1. Background
Brain cancer is one of the deadliest types of cancer, with a poor prognosis (1). Glioblastoma, also known as GBM, is the most aggressively spreading glioma of astrocytic lineage and corresponds to grade IV based on WHO classification (2). The standard treatment for GBM patients involves surgery, followed by radiotherapy with the simultaneous administration of temozolomide and the use of adjuvant drugs (3). The aggressiveness and high degree of penetration of this disease often result in surgical failure (4).
Epigenetic dysregulation is a common feature of many human diseases, including cancer. Cancer research has focused on identifying epigenetic alterations associated with malignant tumors, and the epigenetic regulation of cancer cells has gradually become more apparent. Epigenetic changes are associated with most phases of tumor development, including carcinogenesis, promotion, progression, and recurrence. Epigenetic drugs may reverse some of these changes (5). DNA methylation, which involves the addition of a methyl group to the promoter region of a gene, is one of the primary causes of cancer formation. This insertion either silences the gene or raises the probability of mutations. Oncogenesis is mainly caused by both hypermethylation and hypomethylation; the former occurs more often near the CpG islands in the gene's promoter region, while the latter occurs globally in various genomic locations (6).
The DNMT3A gene encodes the enzyme that adds methyl groups to cytosine residues in CpG dinucleotides. These dinucleotides often build up in the 5' promoter regions upstream of the genes, and a reduction in the expression of the corresponding downstream gene is linked to an increase in methylation of these CpG islands (7). Tumor cells usually show aberrant DNA methylation patterns during transformation into a malignant mass. Although this phenomenon is generally attributed to various mechanisms, changes in the DNMT gene family and disruption of genomic methylation regulation are the main factors causing it. Several samples with multiple abnormalities in DNMT genes have been studied to identify methylation changes and evaluate cancer development (8). High expression of DNMTs (DNMT1, DNMT3A, and DNMT3B) in all tumors leads to hypermethylation and activation of oncogenes (9).
SUZ12 is a nuclear protein of the PRC2 suppressor complex, including EED, EZH2, RBBP4, and RBBP7 proteins. Biochemical studies have shown that all PRC2 components are required for chromatin compaction. This is probably due to the dependence of EZH2 histone methyltransferase activity on EED and SUZ12 and the role of RbBP4 and RbBP7 in mediating binding to the nucleosome (10). PRC2 plays a role in various biological processes, including differentiation, cell identity, proliferation maintenance, and stem cell plasticity. Recent studies have highlighted PRC2 as an essential functional and regulatory factor and the role of non-coding RNA in employing PRC2 to target genes of interest (11).
A type of transcript classified as long non-coding RNAs (lncRNAs) has a length of more than 200 nucleotides and a limited or no coding ability. The sequencing of full-length cDNA libraries in the human genome has lately shown the great diversity of lncRNAs despite their identification decades ago. According to the GENCODE project, about 16,000 lncRNA genes exist in the human genome (12). Long non-coding RNAs plays an essential role in various regulatory processes, such as regulating gene expression (12). HOTAIR is a lncRNA that plays a vital role as an oncogene in multiple cancers such as breast, gastric, colorectal, and cervical cancer (13). HOTAIR expression level can be an essential biomarker for diagnosing and treating various cancers. This RNA also participates in the epigenetic regulation of genes and plays a vital role in different cellular pathways (13).
Despite various treatments for cancer, especially brain cancer, the focus on treatments with fewer side effects has increased in recent years. Curcumin is one of the known compounds in cancer treatment. curcumin, known by the scientific name diferuloylmethane, is a yellow polyphenol and the active component of the perennial plant Curcuma longa (14). This compound is well known for its anti-cancer activities, and many preclinical studies have confirmed its anti-cancer activity (15). Various studies have also shown that curcumin is an epigenetic modifier. Drugs, nutritional, and environmental factors can induce epigenetic changes. Curcumin can affect the activity of various epigenetic factors, including HATs, DNMTs, and miRNAs, and modulate epigenetic processes. However, some findings have not reported the occurrence of demethylation changes by curcumin in tumor cells (15, 16).
Despite the evidence available for the efficacy of curcumin in GBM, clinical trial reports are scarce, and the molecular mechanisms of curcumin require further studies (17). Additionally, despite many therapeutic benefits, curcumin suffers from drawbacks such as low solubility and partial cellular uptake, limiting its use as a therapeutic agent for clinical purposes. There are various methods for preparing nanoparticles (NPs); however, in a study, the wet milling technique was used to reduce the size of curcumin particles to 2 to 40 nm. As a result, the curcumin nanoparticles (curcumin-NPs) prepared using this method exhibited acceptable chemical and physical stability as well as high solubility in water (18). Research has shown that curcumin becomes more water-soluble, which could increase its utility in pharmaceutical formulations and water-based therapeutic options. The average size of less than 60 nm in the measurement of these nanoparticles indicated that these curcumin-NPs are in the ideal size range for effective cellular uptake. Nanoparticles with a size of less than 100 nm may, in most cases, cross biological barriers such as the blood-brain barrier and cell membranes and may also be effectively absorbed by target organs.
