1. Background
Based on GLOBOCAN statistics, breast cancer was the most prevalent cancer type in 2020 (1). As breast cancer is one of the first diagnosed cancer types in history (2), numerous studies have addressed its causes and treatments. Numerous studies support a strong relationship between estrogen exposure levels and breast cancer risk (3, 4). Estrogens are steroid molecules (e.g., estradiol, estriol, estrone, and estretrol) capable of regulating the physiological processes due to their estrogen receptors (ERs). The overexpression of ERs has been reported in more than 70% of breast cancer cases (5). Two isoforms of ERs, namely estrogen receptor alpha (ERα) and estrogen receptor beta, have been identified, which belong to the type 1 nuclear receptors. Both of these receptors are composed of six functional domains (A-F) with different genetic codes and expression profiles (6). The exact role of ERs in breast cancer is complicated. The ERα can increase cell proliferation in the breast tissue through various signaling pathways, such as genomic and nongenomic pathways, that accelerate gene expression (7, 8).
Recent studies have suggested the role of hydroquinone metabolites from estrogen (mainly estradiol-3, 4-quinone) in deoxyribonucleic acid (DNA) mutations affecting cancer development. These metabolites react with DNA through the Michael reaction, resulting in DNA depurination (9). On the other hand, ER antagonists are the first choice in the endocrine therapy of breast cancer.
Selective estrogen receptor modulators (SERMs) inhibit proliferation as a full antagonist in breast tissue; however, they play an agonist role in the bone and cardiovascular systems, which is favorable (10, 11). Five categories of SERMs have been explored based on their chemical structures, namely triphenylethylenes (tamoxifen-like), benzothiophenes (e.g., raloxifene and arzoxifene), tetrahydronaphthalene (e.g., lasofoxifene), indoles (e.g., bazedoxifene), and benzopyrans (e.g., EM-800) (12-14). Structure-Activity Relationship studies have revealed that two or three phenyl rings are connected to a central core of most SERMs; an aminoethoxy pharmacophore is placed on one of the rings (15-18). These compounds insert into the active site of the ER located in the ligand-binding domain or E domain. The presence of an aminoethoxy is essential to inhibit the ER. When a SERM compound binds to the active site, the aminoethoxy side chain prevents receptor activation by Helix-12 (17, 19).
Recent studies have introduced benzo[d]imidazo[2,1-b]thiazole derivatives as prominent anticancer agents with high cytotoxic effects on diverse cell lines (including MCF-7) (20, 21).
2. Objectives
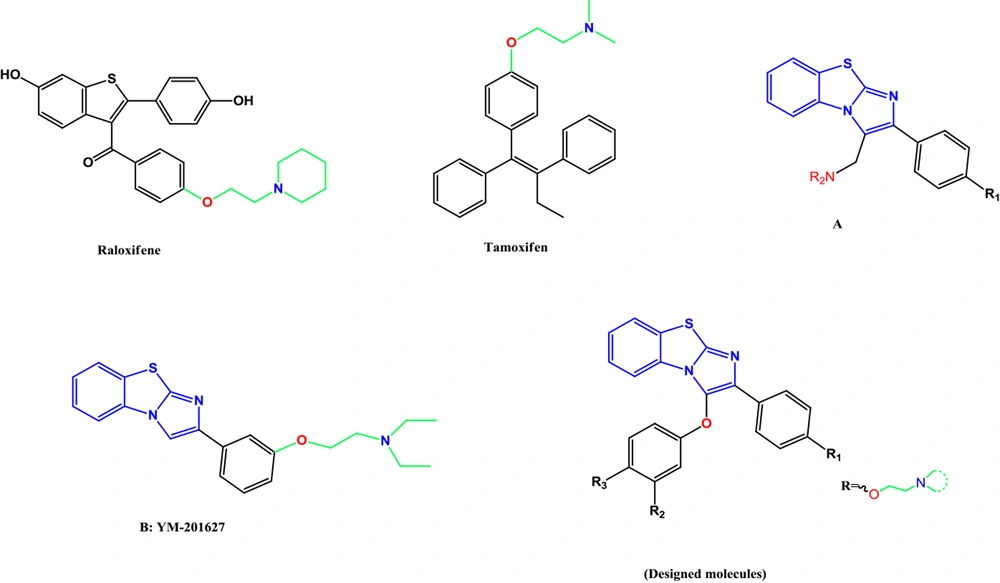
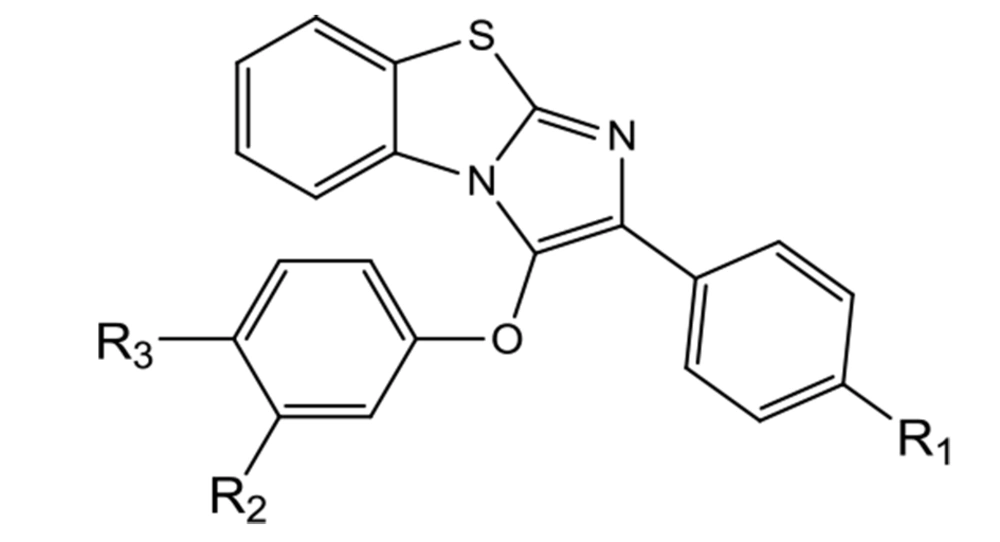
This study designed a new series of diaryl benzo[d]imidazo[2,1-b]thiazole derivatives. These compounds have various aminoethoxy side chains on one of their phenyl rings (para- or meta-positions) (Figure 1).
