1. Background
Respiratory tract infections (RTIs) impose a significant burden on health systems worldwide. However, current diagnostic tools may detect causative pathogens in only 30 - 40% of RTI patients. Therefore, the causative agents remain unrecognized in most clinical cases, and nonspecific respiratory symptoms are usually treated empirically. By application of current molecular techniques, it is now established that about 25% of community-acquired Pneumonia (CAP) has viral etiology (1, 2). Moreover, disease severity and hospital stay length may increase by viral or viral-bacterial coinfections in CAP (3), escalating the need for intensive care and increased mortality rate in pneumonia patients (2).
Application of sensitive diagnostic assays could identify more than one pathogen in 4 - 30% of adults and 23 - 33% of children in prospective studies of CAP (4-8). The detection of respiratory and non-respiratory viruses in RTI patients is challenging. They could be a sole cause may be of viral origin or a coinfection; besides, they can facilitate or worsen the bacterial infection. Differentiating a viral origin from bacterial infection or mixed viral-bacterial infection could substantially decrease antibiotic use, avoid the associated risk of adverse reactions, reduce antibiotic resistance development, decrease hospital-acquired infections, and improve clinical outcomes (9).
Respiratory multiplex real-time kits are in vitro tests for the quantitative/qualitative detection of pathogens’ nucleic acids in throat/nasal swabs, bronchoalveolar lavage, sputum, and culture of human origins for the evaluation of infections with respiratory pathogens detected by commercial kits. These tests contain separate primers and probes, each targeting the compatible sequence on the desired genes (target of interest) with high sensitivity and specificity. These assays have been optimized for the simultaneous detection of pathogens in unknown samples. Currently, the development and improvement of multiplex PCR tests allow accurate characterization of a wide range of viral and bacterial pathogens in acute and non-acute samples (10-12). At present, multiplex PCR assays are the gold standard for diagnosing viral RTI (13), making it possible to diagnose viral and bacterial RTIs rapidly and simultaneously. In addition to the quantitative measurement ability of real-time PCR assays, semi-quantitative microbial load data could be identified to differentiate colonization from true infection by pathogens in symptomatic and asymptomatic patients (14).
2. Objectives
The objectives of the present study were (1) to evaluate a multiplex technique as a "syndromic approach for the detection a wide range of bacteria and non-respiratory viruses in different clinical settings of RTIs, (2) to identify the frequency of each respiratory pathogen alone or in combination (co-detection) among patients with RTIs, (3) to evaluate the clinical presentation of patients with regards to the pathogens detected, and (4) to assess the age specific to disease distribution in different viral and bacterial pathogens.
3. Methods
3.1. Study Design
This cross-sectional, double-center investigation was conducted at the Research Center for Clinical Virology (Tehran University of Medical Sciences), collaborating with Laleh general hospital and Aramesh Medical Laboratory, Tehran, Iran, during 2019 - 2020. Patients were referred to Loghman and Laleh Hospitals, related to the Research Center for Clinical Virology Department, with a history of respiratory tract infection associated with the exacerbation of pulmonary symptoms. There was no age limit for the participants. The inclusion criteria included patients with moderate to severe respiratory symptoms such as cough, dyspnea, hemoptysis, and wheezing, already known for respiratory diseases (small airway diseases) superimposed with acute episodes, and patients who were on antibiotic treatment previously for respiratory infections with unsatisfactory clinical outcomes.
All patients had already been tested for the influenza virus using a triplex real-time PCR assay to detect Flu-A, Flu-B, and H1N1 viruses, all with negative results. The primary clinical manifestations were bronchial asthma, bronchiectasis, tracheobronchitis, and bronchiolitis. A standard chest X-ray showed diffuse and focal reticular patterns, consolidation with lower lobe distribution, air trapping, and congested features in all subjects. Acutely ill patients without previous respiratory tract infection history were excluded from the study.