2. Objectives
In this study, we used curcumin-NPs synthesized by the ultrasound technique as potential inhibitors of the GBM cancer cell line. The study aimed to analyze the effect of these particles on the expression of important epigenetic regulators in the GBM cell line. To our knowledge, this is the first study showing that these NPs can inhibit the proliferation and invasion of the GBM cell line by modulating the master epigenetic regulators and inducing demethylation of the genome.
3. Methods
3.1. Reagents and Materials
Curcumin (CAS 458-37-7) was purchased from Samchun (Korea). A transmission electron microscope (Zeiss-EM10C-80 KV, Germany) was used to investigate the morphology and size of curcumin-NPs. An ultrasonic device (DSA 100-SK2, China) operating at 100 watts and 40 kHz was used to prepare curcumin-NPs. Fetal bovine serum, penicillin-streptomycin, trypsin-EDTA, and DMEM medium were obtained from Bio-idea (Iran). Dichloromethane, Triton X-100, and DMSO were provided by Merck (USA). Cell culture plates were sourced from SPL Life Sciences (South Korea). The cell line 1321N1 was obtained from the National Cell Bank of Iran (Pasteur Institute of Iran, Tehran). Agarose gel powder and a cDNA synthesis kit were prepared by Sina Clon (Iran). Master Mix PCR and Green RT-qPCR were provided by Ampliqon (Denmark). DNMT3A, SUZ12, and MecP2 primary antibodies (DNMT3a (C-12): sc-365769, SUZ12 (D-10): sc-271325, β-Actin (C4): sc-47778, and MecP2 (G-6): sc-137070) and secondary antibodies (m-IgGκ BP-HRP: sc-516102 and mouse anti-rabbit IgG-HRP: sc-2357) were purchased from Santa Cruz Biotechnology (USA).
3.2. Preparing Curcumin Nanoparticles
Curcumin-NPs were prepared using ultrasonic methods (19, 20). In summary, 125 mg of Curcumin was weighed and dissolved in 25 mL of dichloromethane (organic phase). In the next step, 2 mL of the organic phase was added to the aqueous phase under ultrasonic conditions. The color of the solution turned yellow once this step was completed. The prepared solution was placed in an ultrasound machine for 20 minutes to ensure the complete dispersion of Curcumin. Then, the content was placed on a magnetic heater for 20 minutes to ensure thorough dissolution of the solution. Finally, the solution was subjected to reduced pressure in a rotary device to remove dichloromethane. The concentration of the final solution was 0.271 mM. The prepared solution was poured into a dark-colored bottle and kept in the refrigerator (20).
3.3. Examining the Structure of the Curcumin Nanoparticles
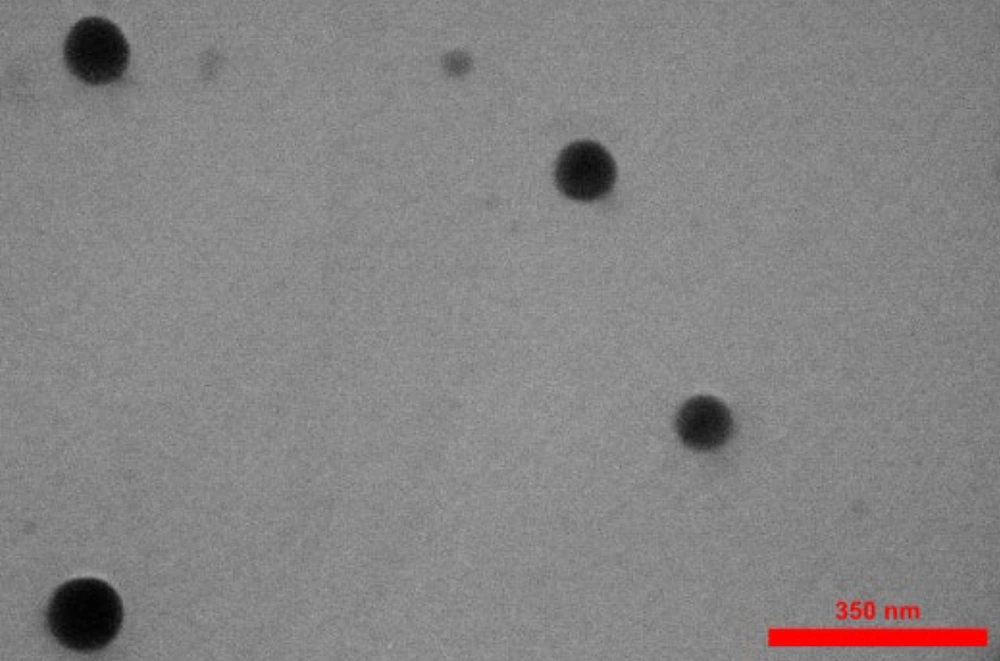
In brief, 1 mL of prepared curcumin-NPs was dissolved in double-distilled water to reach a total volume of 10 mL. This solution was then used to measure the size of the particles and capture images. The final image obtained from the electron microscope was evaluated using ImageJ software (Maryland, USA).