3. Methods
3.1. Materials and Methods
All the applied chemicals were supplied from Merck AG and Sigma-Aldrich. If necessary, solvents were dried by standard methods. The Thomas-Hoover capillary apparatus was employed to measure the melting point. Fourier-transform infrared spectroscopy (FTIR) studies were completed by Cary 630 FTIR spectrometer using potassium bromide (KBr) pellets for solid samples. 1H-Nuclear magnetic resonance (NMR) and 13C-NMR spectra were recorded by a Bruker FT-500 MHz instrument (Brucker Biosciences, USA). Chloroform-D was employed as the solvent; however, tetramethylsilane served as the internal standard. Coupling constant (J) values were reported in hertz (Hz); nonetheless, spin multiples were presented as s, d, t, q, bs, and m corresponding to the singlet, doublet, triplet, quartet, broad singlet, and multiplet, respectively. A 6410 Agilent LC-MS triple quadrupole mass spectrometer [to perform liquid chromatography-mass spectrometry (LC-MS)] equipped with electrospray ionization (ESI) interface was also employed to obtain mass spectral measurements.
3.2. Chemistry
3.2.1. Synthesis of 2-Bromoacetophenone Derivatives (2a-2b)
1 eq Bromine was diluted in 15 mL methanol, which was then dropwise added into a solution of appropriate acetophenone (20 mmol, 1a or 1b) in methanol (200 mL) at room temperature. When the bromination reaction was completed [monitored by thin-layer chromatography (TLC), mobile phase: methylene chloride (DCM)], the mixture was slowly added to a water solution of saturated NaHCO3. The crystalline product was filtered, repeatedly washed with water, and dried under vacuum to give desired product 2a-2b.
3.2.2. General Procedure for Synthesis of 3a-3c
4-Methoxy phenol (22 mmol, 1.1 eq) and 30 mmol (1.5 eq) K2CO3 were suspended in acetone under stirring at 50°C for 30 minutes. Then, 20 mmol (1 eq) 2a was added into the suspension and allowed to reflux for 5 hours. After completing the reaction [checked by TLC, mobile phase: CHCl3/n-Hexane (95:5)], the solvent evaporated, and the resulting residues were repeatedly washed with water and methanol to give pure compound 3a. The same procedure was used for the synthesis of 3b and 3c. The reaction of 2b with 3-methoxy phenol in the presence of K2CO3 resulted in 3b, and the reaction of 2c with phenol and K2CO3 led to the formation of 3c.
2-(4-methoxyphenoxy)-1-phenylethanone (3a)
Yield, 92%; white crystalline powder; mp: 66 - 67°C; IR (KBr): ν (cm-1) 1692 (CO); LC-MS (ESI) m/z: 265 (M+23).
2-(3-methoxyphenoxy)-1-phenylethanone (3b)
Yield, 78%; pale beige crystalline powder; mp: 86°C; IR (KBr): ν (cm-1) 1696 (CO); LC-MS (ESI) m/z: 265 (M+23).
1-(4-methoxyphenyl)-2-phenoxyethanone (3c)
Yield, 80%; white crystalline powder; mp: 65 - 67°C; IR (KBr): ν (cm-1) 1688 (CO); LC-MS (ESI) m/z: 265 (M+23).
3.2.3. General Procedure for Synthesis of Benzo[d]imidazo[2,1-b]thiazole Derivatives (4a-4c)
2-Aminobenzothiazole (15 mmol, 3 eq) and elemental iodine (10 mmol, 2 eq) were added to the stirring solution of compounds 3a, 3b, or 3c (5 mmol, 1 eq) in 15 ml dimethylformamide. The obtained mixture underwent 24 hours of reflux. When the reaction was completed, as the TLC monitoring indicated, 1 mL of saturated sodium thiosulfate was added to the solution, followed by stirring until the disappearance of the brown color. Then, the solution was poured into the beaker containing ice. For 4b and 4c (solid products), the precipitate was separated and recrystallized in methanol to give the pure product. However, 4a (oily product) was extracted with ethyl acetate, evaporated, and used for the next step without purification.