3.2. Sampling
Respiratory specimens were collected from patients, including the sputum, anterior nasal swabs, and throat swabs using stiff synthetic swabs, and transported in the viral transport media to maintain viral nucleic acids in good condition until analysis.
3.3. DNA Extraction and Polymerase Chain Reaction
Using a viral/bacterial RNA/DNA nucleic acid extraction kit (ROCHE, Mannheim, Germany), the viral and bacterial nucleic acids were extracted from different respiratory samples according to the manufacturer’s instructions. For each multiplex panel, internal control was utilized to exclude false-negative results obtained by PCR inhibitors. Multiplex real-time PCR was carried out on 5 µL of eluted DNA using different panels of respiratory bacterial and respiratory and non-respiratory viral pathogen kits, including Flu, Neuro-9, CAP, and HAP kits (Siemens, Luxembourg), according to the manufacturer’s instructions. Semi-quantitative estimation was calculated for each specimen.
3.4. Statistical Analyses
The data were analyzed by SPSS version 23. The normality of age data was twisted with the Kolmogorov-Smirnov test (P = 0.002). The frequency of variables was presented in percentage. The figures were generated with an excel program. The comparison of means was carried out with a nonparametric chi-square test. A P value < 0.05 was considered statistically significant for all comparisons.
4. Results
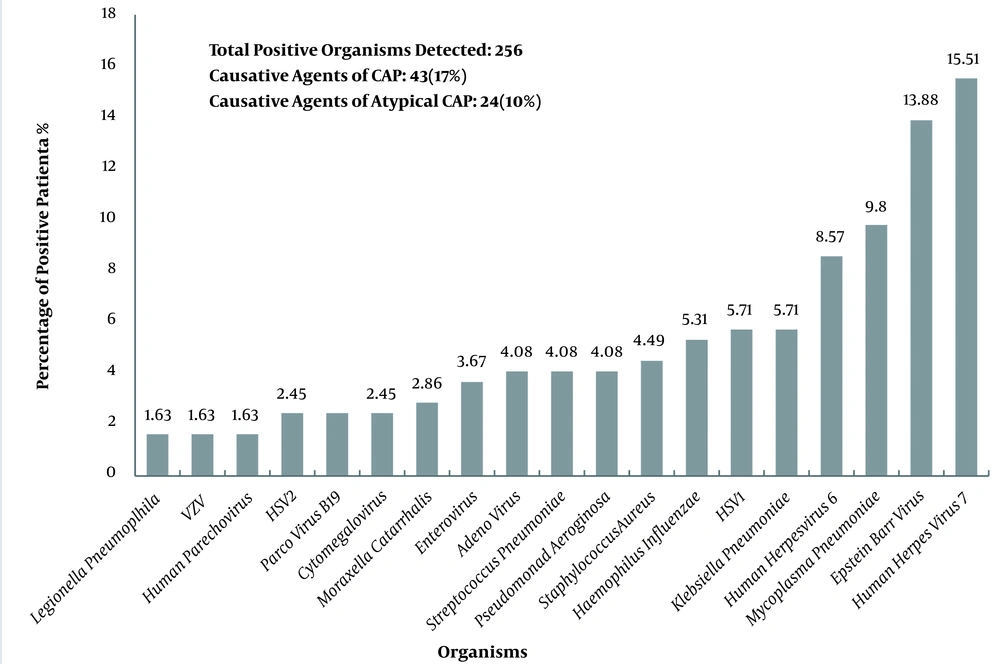
This study enrolled 144 flu-negative outpatients based on the eligibility criteria. The subjects’ age ranged from three to 89 years (mean 42.8, SD 23.7), and 32% of the participants were between 41 and 60 years (Table 1). There were 77 (53.5%) females and 67 (46.5%) males. Thirty-two (22%) patients had a history of comorbidities, including asthma, COPD, and diabetes (data not shown). Thirty-one (21.5%) patients tested negative for all pathogens. Thirty-one (21.5%) patients were infected with only one viral or bacterial agent. Eighty-two (57%) patients were infected with more than one pathogen (viral and/or bacterial). Of a total of 256 multiplex tests, 95 (37%) and 161 (62%) tests were positive for bacterial and viral pathogens, respectively (data not shown). Besides, 23 (16%), 48 (33%), and 61 (42.5%) patients had bacterial, viral, and viral/bacterial co-infections, respectively (Table 1). In 50 (34.7%) cases, we simultaneously detected two organisms (data not shown). The maximum number of organisms detected was six, observed in three cases (data not shown). In addition, CAP and atypical CAP pathogens were found in 17% and 10% of respiratory specimens, respectively (Figure 1).