3.4. Examining the Absorption Spectrum of Curcumin Nanoparticles
Ultraviolet-visible spectrophotometry (Cintra 101, Sydney, Australia) was used to measure the absorption spectrum. In this experiment, 500 µL of curcumin-NPs was added to 4.5 mL of double-distilled water. The absorption spectrum of curcumin-NPs was then measured at wavelengths ranging from 350 to 500 nm. The results were analyzed using Excel software (Microsoft Excel 2022).
3.5. Cell Culture and Viability
The GBM cell line 1321N1 was used for toxicity and cell survival assays. The prepared cells were cultured in DMEM medium containing 10% fetal bovine serum and 1% penicillin and streptomycin (all from Bio-idea). The cultured cells in the flask were then placed in an incubator under 5% CO2 and 98% humidity at 37°C for 48 hours. After reaching approximately 80% confluence, the cells were passaged using 0.25% trypsin-EDTA (x1).
3.6. Examining the Curcumin NPs Cytotoxicity
Briefly, 5,000 cells were cultured in each well of a 96-well plate. After 24 hours, the cells were treated with curcumin-NPs at different concentrations (0.5, 1, 1.5, 2, 2.5, 3, and 6 µM) compared to the control group. After 48 hours, the culture medium containing curcumin-NPs was replaced with 30 µl of MTT salt (Atocel, UK), previously prepared in a dark and sterile environment. To prepare MTT salt, 5 mg of MTT salt was weighed and dissolved in 10 ml of PBS buffer. The plate containing MTT was placed in an incubator for 4 hours, after which 120 µl of DMSO (Merck) was added to each well and placed in the ELISA reader (Bio-Rad). The reading was performed at a wavelength of 570 nm. The data obtained from ELISA were analyzed to determine the IC50 and the percentage of cell survival using GraphPad Prism software (Boston, Massachusetts, USA). The cell viability percentage was also analyzed using the same software with the one-way ANOVA method. The experiment was replicated three times.
3.7. Scratch Wound Healing Assay
In this test, 120,000 cells were cultured in each well of a six-well plate. After 24 hours, the culture medium was drained from each well, and the wells were washed with PBS. Next, a sterile 100-µl pipette tip was used to make a scratch inside each well (treated and control). The plate was then placed in an incubator and treated for 48 hours. After this period, the wells were photographed, and the results were analyzed using ImageJ software.
3.8. Quantitative RT-PCR Assay
The cells were lysed with RNX (Sina Clon) for RNA extraction. Gel electrophoresis and a nanodrop spectrophotometer were used to assess the quality and quantity of the extracted RNA, respectively. Primer design was carried out using Oligo7 software (Molecular Biology Insights, USA), and their specificity was evaluated using NCBI's Primer-BLAST (21) (Table 1). cDNA synthesis was performed using RT-PCR with a cDNA synthesis kit (Sina Clon). The efficiency of the primers was determined using the serial dilution method. In the final step, gene expression was investigated using an RT-qPCR device (ROCHE, Germany). The data obtained from RT-qPCR were analyzed using the Livak method (22).
| Genes and Primers | Sequences (5'→3') | Product Length (bp) | Ta (°C) | Accession Number |
|---|---|---|---|---|
| ACTB | 103 | NM_001101 | ||
| Forward | GAGCATCCCCCAAAGTTCACA | 61 | ||
| Reverse | GGGACTTCCTGTAACAACGCA | 61 | ||
| DNMT3A | 144 | NM_022552 | ||
| Forward | CGACCAGGAATTTGACCCTCC | 63 | ||
| Reverse | ATGTAGCGGTCCACCTGAATG | 61 | ||
| SUZ12 | 132 | NM_015355 | ||
| Forward | ATGCAGGAAATGGAAGAATGTCC | 61 | ||
| Reverse | TGTCCAACGAAGAGTGAACGTC | 62 | ||
| HOTAIR | 109 | NR_186240 | ||
| Forward | TTGTGTAGACCCAGCCCAAT | 58 | ||
| Reverse | AATTCCGGAGCAGCTCAAGT | 59 |
3.9. Western Blotting Assay
After the treatment, the supernatant containing the extracted protein was stored at -20°C. Protein concentration was determined using the Bradford method. Different concentrations of BSA (1, 0.6, 0.5, 0.3, 0.25, 0.15, 0.125 mg/mL) were prepared to draw a standard curve. BSA was used as a standard protein to measure the amount of protein. The exact protein concentration was used for processing with 10% SDS-PAGE. The protein bands were then transferred onto a PVDF membrane (Millipore, Billerica, MA, USA). After preparing a blocking solution, the PVDF membrane was placed inside and incubated. Chemiluminescence kits (ECL advanced reagents) were used for the detection of protein bands. The protein bands were subsequently analyzed using IMAGEJ software.