3-(4-methoxyphenoxy)-2-phenylbenzo[d]imidazo[2,1-b]thiazole (4a)
Yield, 65%; dark orange oil; IR: ν (cm-1) 1632 (C=N); LC-MS (ESI) m/z: 373 (M+1).
3-(3-methoxyphenoxy)-2-phenylbenzo[d]imidazo[2,1-b]thiazole (4b)
Yield, 71%; yellow crystalline powder; mp: 98 - 100 °C; IR (KBr): ν (cm-1) 1692 (C=N); LC-MS (ESI) m/z: 373 (M+1).
2-(4-methoxyphenyl)-3-phenoxybenzo[d]imidazo[2,1-b]thiazole (4c)
Yield, 75%; yellow crystalline powder; mp: 132 °C; IR (KBr): ν (cm-1) 1632 (C=N); LC-MS (ESI) m/z: 373 (M+1).
3.2.4. General Procedure for Synthesis of Phenolic Benzo[d]imidazo[2,1-b]thiazole Derivatives (5a-5c)
In this study, 4 mmol benzoimidazo[2,1-b]thiazole compound 4a (or 4b or 4c) was dissolved in anhydrous DCM (50 mL) and cooled to -10°C utilizing an ice-NaCl-acetone bath. Then, 24 mmol BBr3 was added and stirred under an argon atmosphere for 2 hours. Then, the mixture was quenched by adding a few drops of methanol; subsequently, it was poured into the ice-containing beaker and stirred for 10 minutes. The residues were collected and dried in the oven to obtain 5a-5c.
4-((2-phenylbenzo[d]imidazo[2,1-b]thiazol-3-yl)oxy)phenol (5a)
Yield, 95%; white powder; mp: 272 - 275°C (decomposed); IR (KBr): ν (cm-1) 3023 (OH); LC-MS (ESI) m/z: 359 (M+1).
3-((2-phenylbenzo[d]imidazo[2,1-b]thiazol-3-yl)oxy)phenol (5b)
Yield, 90%; light beige powder; mp: 288 - 293°C (decomposed); IR (KBr): ν (cm-1) 3049, 3325 (OH); LC-MS (ESI) m/z: 359 (M+1).
4-(3-phenoxybenzo[d]imidazo[2,1-b]thiazol-2-yl)phenol (5c)
Yield, 94%; white powder; mp: 297°C (decomposed); IR (KBr): ν (cm-1) 3056, 3433 (OH); LC-MS (ESI) m/z: 359 (M+1), 739 (2M+23).
3.2.5. General Procedure for Synthesis of 6a-6l
In this study, 1 mmol of 5a (or 5b or 5c) and 3 mmol K2CO3 were added into acetone (30 mL) and vigorously stirred at 60°C for 1 hour. Then, 3 mmol of appropriate 2-chloroethylamine was transferred into the suspension and stirred for 5 hours under reflux conditions. After the reaction was completed (TLC monitoring), the solvent was evaporated, and the final product was extracted using ethyl acetate. Finally, the crude products were purified using plate chromatography (mobile phase: CHCl3/CH3OH, 95:5, and ethyl acetate).
N,N-Dimethyl-2-(4-((2-phenylbenzo[d]imidazo[2,1-b]thiazol-3-yl)oxy)phenoxy)ethanamine (6a)
Yield, 69%; yellow oil; IR: ν (cm−1) 1029 (C-N), 1498, 1603 (aromatic C=C); 1H-NMR (CDCl3): δ ppm 2.325 (s, 6H, CH3), 2.697-2.720 (t, 2H, -NCH2), 3.981-4.003 (t, 2H, -OCH2), 6.850 - 6.868 (d, 2H, phenoxy H3 & H5, J= 9 Hz), 6.985 - 7.003 (d, 2H, phenoxy H2 & H6, J = 9 Hz), 7.208 - 7.272 (m, 3H, phenyl H3 & H4 and H5), 7.333 - 7.363 (t, 2H, phenyl H2 & H6), 7.457-7.475 (m, 1H, imidazobenzothiazole H7), 7.643 - 7.649 (m, 1H, imidazobenzothiazole H6), 7.897 - 7.913 (d, 2H, imidazobenzothiazole H5 & H8, J = 7.9 Hz); 13C-NMR (CDCl3): δ ppm 45.90, 45.97, 58.35, 66.38, 115.86, 116.10, 124.03, 124.21, 124.95, 125.55, 125.71, 126.40, 127.19, 128.77, 130.08, 131.78, 132.18, 132.78, 134.28, 142.25, 150.65, 155.13; LC-MS (ESI) m/z: 430 (M+1).
