient was treated with linezolid (600 mg twice a day), amikacin (15 mg/kg/d), and cefoxitin (200 mg/kg/d) for six months. The patient was completely recovered after two weeks.
2.1. Ethical Statement
Formal consent for publication was obtained from the patient.
3. Discussion
Given the lack of expert technicians in TB endemic developing countries, mycobacterial infection could be diagnosed as M. tuberculosis based on the presence of acid fast bacilli in direct smear microbiology. Consequently, the patients would be treated with anti-tuberculosis drugs and regarding no response to TB-drugs, the disease may be presumed as drug-resistant TB (3). Identification and differentiation of NTM using phenotypic test are difficult to standardize, in addition to its high cost, need to expertise staff, being time-consuming, labor-intensive; it may be even wrong in some cases. However, molecular methods such as PCR-restriction fragment length polymorphism (PCR-RFP) and sequencing using housekeeping genes (for example 16S rRNA, 16S-23S rRNA internal transcribed spacer (ITS), rpoB, hsp65, recA, …) are inexpensive, rapid, accurate, and reliable for identification of NTM to species level (1).
M. abscessus was first identified by Moore and Frerichs from an infection of the knee with subcutaneous abscess-like lesions as subspecies of M. chelonae. However, according to the studies of Kusunoki and Ezaki, it was determined as novel distinct species as M. abscessus. Nowadays, the taxonomic scientists declared that Mycobacterium abscessus comprise three closely related subspecies of M. abscessus subsp. abscessus, M. abscessus subsp. Massiliense, and M. abscessus subsp. bolletii. All members of M. ascessus complex were isolated from clinical specimens, particularly lung disease (6, 8). According to the literature, isolation rate of M. abscessus and M. massiliense are approximately equal, however, M. bolleti is rare; clinical manifestation of M. abscessus complex are similar, nonetheless, antibiotic resistance (particularly, clarithromycin) in M. abscessus subsp. abscessus is more common (6, 9).
M. abscessus is one of the important groups of RGM (Rapidly growing mycobacteria), which are able to cause serious infection in immune-compromised patients as well as healthy people. This group of mycobacteria can contaminate medical instruments and cause disseminated infection in patients who use catheters and other medical equipment (2, 10). One of the main points about M. abscessus-caused human infection is its antibiotic resistance. Many NTM species are susceptible to new macrolides and can be treated with clarithromycin and azithromycin. However, unfortunately, M. abscessus infections are inefficient to macrolide-based chemotherapy due to overexpression of erm (erythromycin ribosomal methylase gene) or production of biofilm (9-12). Combinational therapy of amikacin with cefoxitin or imipenem and oral macrolides has been recommended by American thoracic society and infectious diseases society of America. However, insufficient treatment response rates of M. abscessus infections, optimal therapeutic regimens, and treatment durations are not well established and challengeable (4-6). Paradoxically, there has been evidence for person-to-person transmission of M. abscessus between patients with cystic fibrosis (12, 13).
In conclusion, M. abscessus can cause pulmonary infection mimicking pulmonary tuberculosis. The present study emphasized the problems about identification of NTM in developing countries, and highlighted misdiagnosis of NTM pulmonary infection with tuberculosis. Precise identification should be determined based on the standard microbiologic protocols, however, previous history of patients and clinical manifestation are not completely reliable. Furthermore, molecular methods such as sequencing of the 16S rRNA, rpoB, and hsp65 can help us in rapid (in urgent cases), reliable, and efficient identification of NTM at the species level.