3.10. Statistical Analysis
All calculations were performed using GraphPad Prism v9.5.1 (GraphPad Software, San Diego, USA). Statistical significance between groups in both real-time PCR and Western blot tests was determined using t-test analysis. Additionally, statistical significance between survival percentages in GBM cell line groups was determined using one-way ANOVA. A P-value of less than 0.05 (*) was considered the criterion for defining statistical significance. The values are expressed as the mean ± SD, and n represents the number of independent experiments, with each independent sample analyzed in duplicate.
4. Results
4.1. Morphological Evaluation of the Curcumin Nanoparticles
The results obtained from this experiment indicate that the curcumin-NPs have a spherical shape and a homogeneous particle size distribution, with an average size of less than 100 nm. These features are particularly important in biological applications, as the physical properties of nanoparticles can significantly affect their behavior and efficacy. The average size of less than 60 nm suggests that these curcumin-NPs are in the ideal size range for effective cellular uptake (Figure 1).
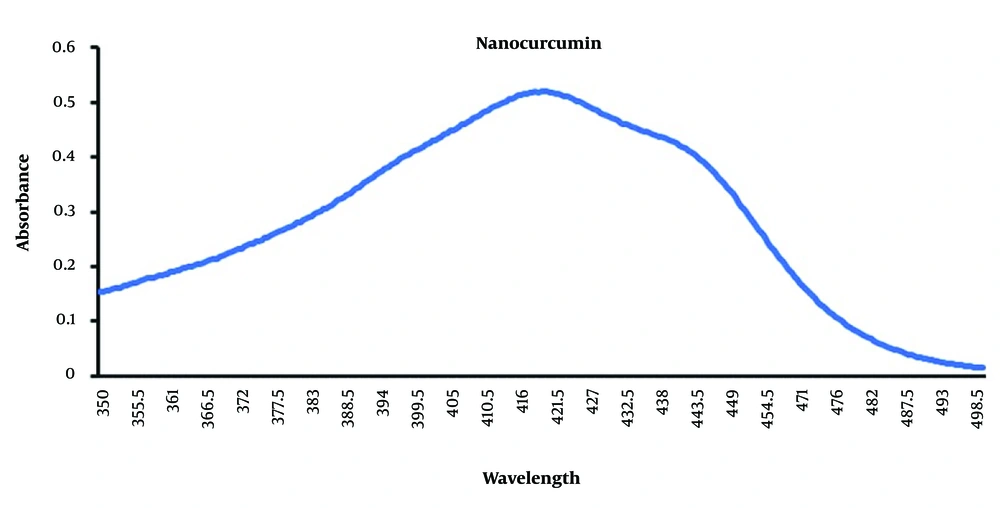
4.2. Absorbance Spectroscopy Analysis of Curcumin Nanoparticles
The results of the absorption spectrum measured with an ultraviolet-visible spectrometer were analyzed using Excel software. The highest absorption of NPs was observed at a wavelength of 426 nm, with a value of 0.51 (Figure 2). This maximum absorption value indicates a strong interaction between the NPs and the incoming light. As shown by the absorption peak, the unique optical properties of NPs are influenced by their size, composition, and surface properties. The observed absorption value of 0.51 also reflects the extent to which the NPs can absorb light at this specific wavelength.