N,N-Diethyl-2-(4-((2-phenylbenzo[d]imidazo[2,1-b]thiazol-3-yl)oxy)phenoxy)ethanamine (6b)
Yield, 77%; yellow oil; IR: ν (cm−1) 1021 (C-N), 1498, 1603 (aromatic C=C); 1H-NMR (CDCl3): δ ppm 1.070 - 1.098 (t, 6H, diethylamine CH3), 2.664 - 2.706 (q, 4H, diethylamine CH2), 2.886-2.909 (t, 2H, -NCH2), 4.012-4.036 (t, 2H, -OCH2), 6.831 - 6.849 (d, 2H, phenoxy H3 & H5, J = 9.1 Hz), 6.982 - 7.001 (d, 2H, phenoxy H2 & H6, J = 9.1 Hz), 7.205-7.266 (m, 3H, phenyl H3 & H4 and H5), 7.330 - 7.361 (t, 2H, phenyl H2 & H6), 7.456 - 7.475 (m, 1H, imidazobenzothiazole H7), 7.630 - 7.648 (m, 1H, imidazobenzothiazole H6), 7.893 - 7.912 (d, 2H, imidazobenzothiazole H5 & H8, J = 7.1 Hz); 13C-NMR (CDCl3): δ ppm 11.49, 47.84, 51.73, 66.69, 113.61, 115.87, 116.10, 124.09, 124.92, 125.65, 126.38, 127.16, 128.73, 130.09, 131.79, 132.20, 132.79, 134.28, 142.23, 150.70, 155.02; LC-MS (ESI) m/z: 458 (M+1).
2-Phenyl-3-(4-(2-(piperidin-1-yl)ethoxy)phenoxy)benzo[d] imidazo[2,1-b]thiazole (6c)
Yield, 58%; yellow oil; IR: ν (cm−1) 1036 (C-N), 1498, 1603 (aromatic C=C); 1H-NMR (CDCl3): δ ppm 1.431 (bs, 2H, piperidine -CH2-γ), 1.588 - 1.610 (t, 4H, -CH2-β), 2.498 (s, 4H, piperidine -NCH2-α), 2.736 - 2.759 (t, 2H, -NCH2), 4.023 - 4.047 (t, 2H, -OCH2), 6.833-6.851 (d, 2H, phenoxy H3 & H5 , J = 9.1 Hz), 6.981-6.999 (d, 2H, phenoxy H2 & H6, J = 9.1 Hz), 7.206-7.263 (m, 3H, phenyl H3 & H4 and H5), 7.332-7.363 (t, 2H, phenyl H2 & H6), 7.458-7.477 (m, 1H, imidazobenzothiazole H7), 7.627 - 7.649 (m, 1H, imidazobenzothiazole H6), 7.900 - 7.914 (d, 2H, imidazobenzothiazole H5 & H8 , J= 7.2 Hz); 13C-NMR (CDCl3): δ ppm 25.90, 55.15, 58.02, 66.39, 113.70, 115.86, 116.14, 124.01, 124.22, 125.71, 126.40, 127.19, 128.77, 130.08, 131.78, 132.18, 132.79, 134.28, 142.25, 150.63, 155.13; LC-MS (ESI) m/z: 470 (M+1).
4-(2-(4-((2-Phenylbenzo[d]imidazo[2,1-b]thiazol-3-yl)oxy)phenoxy)ethyl)morpholine (6d)
Yield, 80%; light beige crystalline powder; mp: 82°C; IR(KBr): ν (cm−1) 1021 (C-N), 1498, 1603 (aromatic C=C); 1H-NMR (CDCl3): δ ppm 2.555 (bs, 4H, morpholine -NCH2), 2.755 - 2.777 (t, 2H, -NCH2), 3.709 - 3.727 (t, 4H, morpholine -OCH2), 4.021 - 4.043 (t, 2H, -OCH2), 6.835 - 6.854 (d, 2H, phenoxy H3 & H5, J = 9.1 Hz), 6.987 - 7.005 (d, 2H, phenoxy H2 & H6, J = 9.1 Hz), 7.207 - 7.276 (m, 3H, phenyl H3 & H4 and H5), 7.331 - 7.362 (t, 2H, phenyl H2 & H6), 7.463 - 7.481 (m, 1H, imidazobenzothiazole H7), 7.636 - 7.655 (m, 1H, imidazobenzothiazole H6), 7.895 - 7.910 (d, 2H, imidazobenzothiazole H5 & H8, J = 7.2 Hz); 13C-NMR (CDCl3): δ ppm 54.18, 57.79, 66.34, 66.98, 113.58, 115.89, 116.18, 124.12, 124.93, 125.64, 126.36, 127.18, 128.74, 130.12, 131.80, 132.21, 132.77, 134.27, 142.24, 150.75, 155.04 ; LC-MS (ESI) m/z: 472 (100), 965 (2M+23).