| Group | All Patients | Any Comorbidity | Negative for All Microbes | 1 Microbe Detected | > 1 Microbe Detected | Bacteria All | Bacteria Median CT | Viruses All | Viruses Median CT | Bacterial Co-detection | Viral Co-detection | Bac/Vir Co-detection |
|---|---|---|---|---|---|---|---|---|---|---|---|---|
| Age (y) | ||||||||||||
| 2 - 7 | 16 (11.1) | 4 (25) | 3 (18.7) | 2 (12.5) | 11 (68.7) | 13 (14.2) | 31.00 | 22 (13.6) | 32.00 | 4 (17.3) | 7 (14.5) | 9 (14.7) |
| 8 - 15 | 11 (7.6) | 4 (36) | 2 (18) | 1 (9) | 8 (72.7) | 6 (6.5) | 31.50 | 15 (9.3) | 32.00 | 1 (4.3) | 4 (8.3) | 6 (9.8) |
| 16 - 40 | 29 (20.1) | 3 (10) | 8 (27.5) | 5 (17.2) | 16 (55) | 15 (16.4) | 33.00 | 34 (21.1) | 32.00 | 4 (17.3) | 9 (18.7) | 11 (18) |
| 41 - 60 | 46 (31.9) | 13 (28) | 10 (21.7) | 11 (23.9) | 25 (54.3) | 25 (27.4) | 32.00 | 51 (31.6) | 31.50 | 6 (26) | 18 (37.5) | 17 (27.8) |
| > 60 | 42 (29.1) | 8 (19) | 8 (19) | 12 (28.5) | 22 (52.3) | 32 (35.1) | 32.00 | 39 (24.2) | 32.00 | 8 (34.7) | 10 (20) | 18 (29.5) |
| Gender | ||||||||||||
| Female | 77 (53) | 18 (23) | 18 (58) | 14 (45) | 45 (54.8) | 38 (54.2) | 32.00 | 53 (53) | 32.00 | 10 (43.4) | 25 (52) | 34 (55.7) |
| Male | 67 (47) | 14 (21) | 13 (42) | 17 (54.8) | 37 (45.1) | 32 (45.7) | 32.00 | 47 (47) | 32.00 | 13 (56.5) | 23 (48) | 27 (44.2) |
| Antibiotic therapy | 23 (15.9) | - | 8 (34) | 5 (21) | 10 (43) | 11 (33.3) | 34 | 22 (66.6) | 31.5 | 4 (17) | 7 (30) | 6 (26) |
| Total | 144 | 32 (22) | 31 (21.5) | 31 (21.5) | 82 (57) | 95 (37) | 32.00 | 161 (62) | 32.00 | 23 (16) | 48 (33) | 61 (42.5) |
The distribution of pathogens was higher in females than males; however, this difference was not significant (P value: 0.56, data not shown). Although patients with previous underlying diseases showed a higher rate of bacterial, viral, and viral/bacterial coinfections, it did not reach statistical significance (data not shown). Regarding the history of antibiotic consumption before the tests, 23 had a history of antibiotic therapy. Fourteen and seven of these patients were positive for viral and bacterial infections, respectively, but the positivity rate was not different between those with and without a history of antibiotic therapy (P value: 0.09). The minimum detected Ct was significantly higher in those with a history of antibiotic therapy than others (31 vs. 27, P value: 0.005, data not shown). No other findings were correlated with a history of antibiotic therapy (Table 1). Subjects > 41 years old had more coinfections and multi-infections than other age groups. On the other hand, children (< 15 years old) showed lower rates of coinfections and multi-infections, especially those in the 8-15-year-old group (Table 1).