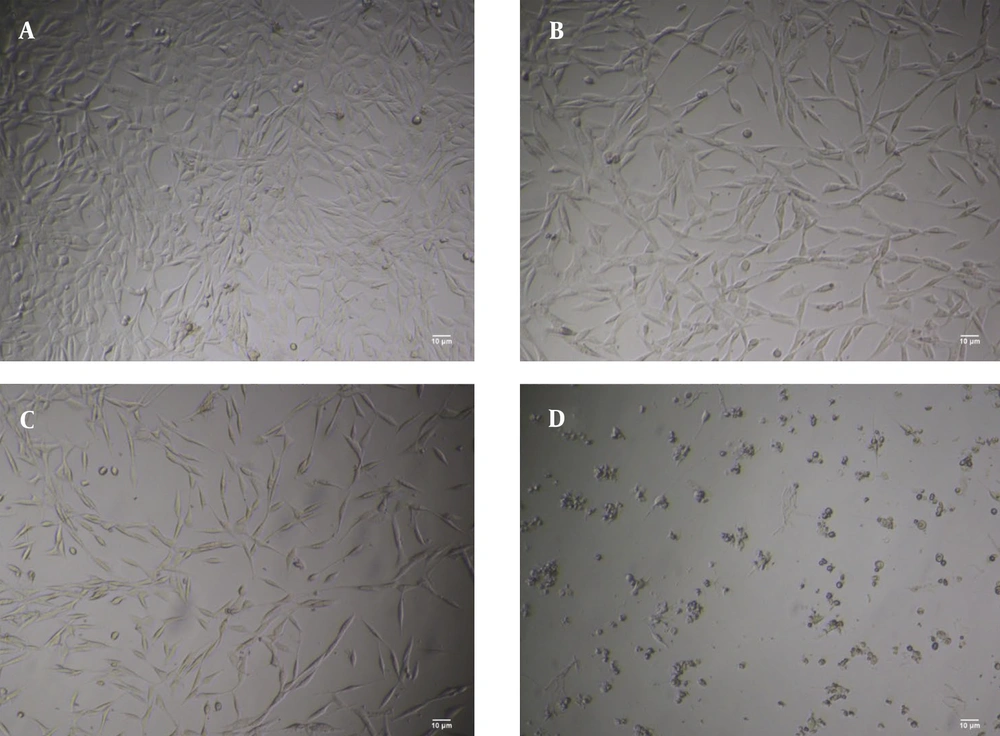
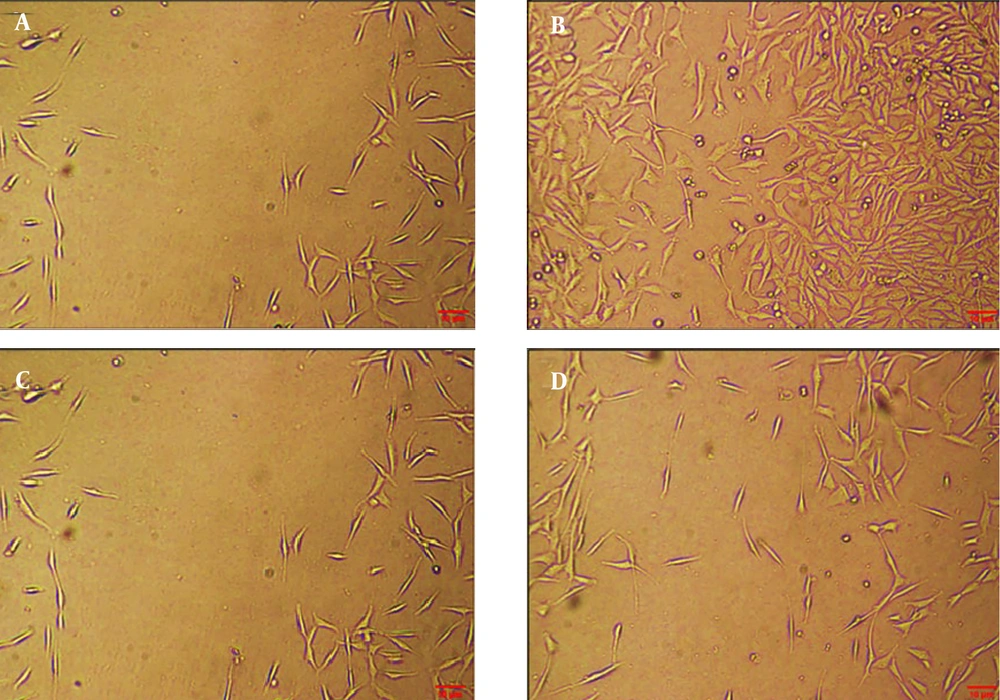
4.3. Investigation of the Morphology of Glioblastoma
The results of different concentrations of curcumin-NPs on GBM cell lines after 48 hours showed that increasing the concentration of curcumin-NPs led to changes in the shape and size of the cells, resulting in a loss of their morphology. The control group was compared with groups treated with curcumin-NPs at concentrations of 0.8, 1.7, and 3.4 µM. The results indicated a decrease in cell viability with increasing drug concentration. The percentage of living cells in the control group was 100%, while in the groups treated with 0.8, 1.7, and 3.4 μM of curcumin-NPs, the viability was approximately 80%, 50%, and 20%, respectively (Figure 3).