N,N-Dimethyl-2-(3-((2-phenylbenzo[d]imidazo[2,1-b]thiazol-3-yl)oxy)phenoxy)ethanamine (6e)
Yield, 80%; yellow oil; IR: ν (cm−1) 1252 (C-N), (1677 C=N); 1H-NMR (CDCl3): δ ppm 2.385 (s, 6H, CH3), 2.776 - 2.796 (t, 2H, -NCH2), 4.057 - 4.078 (t, 2H, -OCH2), 6.639 - 6. 677 (m, 3H, phenoxy H4-H6), 7.192 - 7.277 (m, 4H, phenoxy H2 and phenyl H3-H5), 7.330 - 7.361 (t, 2H, phenyl H2 & H6), 7.437 - 7.455(m, 1H, Imidazobenzothiazole H7), 7.635 - 7.654 (m, 1H, imidazobenzothiazole H6), 7.879 - 7.893 (d, 2H, imidazobenzothiazole H5 & H8, J = 7.3 Hz); 13C-NMR (CDCl3): δ ppm 45.64, 58.02, 65.72, 125.00, 125.57, 125.73, 126.45, 127.25, 128.78, 130.08, 131.03, 131.73, 132.29, 132.70, 133.54, 142.42, 157.82, 160.42; LC-MS (ESI) m/z: 430 (M+1)
N,N-Diethyl-2-(3-((2-phenylbenzo[d]imidazo[2,1-b]thiazol-3-yl)oxy)phenoxy)ethanamine (6f)
Yield, 74%; yellow oil; IR: ν (cm−1) 1260 (C-N), 1491, 1588 (aromatic C=C); 1H-NMR (CDCl3): δ ppm 1.039 - 1.067 (t, 6H, diethylamine CH3), 2.619 - 2.661 (q, 4H, diethylamine CH2), 2.835 - 2.859 (t, 2H, -NCH2), 4.003 - 4.027 (t, 2H, -OCH2), 6.636 - 6.667 (m, 3H, phenoxy H4-H6), 7.182 - 7.271 (m, 4H, phenoxy H2 and phenyl H3-H5), 7.333 - 7.364 (t, 2H, phenyl H2 & H6), 7.346 - 7.455(m, 1H, imidazobenzothiazole H7), 7.629 - 7.647 (m, 1H, imidazobenzothiazole H6), 7.888 - 7.903 (d, 2H, imidazobenzothiazole H5 & H8, J = 7.6 Hz); 13C-NMR (CDCl3): δ ppm 11.60, 47.91, 51.61, 66.61, 102.36, 107.44, 113.67, 124.19, 124.97, 125.57, 125.73, 126.44, 127.23, 128.78, 130.08, 130.98, 131.74, 132.31, 132.73, 133.57, 142.39, 157.82, 160.63; LC-MS (ESI) m/z: 458 (M+1)
2-Phenyl-3-(3-(2-(piperidin-1-yl)ethoxy)phenoxy)benzo[d] imidazo[2,1-b]thiazole (6g)
Yield, 60%; yellow oil; IR: ν (cm−1) 1260 (C-N), 1595 (aromatic C=C); 1H-NMR (CDCl3): δ ppm 1.441 (bs, 2H, piperidine -CH2-γ), 1.606 - 1.628 (t, 4H, piperidine -CH2-β), 2.513 (bs, 4H, piperidine -NCH2-α), 2.745 - 2.767 (t, 2H, -NCH2), 4.061-4.084 (t, 2H, -OCH2), 6.636 - 6.666 (m, 3H, phenoxy H4-H6), 7.164 - 7.283 (m, 4H, phenoxy H2 and phenyl H3-H5), 7.336 - 7.366 (t, 2H, phenyl H2 & H6), 7.446 - 7.464 (m, 1H, imidazobenzothiazole H7), 7.642 - 7.660 (m, 1H, imidazobenzothiazole H6), 7.886 - 7.901 (d, 2H, imidazobenzothiazole H5 & H8, J = 7.4 Hz); 13C-NMR (CDCl3): δ ppm 25.67, 55.14, 57.80, 65.99, 124.98, 125.57, 125.74, 126.44, 127.24, 128.78, 130.09, 130.98, 131.75, 132.31, 132.71, 133.58, 142.40, 157.83, 160.58; LC-MS (ESI) m/z: 470 (M+1).
4-(2-(3-((2-Phenylbenzo[d]imidazo[2,1-b]thiazol-3-yl)oxy)phenoxy)ethyl)morpholine (6h)
Yield, 48%; yellow oil; IR: ν (cm−1) 1021 (C-N), 1498, 1603 (aromatic C=C), 1677 (C=N); 1H-NMR (CDCl3): δ ppm 2.534 (bs, 4H, morpholine -NCH2), 2.731 - 2.752 (t, 2H, -NCH2), 3.703 - 3.720 (t, 4H, morpholine -OCH2), 4.028 - 4.050 (t, 2H, -OCH2), 6.637 - 6.690 (m, 3H, phenoxy H4-H6), 7.197 - 7.279 (m, 4H, phenoxy H2 and phenyl H3-H5), 7.335 - 7.366 (t, 2H, phenyl H2 & H6), 7.442 - 7.461 (m, 1H, Imidazobenzothiazole H7), 7.641 - 7.660 (m, 1H, Imidazobenzothiazole H6), 7.886 - 7.901 (d, 2H, Imidazobenzothiazole H5 & H8, J = 7.5 Hz); 13C-NMR (CDCl3): δ ppm 54.19, 57.62, 66.92, 67.05, 102.15, 109.82, 113.56, 124.23, 124.98, 125.55, 125.72, 126.40, 127.24, 128.76, 130.09, 131.00, 131.72, 132.30, 132.69, 133.54, 142.40, 157.83, 160.54; LC-MS (ESI) m/z: 472 (M+1)