Figure 1 presents the frequency distribution of viral and bacterial infections. The four predominant pathogens were Human Herpes Virus 7 (HHV-7) (n = 38, 15.5%), Epstein-Barr Virus (EBV) (n = 34, 13.8%), Mycoplasma pneumoniae (n = 24, 9.8%), and Human Herpes Virus 6 (HHV-6) (n = 21, 8.5%). Bacterial pathogens responsible for CAP were among the commonest pathogens. Accordingly, Pseudomonas aeruginosa and Klebsiella pneumoniae responsible for atypical CAP pathogens composed 10% of the pathogens. Table 2 shows the frequency of viral and bacterial infections against age categories. Klebsiella pneumoniae and P. aeruginosa (both belonging to atypical CAP) were found mainly in those > 41 years old, especially the elderly (P values: 0.158 and 0.09, respectively). Mycoplasma pneumoniae, Haemophilus influenzae, HHV-6, and HHV-7 were seen mostly in patients < 7 and 16 - 40 years old (Table 2). Haemophilus influenzae, HHV-6, and HHV-7 were mostly observed for those between 8 and 15 years. Mycoplasma pneumoniae, HHV-6, HHV-7, EBV, Adenovirus, Parvovirus B19, K. pneumoniae, and Staphylococcus aureus were dominantly detected in the 41 - 60 age group. Lastly, EBV, P. aeruginosa, K. pneumoniae, HHV-7, HSV-1, and S. pneumonia were detected in the > 60 age group (Table 2).
| Pathogens | 2 - 7 Years | 8 - 15 Years | 16 - 40 Years | 41 - 60 Years | > 60 Years |
|---|---|---|---|---|---|
| Klebsiella pneumoniae | 1 (3.4) | 0 (0.0) | 2 (4.1) | 6 (7.5) | 5 (7.2) |
| Pseudomonas aeruginosa | 0 (0.0) | 0 (0.0) | 2 (4.1) | 8 (9.7) | 8 (11.6) |
| Mycoplasma pneumoniae | 6 (20.6) | 1 (5.2) | 7 (14.2) | 8 (9.7) | 2 (2.8) |
| Moraxella catarrhalis | 2 (6.8) | 1 (5.2) | 0 (0.0) | 1 (1.2) | 3 (4.3) |
| Haemophilus influenzae | 0 (0.0) | 3 (16.0) | 4 (8.1) | 2 (2.4) | 4 (5.7) |
| Staphylococcus aureus | 1 (3.4) | 0 (0.0) | 2 (4.1) | 4 (4.7) | 4 (5.7) |
| Chlamydia pneumoniae | 0 (0.0) | 0 (0.0) | 0 (0.0) | 0 (0.0) | 0 (0.0) |
| Streptococcus pneumoniae | 3 (10.6) | 1 (5.2) | 0 (0.0) | 2 (2.4) | 4 (5.7) |
| Legionella pneumophila | 0 (0.0) | 1 (5.2) | 1 (2.0) | 1 (1.2) | 1 (1.4) |
| Enterovirus | 2 (6.8) | 1 (5.2) | 2 (4.1) | 2 (2.4) | 2 (2.8) |
| Human parechovirus | 0 (0.0) | 1 (5.2) | 1 (2.0) | 1 (1.2) | 1 (1.4) |
| Human Herpesvirus 6 | 4 (14.0) | 2 (11.0) | 4 (8.1) | 7 (8.4) | 4 (5.7) |
| Human herpesvirus 7 | 4 (14.0) | 4 (21.0) | 9 (18.3) | 15 (18) | 6 (9.6) |
| Parvovirus B19 | 2 (6.8) | 0 (0.0) | 0 (0.0) | 4 (4.7) | 0 (0.0) |
| Adenovirus | 0 (0.0) | 1 (5.2) | 1 (2.0) | 6 (7.5) | 2 (2.8) |
| Cytomegalovirus | 0 (0.0) | 1 (5.2) | 2 (4.1) | 2 (2.4) | 1 (1.4) |
| Epstein-Barr virus | 2 (6.8) | 1 (5.2) | 4 (8.1) | 9 (10.9) | 18 (26.2) |
| Herpes simplex virus 1 | 1 (3.4) | 1 (5.2) | 5 (10.6) | 3 (3.3) | 4 (5.7) |
| Herpes simplex virus 2 | 1 (3.4) | 0 (0.0) | 3 (6.1) | 2 (2.4) | 0 (0.0) |
| Total | 29 (100) | 19 (100) | 49 (100) | 83 (100) | 69 (100) |
Prevalence of Pathogens in Different Age Groups a