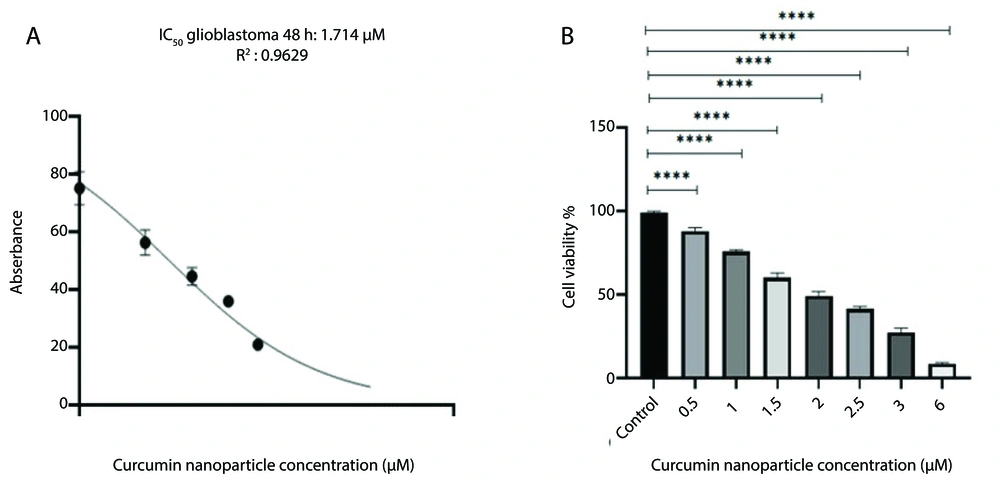
4.4. Investigation of Cell Viability
In this study, we investigated the cell viability of the GBM cell line using an MTT assay. The viability of GBM cells was examined after 48 hours at different doses of 0.5, 1, 1.5, 2, 2.5, 3, and 6 μM. The results showed that with increasing concentration of curcumin-NPs, the cell viability was decreased (P < 0.0001). Curcumin-NPs interfere with cell growth in a dose-dependent manner, so cell growth and survival decrease with increasing concentration of curcumin-NPs. The results indicate that cell death rates increase at high concentrations of curcumin-NPs. However, the lack of toxic effects of curcumin-NPs on the GBM cell line suggests that using these nanoparticles to deliver curcumin to GBM cells is safe (Figure 4).
4.5. Effect of Curcumin NPs on the Migration
In this study, we used a scratch wound healing model to evaluate the effect of curcumin-NPs on GBM cell migration. The findings showed a reduction in cell migration and invasion in the group treated with the IC50 concentration of curcumin NPs (1.7 μM) compared to the control group. These results indicate that curcumin NPs have a significant effect on cell migration and can substantially reduce cancer cell migration and invasion (Figure 5).
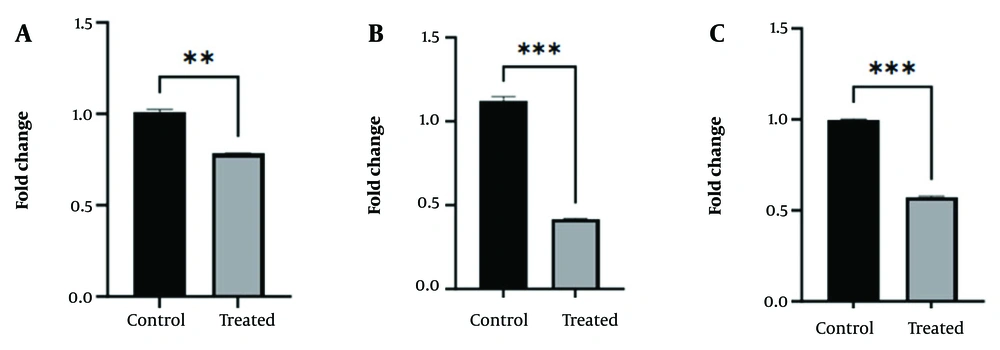
4.6. Downregulation of the RNA Expression Levels of HOTAIR, SUZ12, and DNMT3A
HOTAIR, SUZ12, and DNMT3A expression levels were analyzed using RT-qPCR. The effects of curcumin-NPs on the expression of DNMT3A, HOTAIR, and SUZ12, as important epigenetic markers, were investigated in GBM (Figure 6). The gene expression results in the group treated with curcumin-NPs showed a significant decrease in the expression of the DNMT3A gene in the group treated with the IC50 concentration (1.7 μM) of curcumin-NPs compared to the control group (P < 0.01). Additionally, the gene expression results showed a significant decrease in the expression of SUZ12 and HOTAIR genes in the group treated with the IC50 concentration of curcumin-NPs compared to the control group (P < 0.001).
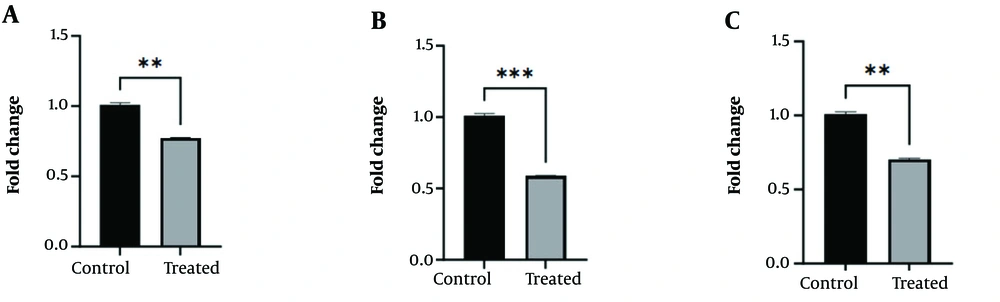
4.7. Downregulation of SUZ12, DNMT3A, and MecP2 Protein Levels
The Western blot method was employed to compare the protein expression of DNMT3A, SUZ12, and MecP2 in the GBM cell line after treatment with curcumin-NPs for 48 hours, compared to the control group (Figure 7). The results of the Western blot analysis showed a decrease in DNMT3A protein levels in the group treated with 1.7 μM curcumin-NPs compared to the control group (P < 0.01). Additionally, the expression of SUZ12 and MecP2 proteins at the IC50 concentration of curcumin-NPs showed decreased expression in the treated group compared to the control group (P < 0.01, P < 0.001).