N,N-Dimethyl-2-(4-(3-phenoxybenzo[d]imidazo[2,1-b]thiazol-2-yl)phenoxy)ethanamine (6i)
Yield, 88%; yellow oil; IR: ν (cm−1) 1692 (C=N); 1H-NMR (CDCl3): δ ppm 2.329 (s, 6H, CH3), 2.712 - 2.734 (t, 2H, -NCH3), 4.048 - 4.071 (t, 2H, -OCH3), 6.894 - 6.912 (d, 2H, phenoxyethylamine H2 & H6, J = 8.8 Hz), 7.068-7.108 (m, 3H, phenoxyethylamine H3 & H5 and phenoxy H4), 7.232 - 7.261 (m, 2H, phenoxy H3 & H5), 7.306 - 7.338 (t, 2H, phenoxy H2 & H6), 7.422 - 7.440 (m, 1H, imidazobenzothiazole H7), 7.622 - 7.640 (m, 1H, imidazobenzothiazole H6), 7.803 - 7.820 (d, 2H, imidazobenzothiazole H5 & H8, J = 8.7 Hz); 13C-NMR (CDCl3): δ ppm 45.98, 46.05, 58.38, 65.99, 113.40, 114.86, 115.16, 123.85, 124.80, 125.58, 126.35, 126.82, 127.03, 130.02, 130.45, 131.83, 132.29, 132.87, 142.13, 156.82, 158.13; LC-MS (ESI) m/z: 430 (M+1)
N,N-Diethyl-2-(4-(3-phenoxybenzo[d]imidazo[2,1-b]thiazol-2-yl)phenoxy)ethanamine (6j)
Yield, 78%; yellow oil; IR: ν (cm−1) 1029 (C-N), 1491 (C=C), 1692 (C=N); 1H-NMR (CDCl3): δ ppm 1.120 - 1.148 (t, 6H, diethylamine CH3), 2.707 - 2.750 (q, 4H, diethylamine CH2), 2.950 - 2.974 (t, 2H, -NCH2), 4.112 - 4.137 (t, 2H, -OCH2), 6.919 - 6.936 (d, 2H, phenoxyethylamine H2 & H6, J = 8.8 Hz), 7.114-7.156 (m, 3H, phenoxyethylamine H3 & H5 and phenoxy H4 ), 7.275 - 7.306 (m, 2H, phenoxy H3 & H5), 7.354 - 7.387 (t, 2H, phenoxy H2 & H6), 7.476 - 7.494 (m, 1H, imidazobenzothiazole H7), 7.676 - 7.694 (m, 1H, imidazobenzothiazole H6), 7.845 - 7.863 (d, 2H, imidazobenzothiazole H5 & H8, J = 8.8 Hz ); 13C-NMR (CDCl3): δ ppm 11.68, 47.86, 51.68, 66.22, 113.40, 114.82, 115.15, 123.83, 124.09, 124.78, 125.61, 126.33, 126.97, 130.04, 130.43, 131.85, 132.30, 132.91, 142.12, 156.84, 158.03; LC-MS (ESI) m/z: 458 (M+1), 937 (2M+23).
3-Phenoxy-2-(4-(2-(piperidin-1-yl)ethoxy)phenyl)benzo[d] imidazo[2,1-b]thiazole (6k)
Yield, 50%; yellow oil; IR: ν (cm−1) 1692 (C=N); 1H-NMR (CDCl3): δ ppm 1.460 (bs, 2H, piperidine -CH2-γ), 1.650 - 1.672 (t, 4H, piperidine -CH2-β), 2.604 (s, 4H, piperidine -NCH2-α), 2.843 - 2.865 (t, 2H, -NCH2 ), 4.147 - 4.170 (t, 2H, -OCH2), 6.872 - 6.890 (d, 2H, phenoxyethylamine H2 & H6, J = 8.8 Hz), 7.065 - 7.109 (m, 3H, phenoxyethylamine H3 & H5 and phenoxy H4), 7.237 - 7.266 (m, 2H, phenoxy H3 & H5), 7.307 - 7.339 (t, 2H, phenoxy H2 & H6 ), 7.425 - 7.444 (m, 1H, imidazobenzothiazole H7), 7.627 - 7.645 (m, 1H, imidazobenzothiazole H6), 7.799 - 7.817 (d, 2H, imidazobenzothiazole H5 & H8, J = 8.9 Hz); 13C-NMR (CDCl3): δ ppm 25.57, 54.99, 57.78, 65.56, 113.43, 114.87, 115.14, 123.87, 124.82, 125.72, 126.37, 126.86, 127.07, 130.02, 130.46, 131.82, 132.22, 132.91, 142.15, 156.80, 157.84; LC-MS (ESI) m/z: 470 (M+1), 939 (2M+1).