Next, we compared the association between clinical symptoms and microbial findings. We found that acute cough, chronic cough (more than 35 days), and dyspnea were the common main symptoms between patients. Except for dyspnea that was more common in adults > 60 years old, the other symptoms were proportionally distributed between different age groups and genders (data not shown). Accordingly, the frequency of wheezing was higher in patients infected with EBV and M. pneumoniae (27.78% and 16.67%, respectively), although both were statistically insignificant (Table 3). Hemoptysis was more observed in patients with M. pneumoniae (the highest frequency), followed by adenovirus, HSV-1, and Moraxella catarrhalis (with equal frequencies) (Table 3). Moreover, fever was predominant in patients infected with EBV, HHV-1, HHV-7, and M. pneumonia (Table 3). Furthermore, dyspnea was more associated with detecting EBV, HHV-7, S. aureus, and HHV-6 (listed in the order of significance from the highest to lowest) (Table 3).
| Pathogens | Clinical Symptoms | ||||||
|---|---|---|---|---|---|---|---|
| Fever (N = 17) | Wheezing (N = 18) | Hemoptysis (N = 16) | Dyspnea (N = 83) | Chronic Cough (N = 37) | Acute Cough (N = 148) | Chronic and Acute Cough (N = 14) | |
| Legionella pneumophila | 0 | 1 (5.56) | 0 | 1 (1.15) | 1 (2.44) | 2 (1.25) | 0 |
| VZV | 1 (5.26) | 1 (5.56) | 0 | 0 | 0 | 3 (1.88) | 0 |
| Human parechovirus | 1 (5.26) | 0 | 0 | 0 | 0 | 4 (2.5) | 0 |
| Herpes simplex virus 2 | 0 | 1 (5.56) | 1 (6.25) | 1 (1.15) | 1 (2.44) | 5 (3.12) | 1 (6.25) |
| Parvovirus B19 | 0 | 0 | 1 (6.25) | 2 (2.3) | 0 | 5 (3.12) | 0 |
| Cytomegalovirus | 0 | 1 (5.56) | 0 | 3 (3.45) | 4.88 | 1 (0.52) | 0 |
| Moraxella catarrhalis | 0 | 0 | 2 (12.5) | 3 (3.45) | 0 | 4 (2.5) | 0 |
| Enterovirus | 0 | 1 (5.56) | 0 | 0 | 0 | 6 (3.75) | 0 |
| Adenovirus | 0 | 1 (5.56) | 2 (12.5) | 4 (4.6) | 4 (9.76) | 3 (1.88) | 0 |
| Streptococcus pneumoniae | 2 (10.53) | 0 | 1 (6.25) | 4 (4.6) | 1 (2.44) | 6 (3.75) | 0 |
| Pseudomonas aeruginosa | 0 | 1 (5.56) | 1 (6.25) | 7 (8.05) | 0 | 5 (3.12) | 0 |
| Staphylococcus aureus | 0 | 1 (5.56) | 0 | 8 (9.2) | 1 (2.44) | 7 (4.38) | 0 |
| Haemophilus influenzae | 0 | 0 | 1 (6.25) | 4 (4.6) | 1 (2.44) | 8 (5) | 0 |
| Herpes simplex virus 1 | 3 (15.79) | 0 | 2 (12.5) | 5 (5.75) | 3 (7.32) | 6 (3.75) | 0 |
| Klebsiella pneumoniae | 0 | 1 (5.56) | 0 | 4 (4.6) | 1 (2.44) | 12 (7.5) | 1 (6.25) |
| Human herpesvirus 6 | 0 | 0 | 0 | 6 (6.9) | 4 (9.76) | 13 (8.12) | 1 (6.25) |
| Mycoplasma pneumoniae | 3 (15.79) | 3 (16.67) | 3 (18.7) | 5 (5.75) | 7 (17.07) | 19 (11.88) | 5 (31.25) |
| Epstein-Barr virus | 4 (21.05) | 5 (27.78) | 1 (6.25) | 13 (14.94) | 6 (14.63) | 23 (14.38) | 3 (18.75) |
| Human herpes virus 7 | 3 (15.79) | 1 (5.56) | 1 (6.25) | 13 (14.94) | 7 (17.07) | 21 (13.12) | 3 (18.75) |
Frequency of Clinical Symptoms Concerning Isolated Pathogens a