5. Discussion
Curcumin is a natural polyphenolic compound extracted from the Curcuma longa plant, also known as turmeric (23). This compound has been traditionally used for many years to treat respiratory and infectious allergies. Many studies have shown anti-inflammatory, antimicrobial, anti-tumor, and antioxidant effects for curcumin (24). It has also been shown that curcumin can be used as an adjuvant to secondary drugs, especially in GBM cancer. Additionally, curcumin can affect the proliferation of cancerous cells (25). Nanotechnology-based therapeutic delivery systems, including NPs, nano-emulsions, and liposomes, have emerged to improve the bioavailability, low aqueous solubility, cellular uptake, and antitumor activity of curcumin (26).
This study prepared curcumin-NPs using ultrasonic methods. The results of curcumin-NPs are consistent with previous data on curcumin NPs (19). One of the essential features of curcumin-NPs used in this research is their increased solubility in water, appropriate physical and chemical stability, and suitable particle size (18). Accordingly, the current study shows the effects of curcumin-NPs on the growth and proliferation of GBM cancer cells, and the results demonstrate that curcumin-NPs made by this method have anti-proliferative and inhibitory effects. In general, curcumin-NPs are potentially effective anti-cancer compounds.
Natural compounds like curcumin are known to induce epigenetic changes that may increase the sensitivity of cancer cells to conventional chemotherapeutic agents, thus suppressing tumor growth. The biological effects of curcumin at slightly low concentrations in humans may be attributed to its ability to modulate different pathways through epigenetic mechanisms (27). In an earlier study, DNMT1 gene expression decreased under the influence of curcumin at the mRNA and protein levels. The study suggested that the decrease in gene expression was probably due to the binding of curcumin to the promoter of the desired gene, which may lead to the inhibition of positive regulators such as p65 and Sp1 subunits of NF-κB (16).
Studies investigating the effect of curcumin on DNA methylation have shown that this compound can inhibit the activity of DNMTs and significantly change the pattern of DNA methylation in different tumor cells (16, 28). However, the exact functional mechanism of curcumin and curcumin NPs on the expression of genes involved in epigenetic processes, including DNMT genes, is unknown (29). This research examined the expression of the DNMT3A gene in GBM cancer cells at both mRNA and protein levels under the effect of nanoparticle curcumin. The results of this study show a significant decrease in the expression of the DNMT3A gene at both mRNA and protein levels. This result may provide helpful information about the effect of curcumin on the regulation of epigenetic mechanisms such as DNA methylation. Therefore, curcumin could act as an indirect inhibitor of DNMT3A, helping to alter DNA methylation patterns and prevent the activation of aggressive genes or promote the activation of tumor suppressor genes. Reducing DNMT3A activity may help reduce methylation and thus inhibit tumor growth. However, more research is required to examine the molecular pathways involved in this process.
These epigenetic processes include DNA methylation and demethylation, carried out by methylase and demethylase enzymes, which can help regulate genes related to essential cellular functions such as apoptosis, cancer cell migration, and invasion. This study also revealed that curcumin can down-regulate the migration and invasion of GBM cells and increase the apoptosis rate. Additionally, the results showed that curcumin NPs can affect gene methylation by altering the level of DNMT3A genes and reducing the methylase enzyme, as well as changing the methylation status of genes related to cancer by decreasing the methylase enzyme. This result also shows that curcumin NPs significantly impact the modulation of epigenetic processes.
SUZ12 is a critical component of the PRC2, which drives triple methylation of lysine 27 on histone 3 (H3K27Me3) and gene silencing (29). According to studies, SUZ12 has attracted much attention as a possible oncogene (30, 31). Knockdown in cancer cells with high expression of PRC2 shows that many cancers depend on PRC2 for proliferation (32). The main components of PRC2 are also abnormally expressed in all types of cancers. High levels of SUZ12 are also found in cholangiocarcinoma and ovarian cancer. Hence, SUZ12 expression is essential for the growth and survival of cancers (33). Despite the studies carried out, more research is needed on the role of SUZ12 in cancer, especially GBM. Studies conducted on the role of SUZ12 in GBM cancer indicate the role of this gene in GBM cancer and can be considered a potential factor (34). The more important point is that there are very few direct studies regarding the effect of Curcumin on the expression of the SUZ12 gene, and the effects of Curcumin have been measured directly on PRC2.