4-(2-(4-(3-Phenoxybenzo[d]imidazo[2,1-b]thiazol-2-yl)phenoxy)ethyl)morpholine (6l)
Yield, 78%; yellow oil; IR: ν (cm−1) 1029 (C-N); 1H-NMR (CDCl3): δ ppm 2.574 (bs, 4H, morpholine -NCH2), 2.782 - 2.804 (t, 2H, -NCH2), 3.719 - 3.737 (t, 4H, morpholine -OCH2), 4.096 - 4.118 (t, 2H, -OCH2), 6.880 - 6.898 (d, 2H, phenoxyethylamine H2 & H6, J = 8.8 Hz), 7.069 - 7.112 (m, 3H, phenoxyethylamine H3 & H5 and phenoxy H4), 7.241 - 7.250 (m, 2H, phenoxy H3 & H5), 7.310 - 7.342 (t, 2H, phenoxy H2 & H6), 7.428 - 7.446 (m, 1H, imidazobenzothiazole H7), 7.631 - 7.649 (m, 1H, imidazobenzothiazole H6), 7.805 - 7.823 (d, 2H, imidazobenzothiazole H5 & H8, J = 8.8 Hz ); 13C-NMR (CDCl3): δ ppm 54.20, 57.75, 65.82, 67.02, 113.39, 114.87, 115.13, 123.85, 124.01, 124.22, 124.82, 125.70, 126.35, 126.84, 127.05, 130.01, 130.46, 131.80, 132.21, 132.89, 142.15, 156.80, 157.95; LC-MS (ESI) m/z: 472 (M+1)
3.3. Cytotoxicity Evaluation
The cytotoxicity of the prepared compounds was determined on the MCF-7 cell line (human breast cancer cell line, supplied from the Iranian Biological Resources Center) using the MTT (3-[4,5-dimethylthiazol-2-yl]-2,5-diphenyl tetrazolium bromide) method (22). First, the cells were cultured by incubation in RPMI 1640 medium, 10% fetal bovine serum, and 100 µg/mL streptomycin at 37°C and 5% CO2. The MTT assay was carried out to determine the percentage of dead cells. For this purpose, 100 µlL of culture medium containing MCF-7 cells (10,000 cells per well) was poured into each well of the 96-well plate and incubated at 37°C and 5% CO2 for 24 hours. Then, the synthesized compounds (6a-6l) (10 μM), tamoxifen (positive control), and dimethyl sulfoxide (DMSO) (negative control) were added to the cell-containing wells. After 24 hours of incubation at 37°C, 10 μL MTT solution (0.5 mg/mL) was added to all wells, and the microplates were incubated for an extra 3 hours. Finally, the purple crystals of formazan were dissolved in 100 μL DMSO, and their optical absorbance was recorded at 570 nm using an ELISA reader (Infinite® M200, TECAN). The percentage of inhibition was calculated.
3.4. Molecular Modeling Studies
Docking simulations were carried out utilizing the AutoDock software (version 4) to characterize the interactions between the synthesized compounds and the active site of the ER. The 3D crystal structure of ER in complex with raloxifene [protein data bank (PDB) ID: human estrogen receptor ligand-binding domain in complex with raloxifene (1ERR)] was acquired from the RCSB PDB. For the protein preparation, all the ligands and water molecules were eliminated; Kollman charges and polar hydrogens were also added to the protein structure. Then, the PDB file was converted to a protein data bank, partial charge (Q) and atom type (T) (PDBQT) format with the help of AutoDock tools. All the ligands were sketched using HyperChem software (version 8.0), and the geometry of these compounds was optimized using the MM+ method. Finally, the partial charges were calculated, nonpolar hydrogens were merged, and the PDB ligand files were converted to the PDBQT formats using AutoDock tools.
The receptor grid box was set according to the binding site of raloxifene (PDB ID: 1ERR) with a grid space of 7 Å. The Lamarckian genetic algorithm was used with 100 runs. After molecular docking, the ligand-receptor complexes were further analyzed using PyMOL (version 1.1.7) (23).
4. Results and Discussion
In this study, novel benzo[d]imidazo[2,1-b]thiazole derivatives (6a-6l) were designed as SERMs and synthesized through five reaction steps (Figure 1). The structures were confirmed by FTIR, 1H-NMR, 13C-NMR, and LC-MS techniques. As SERMs are valuable agents in breast cancer treatment, the cytotoxicity of all compounds was evaluated on a breast cancer cell line to reveal their anticancer potentials. Docking studies of these compounds with ER were also included (Figure 2).
Synthesis of benzo[d]imidazo[2,1-b]thiazole derivatives: reagents and conditions: A, Br2, CH3OH, RT; B, K2CO3 (1.5 eq), phenol or 4-methoxyphenol (1.1 eq), acetone, reflux, 5 hours; C, I2 (2 eq), 2-aminobenzothiazole (3 eq), dimethylformamide, reflux, 24 hours; D, BBr3 (5 eq), methylene chloride, -10°C; E, K2CO3 (3 eq), different 2-chloroethylamines (3 eq), acetone, reflux, 5 hours.
The cytotoxicity of 6a-6l was evaluated on the MCF-7 cell line using the MTT assay. According to Figure 3 and Table 1, the 24-hour treatment of MCF-7 cells with all compounds (10 μM) led to the significant inhibition of cells’ growth. Most of these compounds showed higher cytotoxicity against MCF-7 than tamoxifen (i.e., positive control). Among the aforementioned compounds, 6i and 6j showed the highest potency in inhibiting the proliferation of the tested cell line with 81% and 77% inhibition rates, respectively. Moreover, the cytotoxic effects were evaluated on the fibroblast cell line with a concentration of 100 µM. Based on the results, most synthesized compounds demonstrated less cytotoxicity than tamoxifen.