We divided patients into acute, chronic, and acute-chronic cough patients regarding the history of cough. Acute cough was accompanied mainly by M. pneumoniae and HHV-7 with an equal frequency, followed by EBV, HHV-6, and adenovirus. However, EBV was the dominant pathogen for chronic cough, followed by M. pneumoniae, HHV-7, HHV-6, and K. pneumoniae (Table 3). Lastly, three pathogens were mainly associated with cough, regardless of being acute or chronic, including M. pneumoniae, EBV (18.75%), and HHV-7 (18.75%) (Table 3). Dyspnea and cough were associated with multiple pathogen detection (viral, bacterial, and viral-bacterial); however, these correlations were insignificant compared to other symptoms (data not shown). Regarding the Ct values of the tests, there was no substantial difference between viral and bacterial Ct median values concerning gender, age, and comorbidities (Table 1). However, the minimum detected Ct was significantly higher in those with a history of antibiotic therapy than others (31 vs. 27, P value: 0.005)
5. Discussion
The present study was the first Iranian survey of the frequency of pathogens among RTI outpatients referred to Loghman and Laleh hospitals using multiplex assays during 2019 - 2020. The results clearly showed that using targeted panels of real-time multiplex PCR for viral and bacterial pathogens increased the chance of a proper diagnosis in patients with respiratory tract infection and provided valuable information about the distribution of pathogens among different age groups. Of all tests, 36% and 64% were positive for bacterial and viral pathogens, respectively. Although the differences were not statistically significant, we observed higher positivity rates in females, those without a history of antibiotic therapy, and those with underlying disease. Previously, the viral etiology of CAP and the nature of mixed infections with different pathogens had been underestimated due to the limited range of diagnostic methods. However, recent studies on multi-targeted molecular diagnostics have indicated that the viral-bacterial co-detection rate was 59.8% (15). Crotty et al. reported bacterial coinfection and viral coinfection rates of 57.6% and 33%, respectively (16). Another study reported bacterial, viral, and viral-bacterial coinfection rates of 28%, 74%, and 17%, respectively (17).
On the other hand, the above data included the respiratory viral causes of CAP. The present study showed that multiple non-respiratory viruses were detected in at least 62% of moderate cases of CAP. Among non-respiratory viruses, HHV-7, EBV, and HHV-6 were the most frequent ones, comprising 14.9%, 13.3%, and 8.2% of all detected pathogens, respectively. There are several reports on the increased detection of non-respiratory viruses involved in viral CAP in respiratory samples from immunocompetent patients, although at a lower frequency than in immunocompromised hosts. Besides, EBV has been the most commonly detected one, followed by HHV-7, HHV-6, and CMV at different rates among patients with various respiratory diseases (18-20).