In this study, the effect of curcumin NP on the expression of the SUZ12 gene, one of the subunits of the PRC2 complex, was investigated at the mRNA and protein levels in GBM. The results show that SUZ12 gene expression is upregulated in most cancer cells and that decreased SUZ12 expression is associated with apoptosis induction and inhibition of cell proliferation, leading to decreased tumor growth. The findings of our study indicate that curcumin NPs reduce SUZ12 gene expression in cancer cells and subsequently induce apoptosis. Finally, these results suggest that SUZ12 gene expression is positively associated with GBM cancer, so its decrease induces apoptosis and inhibits cell proliferation.
The HOTAIR lncRNA gene is ubiquitously expressed and is associated with cancer metastasis, invasion, and progression (35).
HOTAIR can also alter gene expression at post-transcriptional levels by pairing with translation factors or ribosomes to control translation and binding to splicing-modulating factors (36). Studies have shown an association between the PRC2 complex and HOTAIR, demonstrating that PRC2 binds to the 5′ end of HOTAIR. HOTAIR regulates transcription silencing of HOXD locus genes and other genetic loci by binding to the 5' end of the PRC2 complex and localizing it to a specific site where H3K27 trimethylation and epigenetic silencing of gene expression occur (37). Extensive functional studies have shown that HOTAIR overexpression occurs in most human solid tumors. Current research indicates that HOTAIR is an adverse prognostic factor for the survival of patients with breast, colon, and glioma cancer, and increased expression of HOTAIR in patients is associated with increased metastasis (36, 38). Studies have shown that curcumin can affect the expression of the HOTAIR gene (39). Curcumin, by affecting HOTAIR, can play a role in the expression of different genes involved in cellular mechanisms. HOTAIR can act as a prognostic factor for the survival of glioma patients, as well as a biomarker in glioma and a regulator of cell cycle progression (40, 41).
In this study, we also examined the expression of the HOTAIR gene. The results showed a decrease in the expression of the HOTAIR gene in GBM cells treated with nanoparticle curcumin. HOTAIR is known as a biomarker with a poor prognosis in cancer. This lncRNA also plays a vital role in modulating epigenetic processes through interaction with DNMT enzymes and the PRC2 complex. Considering the interaction of HOTAIR with the PRC2 complex and its role in H3K27 trimethylation, and the effect of curcumin on the expression of these genes, it can be assumed that nanoparticle curcumin can disrupt these interactions and lead to a decrease in the expression of the HOTAIR gene. However, more research is needed to better understand its mechanisms.
In addition, the expression of MecP2 was also measured at the protein level, which was associated with a decrease in expression. The nuclear protein MecP2 is an epigenetic factor essential to cell identity and function (42). MecP2 is a member of the MBD protein family with different domains and isoforms. In general, the expression of MecP2 is increased in various cancers, and studies have shown that knocking down the MecP2 gene reduces the progression of cancer cells. Additionally, studies have shown that the high expression of MecP2 is associated with recruitment to the regulatory regions of tumor suppressor genes (42). In various studies, the expression level of MecP2 in tumor tissue compared to normal tissue has been investigated, and it has been shown that the expression of this gene is high in tumor tissue (43, 44). A study in pancreatic cancer showed that the increased expression of MecP2 in people had a positive relationship with their survival rate. Also, studies have shown that MecP2 expression is higher in normal tissues than in cancer tissues (45). The study of glioma cancer showed that MecP2 could be an essential factor in the epithelial-mesenchymal transition (EMT) process, so its inhibition reduced EMT. For this reason, MecP2 can be considered a potential therapeutic target in cancer treatment and an essential factor in cancer occurrence (46).
In this study, the results of examining MecP2 expression showed a decreased protein level. MecP2, as an epigenetic regulator, plays a vital role in tumorigenesis and cancer progression, especially in the EMT process. However, the study of MecP2 expression in different types of cancer has shown different results. In some studies, increased expression of MecP2 protein has been demonstrated in healthy tissue compared to cancerous tissue, which could indicate a complex and context-dependent role for this protein. Our study showed that curcumin may affect and disrupt cancer-related epigenetic pathways by affecting MecP2 expression. These findings indicate curcumin's clinical potential as a targeted cancer therapy, but further research is needed to better understand its mechanisms.
5.1. Conclusions
Finally, the results of this study show that C-NPs significantly reduce the expression of genes related to epigenetic processes, including DNMT3A, SUZ12, HOTAIR, and MecP2. Changes in the expression of these genes can play an essential role in inhibiting the growth and progression of GBM cancers. By reducing the expression of these genes and modulating epigenetic processes, curcumin NPs can serve as a promising therapeutic option in cancer treatment. Additionally, the effect of curcumin on the regulation of HOTAIR and its association with the PRC2 complex may effectively modulate epigenetic processes. These findings demonstrate the potential of curcumin as a targeted and combined therapy in cancer treatment, but further research is necessary to understand its molecular mechanisms more precisely.