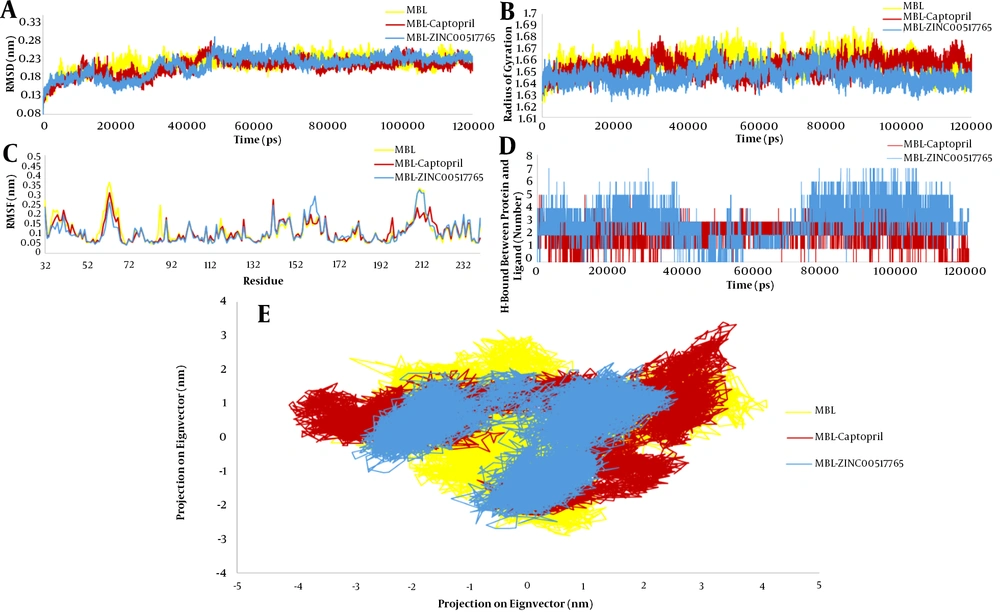
After MD simulation to evaluate the stability of the crystal structure and changes during the simulation, RMSD (
Figure 3A), Rg (
Figure 3B), RMSF (
Figure 3C), Intermolecular H-bonds (
Figure 3D), and PCA (
Figure 3E) analyses were performed on MBL-ZINC00517765, MBL-Captopril, and free MBL. The RMSD results for free MBL, MBL-ZINC00517765, and MBL-Captopril were 2.18, 2.13, and 2.09 Å, respectively. The RMSD results showed that from 50 nm to the end of the simulation, all the systems obtained stable conditions, but the stability of the MBL-ZINC00517765 complex was better than those of the other two systems. Based on the Rg analysis, the protein fluctuated during the simulations, and these changes were fewer for the MBL-ZINC00517765 complex, indicating that this complex had better stability. The Rg values for all simulations were between 16.2 and 16.8 Å, but the Rg value for MBL-ZINC00517765 was between 16.2 and 16.7 Å. In addition, RMSF analysis was performed using the simulation results, which showed that free MBL in the residues Val36 with 2.21 Å, Glu38 with 2.20 Å, Asp63 with 3.62 Å, Tyr67 with 2.32 Å, Trp87 with 2.45 Å, His116 with 1.77 Å, and Val211 with 3.32 had the highest fluctuations. Also, Val41 with 2.20 Å, Arg127 with 1.73 Å, Arg141 with 2.75 Å, Cys198 with 1.81 Å, and Leu216 with 2.37 Å had the most fluctuations in the MBL-Captopril system. The RMSF analysis demonstrated that MBL-ZINC00517765 in the residues including Arg109 with 1.38 Å, Glu156 with 2.24 Å, Ser159 with 2.41 Å, Ser161 with 2.92 Å, and Glu232 with 1.98 Å had the most fluctuations. Inter-molecular H-bond analysis revealed that during simulation, ZINC00517765 formed more hydrogen bonds with the active site of MBL than did Captopril. The PCA displayed the movements of proteins. The PCA outcomes showed that MBL-ZINC00517765 had less motion than the other two systems and confirmed the results of other analyses, including Rg, intramolecular H-bonds, RMSF, and RMSD.