In two different reports on ARDS patients with unknown etiology, Bonizzoli reported a prevalence of 43% for EBV and 28% for HSV-1 and CMV (21), and in the second survey, Tachikawa et al. found CMV (60%), EBV (45%), HSV-1 (31%), HHV-7 (2%), and HHV-6 (5%) (22). Among 108 bronchiectasis patients, 48% were positive for EBV (23). Of 136 patients with chronic obstructive pulmonary disease (COPD), EBV was observed in 65 (48%) severe patients and 31 (46%) moderate patients (24). In patients with interstitial pneumonia, HHV-7 DNA was detected in 79.2%, and HHV-6 DNA was discovered in 12.5% of biopsy tissues (25). On the other hand, EBV and HHV-7 were detected frequently in asymptomatic immunocompetent healthy adults as 9 - 50% (4, 5, 26, 27) and 34% (8, 28), respectively.
A question is raised “What are the roles of viral agents in respiratory secretions either alone or combined with bacterial agents?” Over the past few years, several herpesviruses have been isolated in patients with respiratory tract infections. As known, EBV, HHV-6, and HHV-7 can cause severe pneumonia in immunocompromised hosts and pneumonia in normal individuals without underlying diseases (29, 30). The interaction of these co-pathogens may worsen respiratory disease outcomes. Viral infections may predispose the respiratory tract for secondary bacterial infections by an interaction between bacteria and respiratory viruses (31, 32). However, there is still scarce data on the occurrence of non-respiratory viruses in immunocompetent CAP patients; therefore, further observational data using multiplex assays are needed to fulfill this gap of knowledge.
Patients > 41 years had more coinfections and multi-infections than other age groups. On the other hand, children < 15 years showed lower rates of coinfection and multi-infection, especially those in the 8 - 15 years old group. Despite a high rate of herpesviruses primary infection in immunocompetent children, detecting these viruses in respiratory secretions might not be a common phenomenon. The other reason could be that children < 15 years old comprised less than 20% of the patient populations in the present study. We found some associations between four significant symptoms and pathogen predominance. Wheezing and cough (both acute and chronic) were primarily observed in the presence of M. pneumoniae and EBV. Hemoptysis was mainly associated with M. pneumoniae, adenovirus, and HSV-1, indicating the potential of a correlation between major symptoms and specific pathogens. On the other hand, fever, acute/chronic cough, and dyspnea were strongly associated with versatile lists of pathogens. Likewise, in some surveys, wheezing in children and adults had been meaningfully linked with viral pneumonia compared to mixed viral or bacterial infections (33, 34). However, we appreciate that clinical manifestations may differ depending on specific coinfection patterns, and significant overlaps in the present study and other reports mitigate the utility of these findings in respiratory-infected patients.
Herpesviruses could persist and establish latency; therefore, differentiation between latent infection and active viral replication by diagnostic assays is paramount. Positive results on qualitative assays may indicate respiratory disease etiology, asymptomatic colonization, microorganism shedding, or upcoming infection (35). Using a semi-quantitative approach, we aimed to use the Ct values; however, due to significant overlap in Ct value diversities between different detected pathogens, identifying a certain Ct value threshold to show the cause and effect relatedness was not possible. The present investigation also has several limitations. The obtained results focused only on pathogens included in the employed respiratory assays; therefore, the presence of other non-detected pathogens could not be excluded. Moreover, the obtained data from a double-center study could not be attributed to the whole country.
5.1. Conclusions
In conclusion, given that a substantial proportion of respiratory diseases has been caused by viral pathogens, the unnecessary usage of antibiotics is puzzling due to its unfavorable health consequences and antimicrobial resistance complications. Hence, the multiplex respiratory panels provide the early detection of the pathogen spectrum and benefit patients in the clinical decision-making process. The role of herpesviruses in disease worsening and complications deserved further investigations.