For the investigation of the possible mechanism of action, docking of the most potent compound (6i) was performed using AutoDock software (version 4). The docking results revealed that this compound with a free energy of -9 kJ/mol interacted well with the ER active site and inhibited the receptor (Figure 4). The nitrogen atom of the aminoethoxy side-chain established a hydrogen bond with the aspartate-351 amino acid, which is the most critical interaction to inhibit the ER (distance: 5.81 Å). The phenoxy ring showed a π-π interaction with the phenylalanine-404 residue (distance: 3.17 Å). Moreover, the nitrogen atoms of the imidazobenzothiazole are adjacent to the histidine-524 residue; therefore, they can establish a hydrogen bonding with this amino acid.
Since the ER is highly expressed in MCF-7 cells, these compounds have exerted their antiproliferative effects on this cell line by inhibiting the ER. Docking studies also support this hypothesis. However, binding studies are required to determine the mechanism of action of these compounds.
| Compound | R1 | R2 | R3 | % Inhibition MCF-7 (10 µM) | % Inhibition Fibroblast (100 µM) |
|---|---|---|---|---|---|
| 6a | H | H | Dimethylaminoethoxy | 59.07 | 36.33 |
| 6b | H | H | Diethylaminoethoxy | 69.67 | 49 |
| 6c | H | H | Piperidinoethoxy | 62.67 | 39.33 |
| 6d | H | H | Morpholinoethoxy | 72.4 | 40.33 |
| 6e | H | Dimethylaminoethoxy | H | 55.13 | 40.67 |
| 6f | H | Diethylaminoethoxy | H | 65.27 | 44.67 |
| 6g | H | Piperidinoethoxy | H | 62.67 | 43.5 |
| 6h | H | Morpholinoethoxy | H | 63.4 | 41.33 |
| 6i | Dimethyl aminoethoxy | H | H | 81.93 | 42.17 |
| 6j | Diethyl aminoethoxy | H | H | 72.73 | 42.00 |
| 6k | Piperidinoethoxy | H | H | 65.27 | 81.83 |
| 6l | Morpholinoethoxy | H | H | 69.87 | 48.17 |
| Tamoxifen | 62.13 | 44.67 |
5. Conclusion
In summary, 12 new diaryl benzo[d]imidazo[2,1-b]thiazole derivatives were synthesized that contained aminoethoxy substitutions on one of the phenyl rings. In vitro studies indicated the significant cytotoxic impact of these compounds on the MCF-7 cell line with low toxicity on normal cells. According to docking investigations, the possible mechanism of cytotoxic effects could be the inhibition of ERs. However, it is recommended to carry out further binding studies and cytotoxicity tests on more cell lines.

![Synthesis of benzo[d]imidazo[2,1-b]thiazole derivatives: reagents and conditions: A, Br<sub>2</sub>, CH<sub>3</sub>OH, RT; B, K<sub>2</sub>CO<sub>3</sub> (1.5 eq), phenol or 4-methoxyphenol (1.1 eq), acetone, reflux, 5 hours; C, I<sub>2</sub> (2 eq), 2-aminobenzothiazole (3 eq), dimethylformamide, reflux, 24 hours; D, BBr<sub>3</sub> (5 eq), methylene chloride, -10°C; E, K<sub>2</sub>CO<sub>3</sub> (3 eq), different 2-chloroethylamines (3 eq), acetone, reflux, 5 hours. Synthesis of benzo[d]imidazo[2,1-b]thiazole derivatives: reagents and conditions: A, Br<sub>2</sub>, CH<sub>3</sub>OH, RT; B, K<sub>2</sub>CO<sub>3</sub> (1.5 eq), phenol or 4-methoxyphenol (1.1 eq), acetone, reflux, 5 hours; C, I<sub>2</sub> (2 eq), 2-aminobenzothiazole (3 eq), dimethylformamide, reflux, 24 hours; D, BBr<sub>3</sub> (5 eq), methylene chloride, -10°C; E, K<sub>2</sub>CO<sub>3</sub> (3 eq), different 2-chloroethylamines (3 eq), acetone, reflux, 5 hours.](https://services.brieflands.com/cdn/serve/3170b/1ffa066f3619c28b17f6883b130ceac19ce978b8/ijpr-127041-g002-F2-preview.webp)
![Binding modes of 6i (N,N-dimethyl-2-(4-(3-phenoxybenzo[d]imidazo[2,1-b]thiazol-2- yl)phenoxy)ethanamine) in the active site of estrogen receptor alpha [protein data bank ID: human estrogen receptor ligand-binding domain in complex with raloxifene (1ERR)]. Binding modes of 6i (N,N-dimethyl-2-(4-(3-phenoxybenzo[d]imidazo[2,1-b]thiazol-2- yl)phenoxy)ethanamine) in the active site of estrogen receptor alpha [protein data bank ID: human estrogen receptor ligand-binding domain in complex with raloxifene (1ERR)].](https://services.brieflands.com/cdn/serve/3170b/6dfa46f5937ea196898323d49ec18474a69f92da/ijpr-127041-g004-F4-preview.webp)